ChemDFM-X: Multimodal Foundation Model for Chemistry
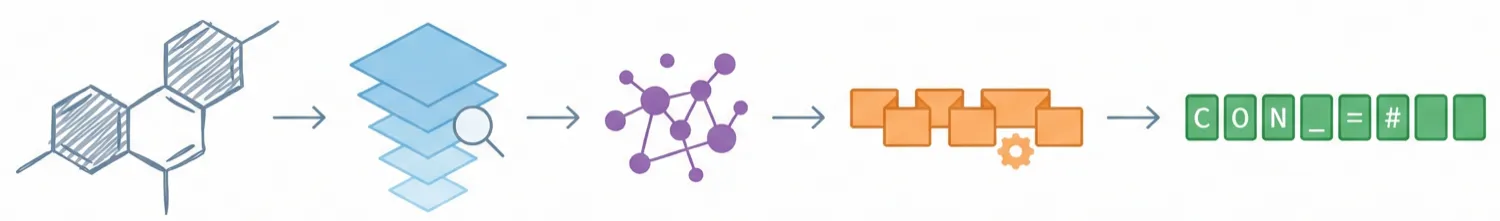
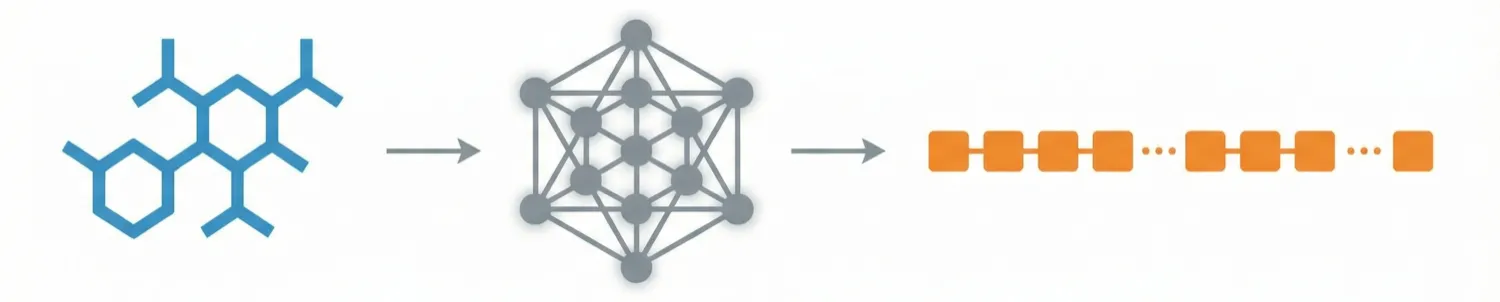
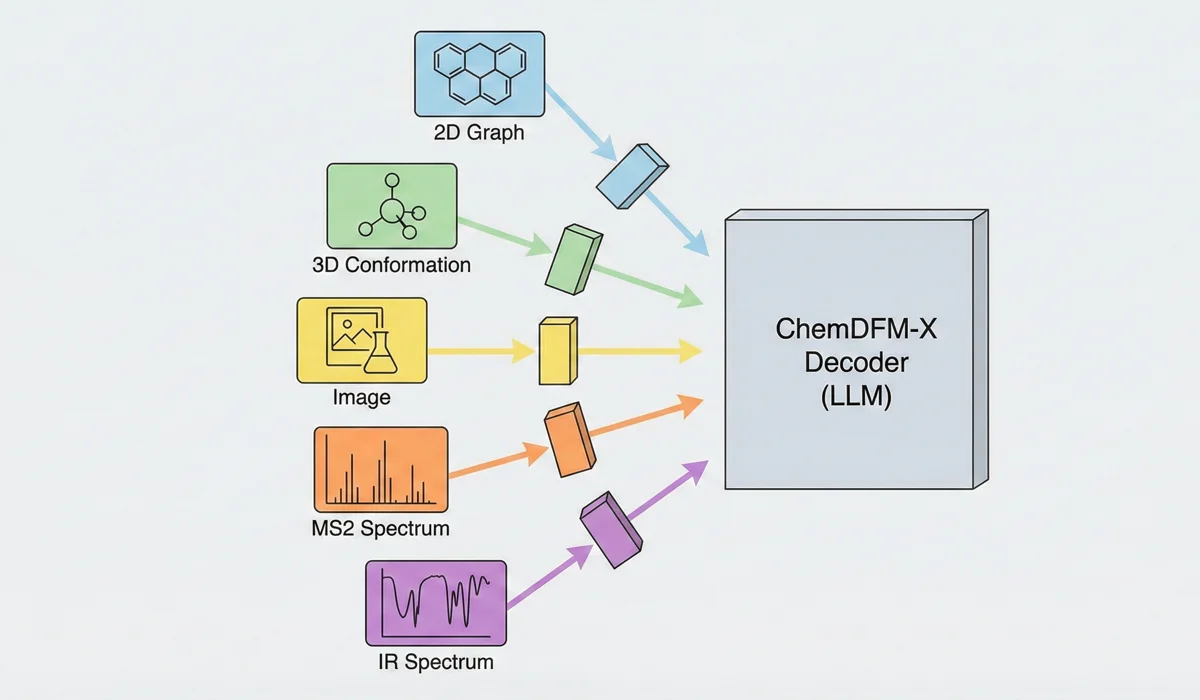
ChemDFM-X is a multimodal chemical foundation model that integrates five non-text modalities (2D graphs, 3D conformations, images, MS2 spectra, IR spectra) into a single LLM decoder. It overcomes data scarcity by generating a 7.6M instruction-tuning dataset through approximate calculations and model predictions, establishing strong baseline performance across multiple modalities.