OCSU: Optical Chemical Structure Understanding (2025)
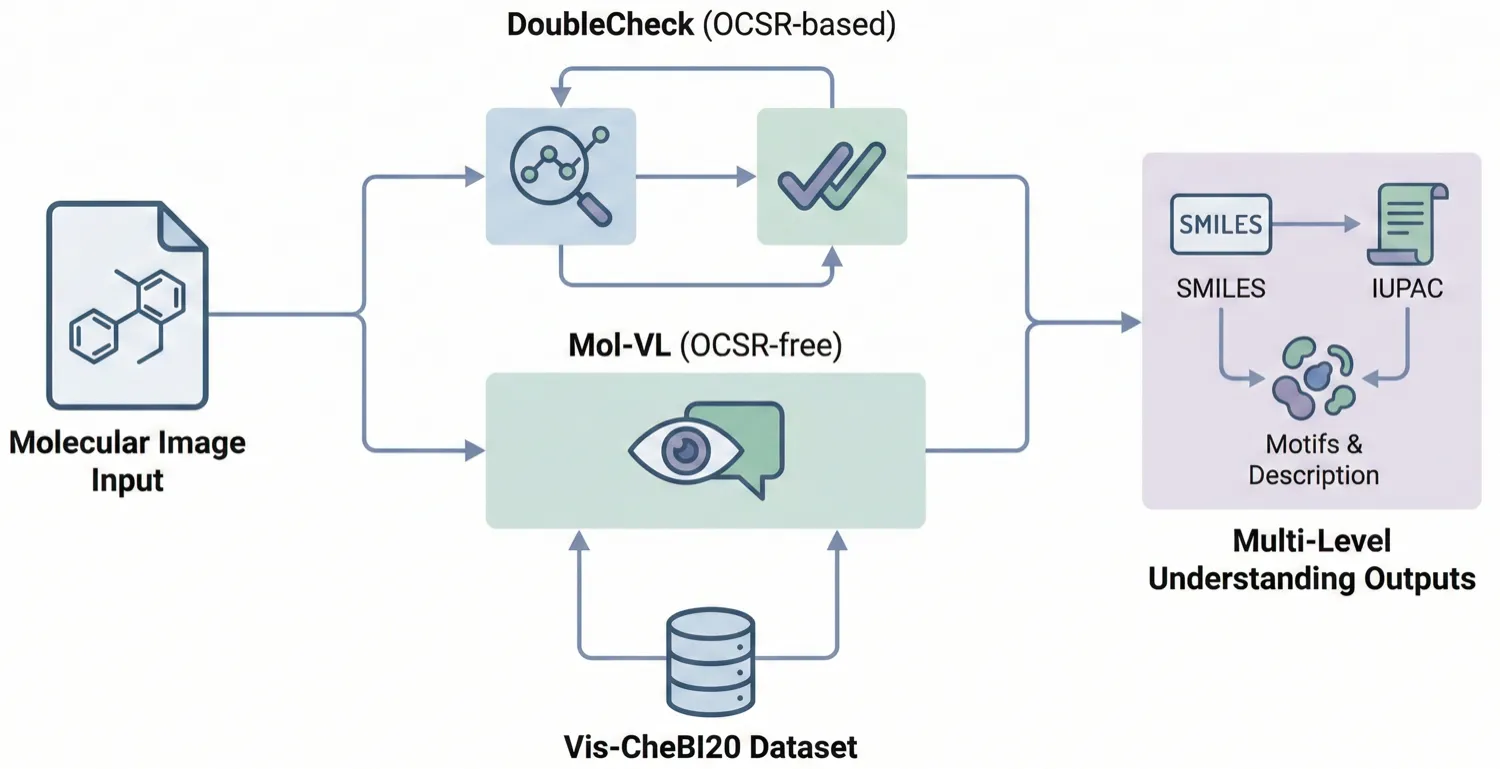
Proposes the ‘Optical Chemical Structure Understanding’ (OCSU) task to translate molecular images into multi-level descriptions (motifs, IUPAC, SMILES). Introduces the Vis-CheBI20 dataset and two paradigms: DoubleCheck (OCSR-based) and Mol-VL (OCSR-free).