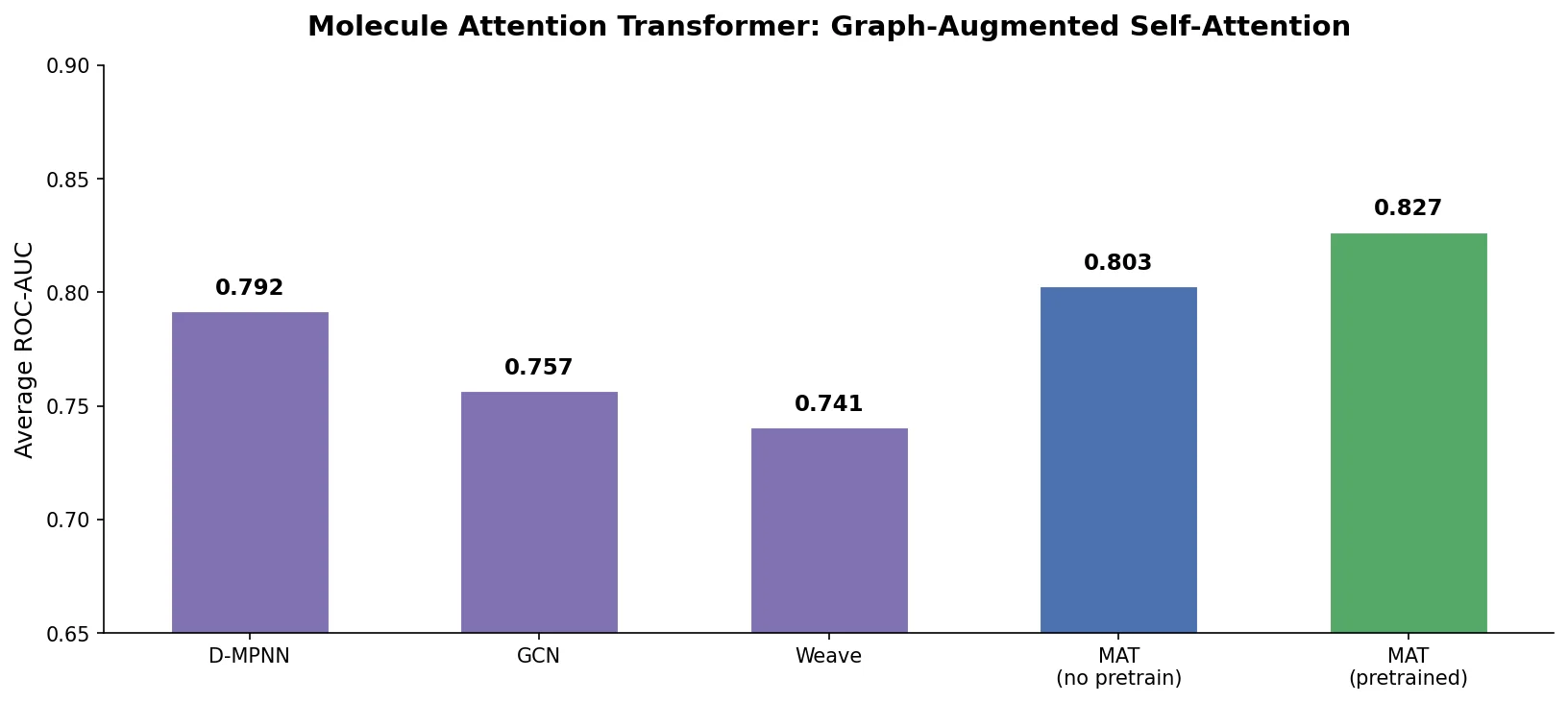
MAT: Graph-Augmented Transformer for Molecules (2020)
Molecule Attention Transformer (MAT) augments Transformer self-attention with inter-atomic distances and graph adjacency, achieving strong property prediction across diverse molecular tasks with minimal hyperparameter tuning after self-supervised pretraining.