MolSight: OCSR with RL and Multi-Granularity Learning
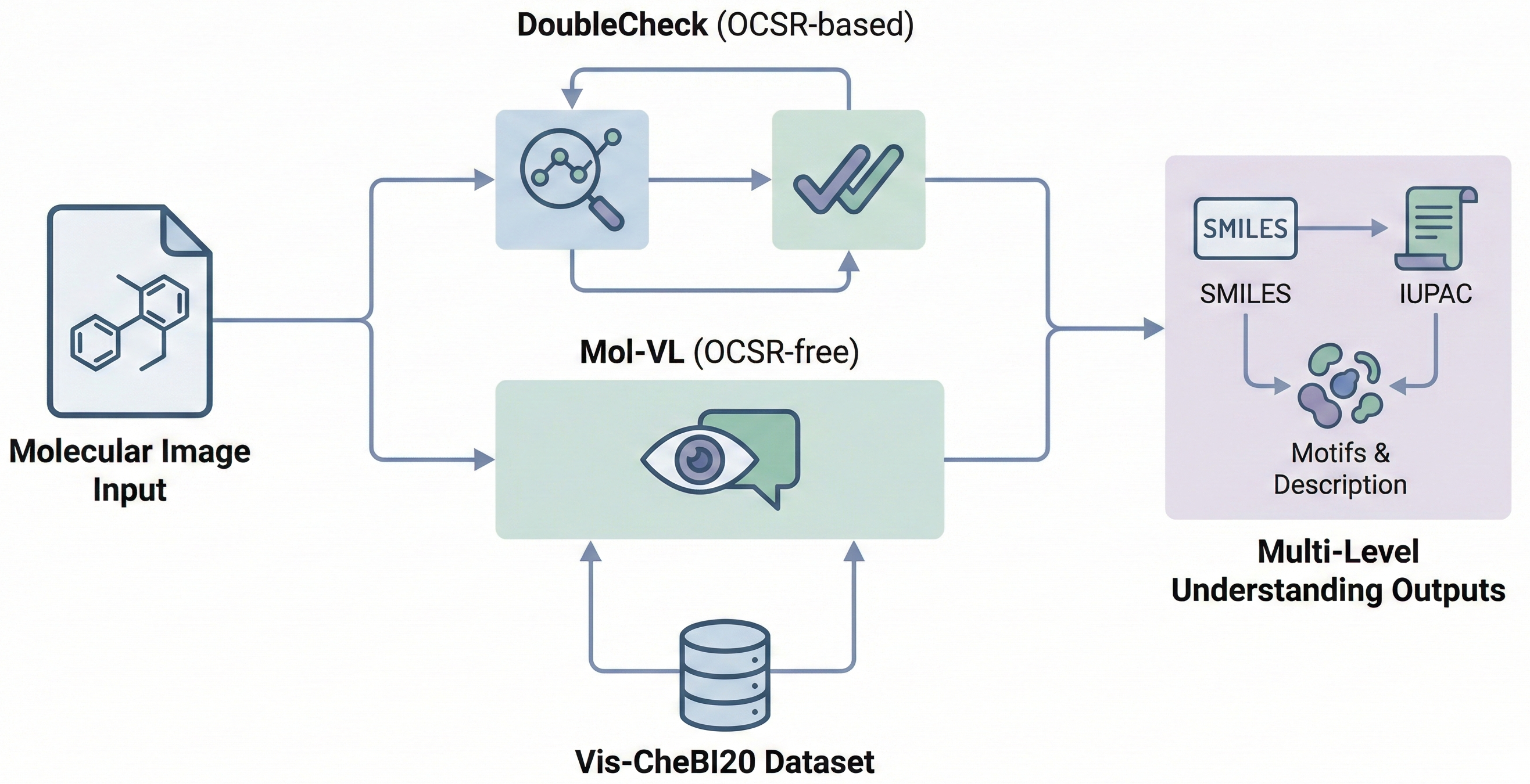
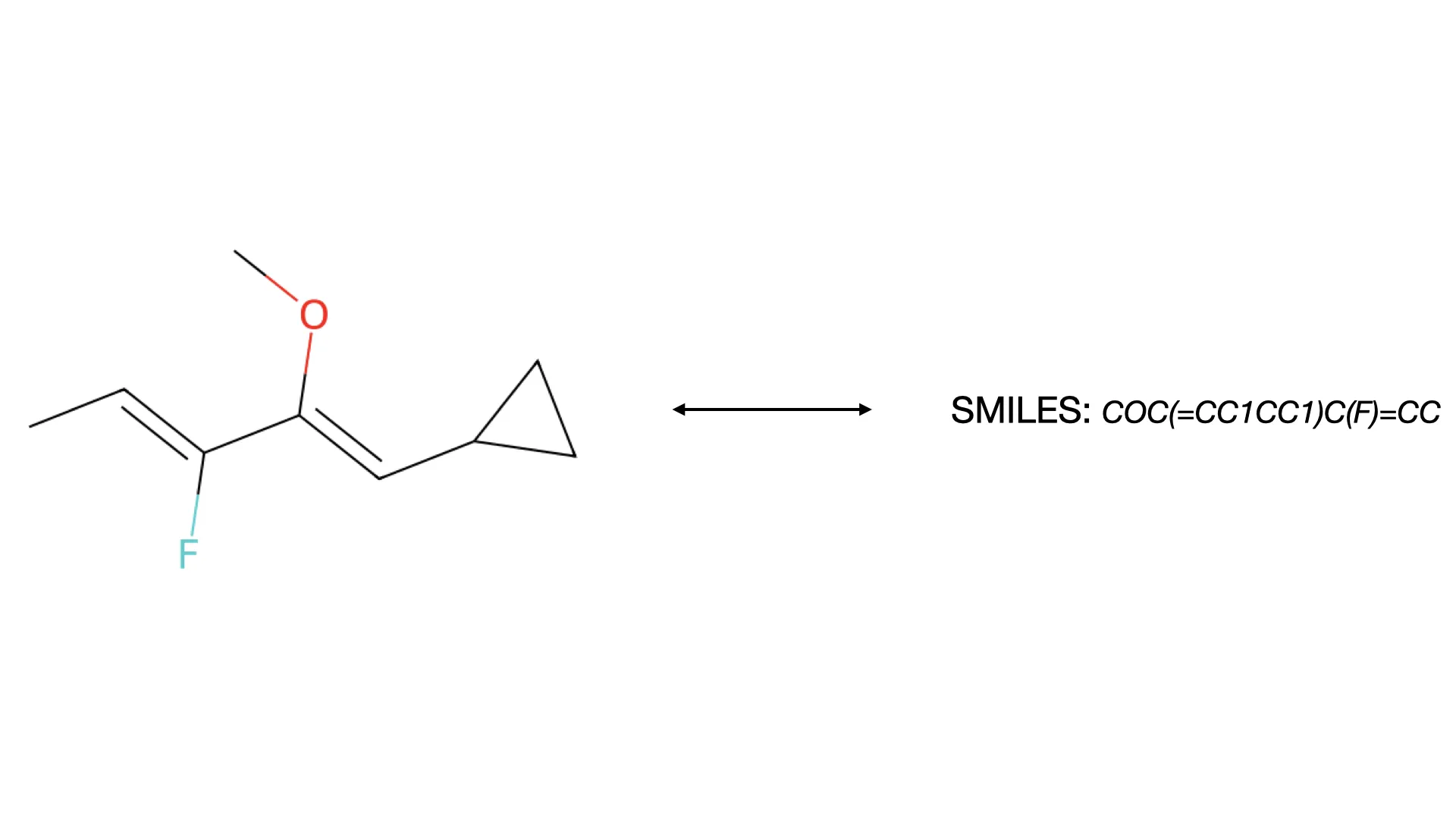
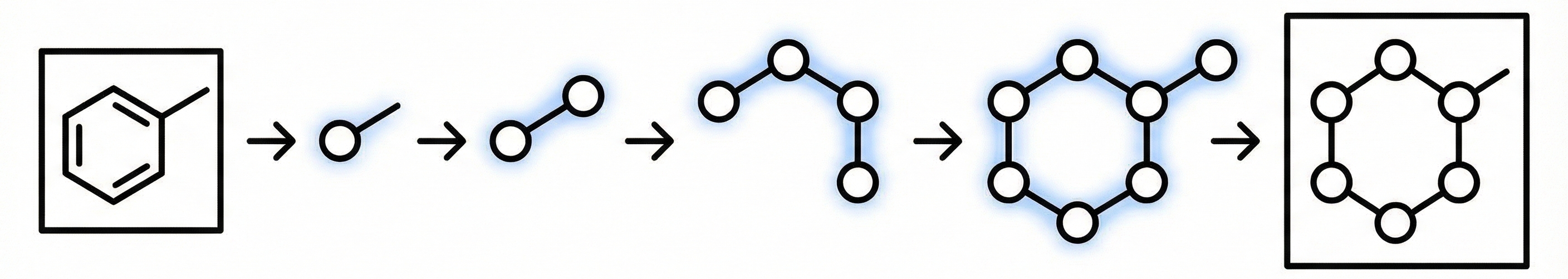
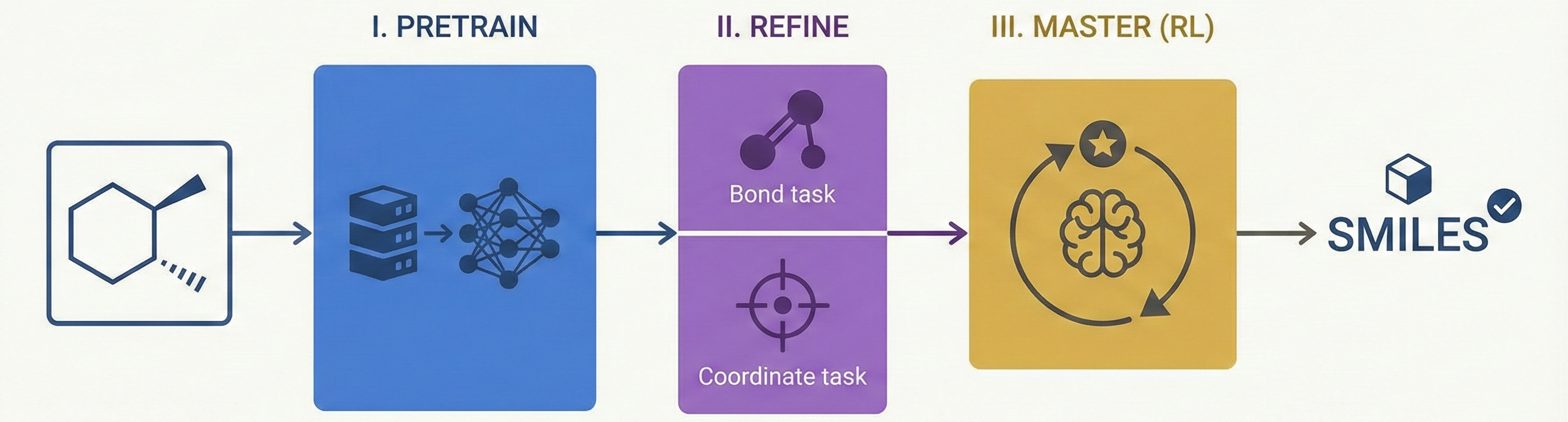
MolSight introduces a three-stage training paradigm for Optical Chemical Structure Recognition (OCSR), utilizing large-scale pretraining, multi-granularity fine-tuning with auxiliary bond and coordinate prediction tasks, and reinforcement learning (GRPO) to achieve state-of-the-art performance in recognizing complex stereochemical structures like chiral centers and cis-trans isomers.