InstructMol: Multi-Modal Molecular Assistant
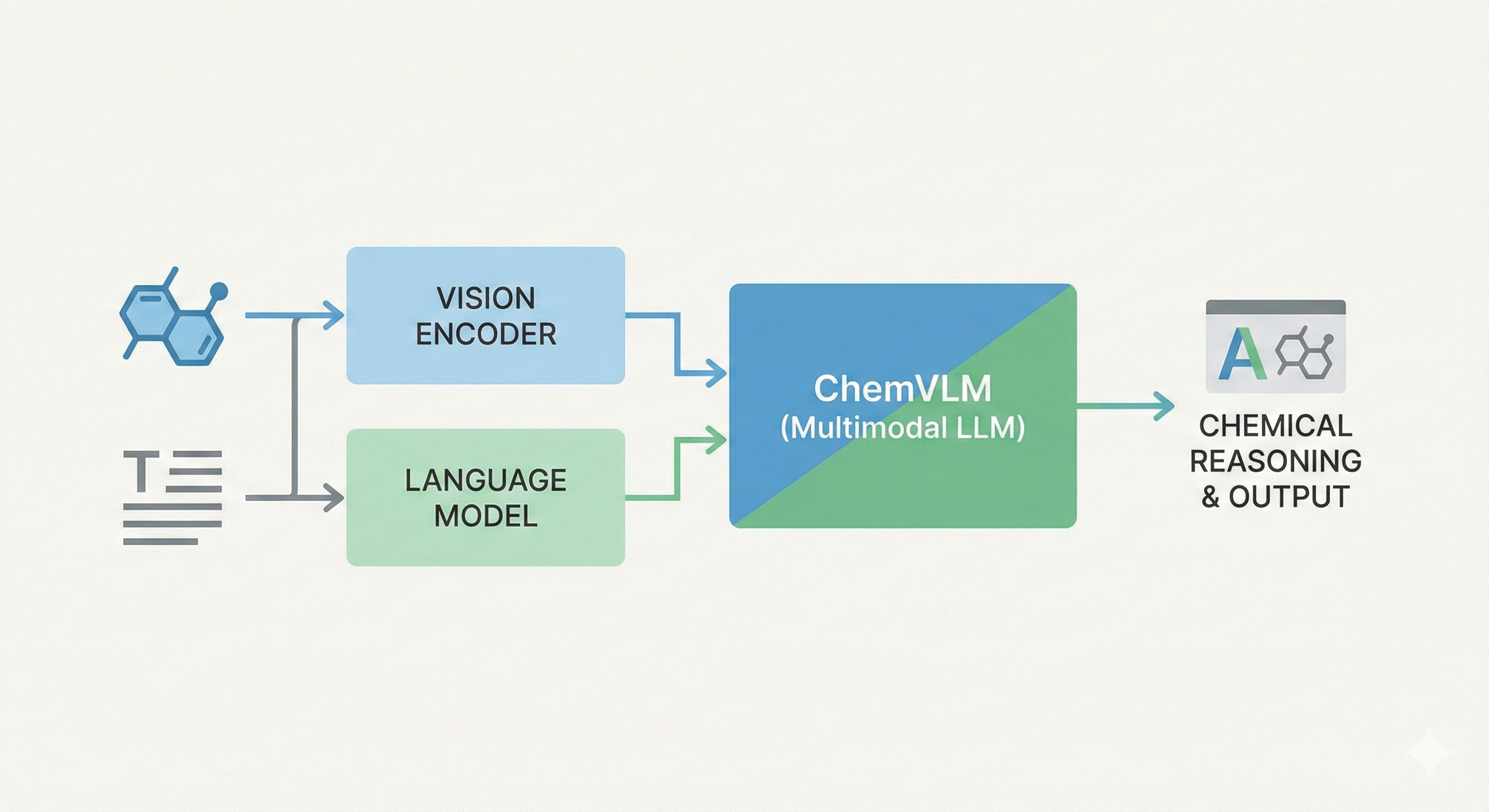
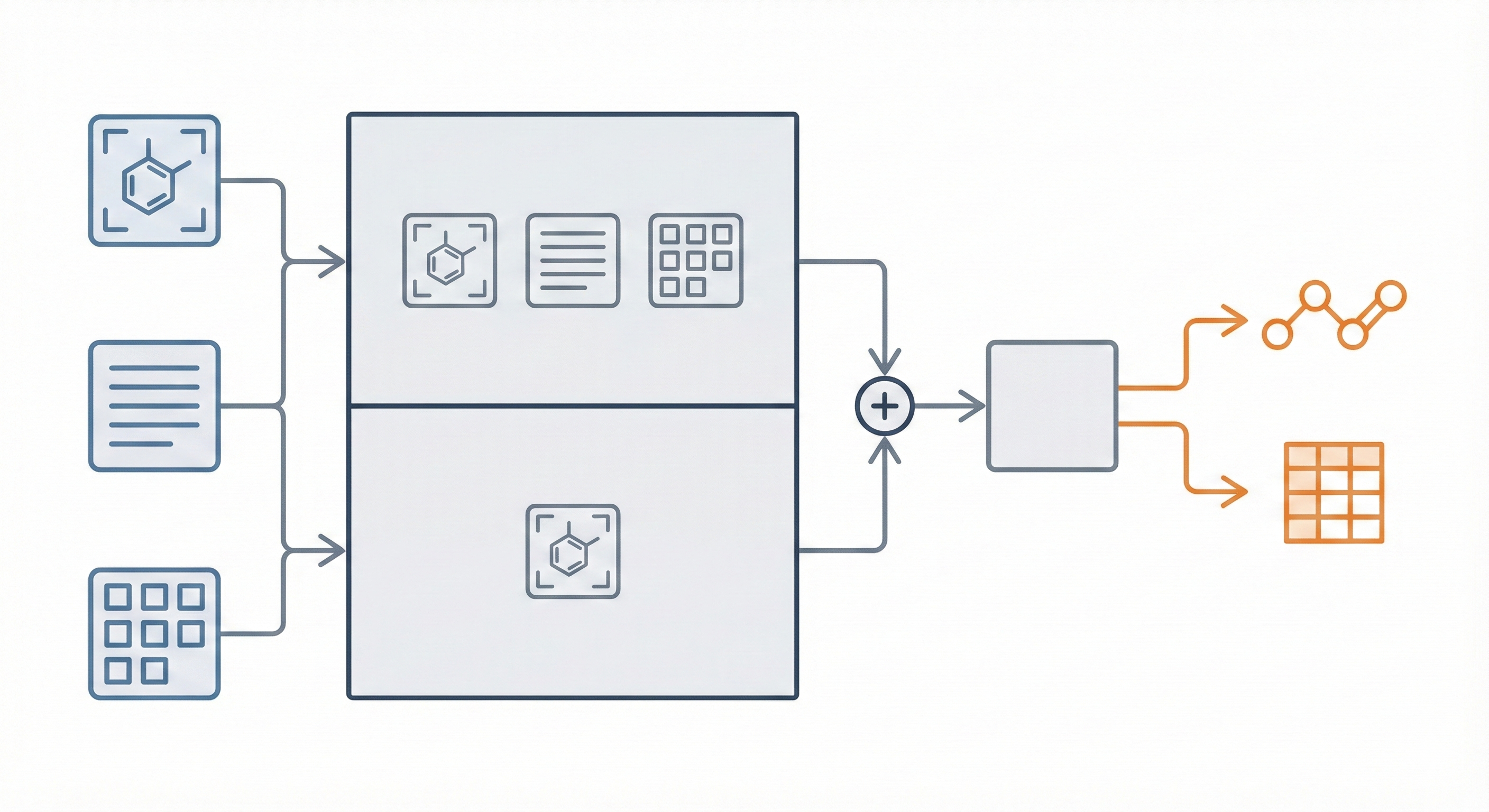
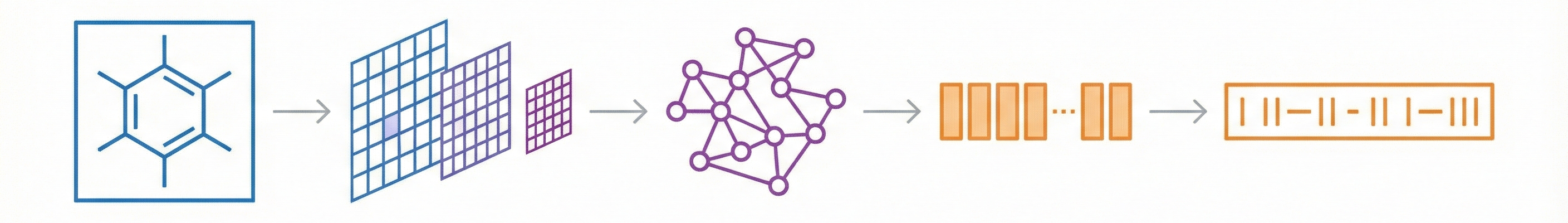
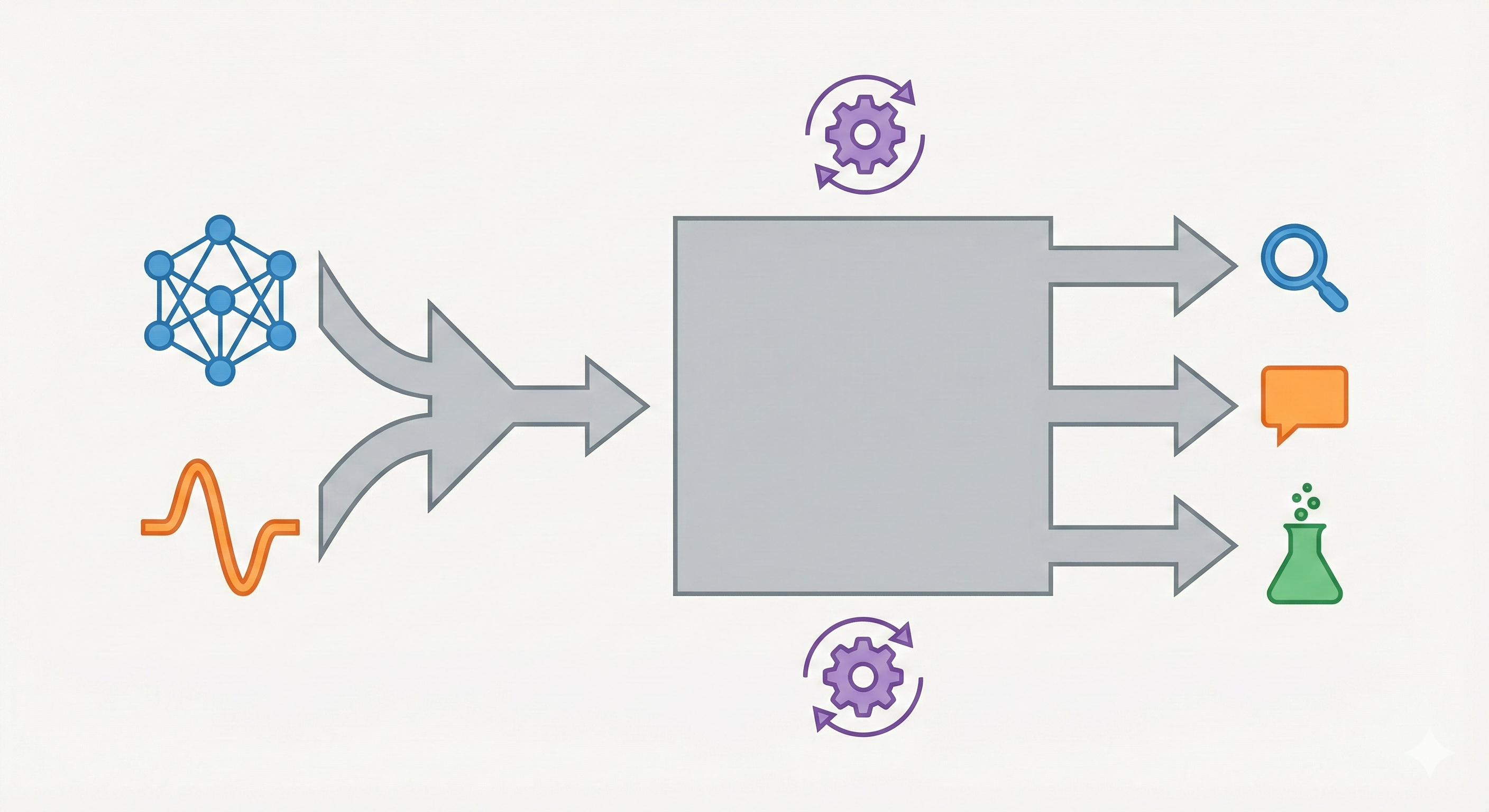
InstructMol integrates a pre-trained molecular graph encoder (MoleculeSTM) with a Vicuna-7B LLM using a linear projector. It employs a two-stage training process (alignment pre-training followed by task-specific instruction tuning with LoRA) to excel at property prediction, description generation, and reaction analysis.