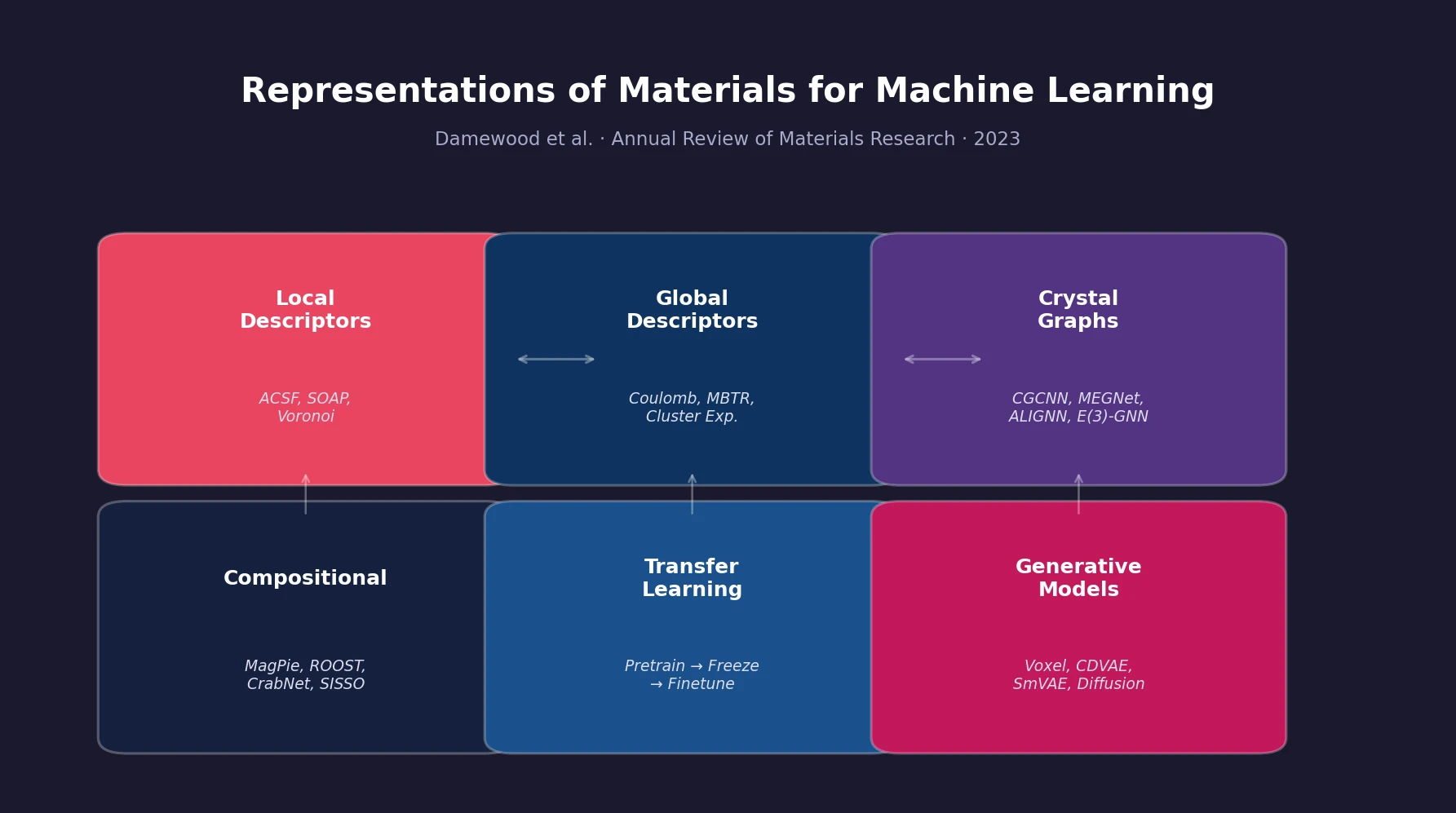
Materials Representations for ML Review
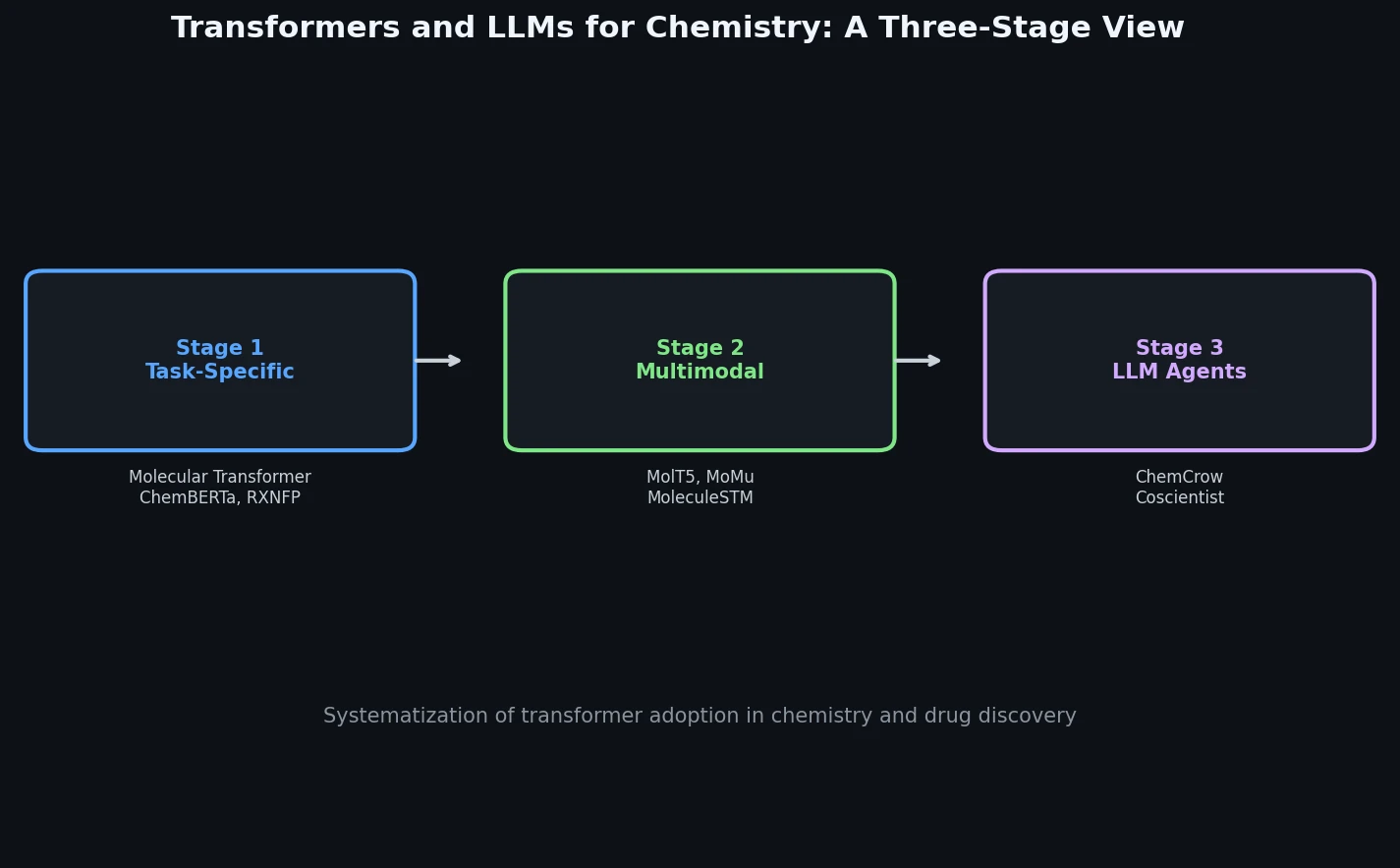
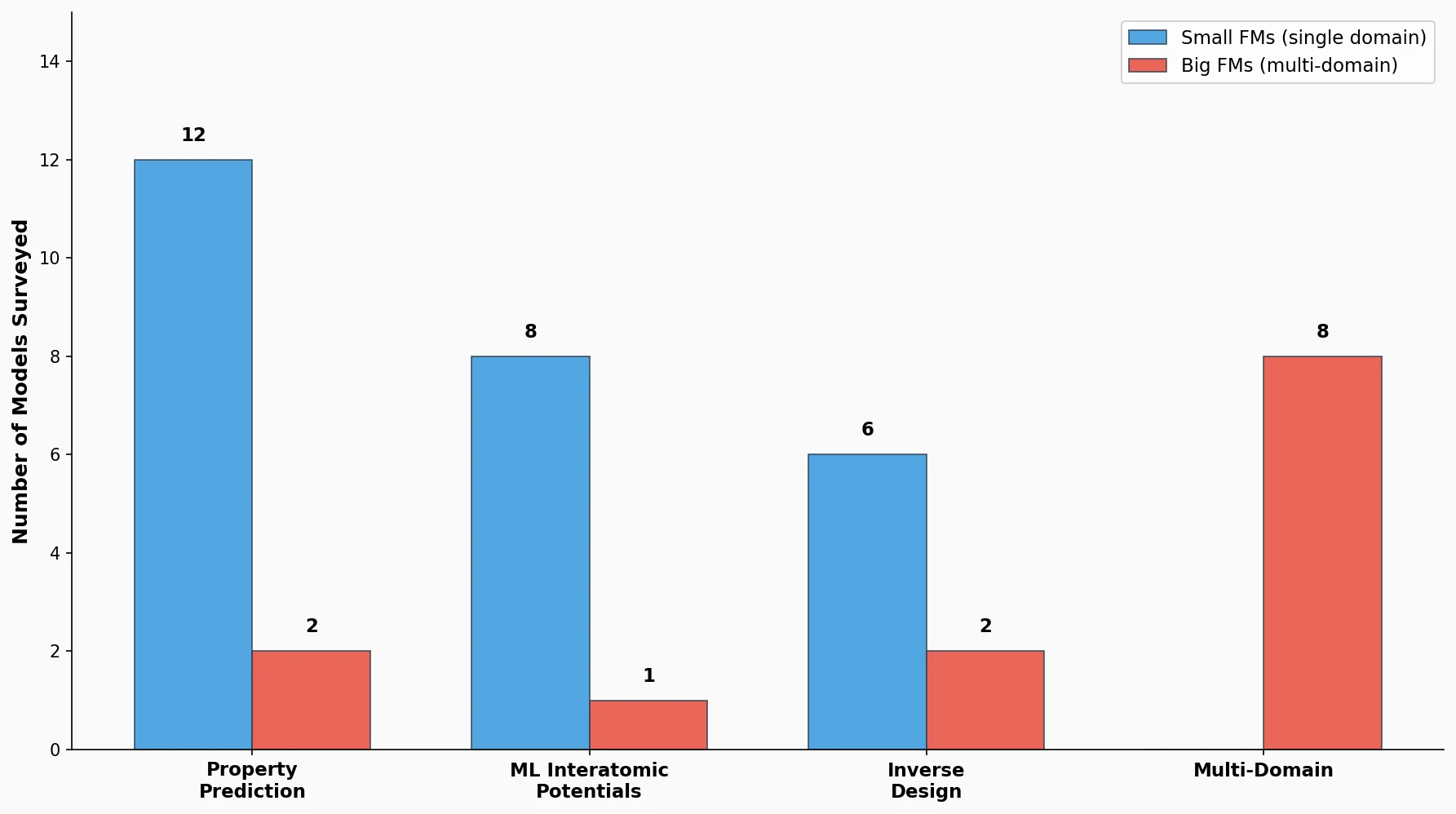
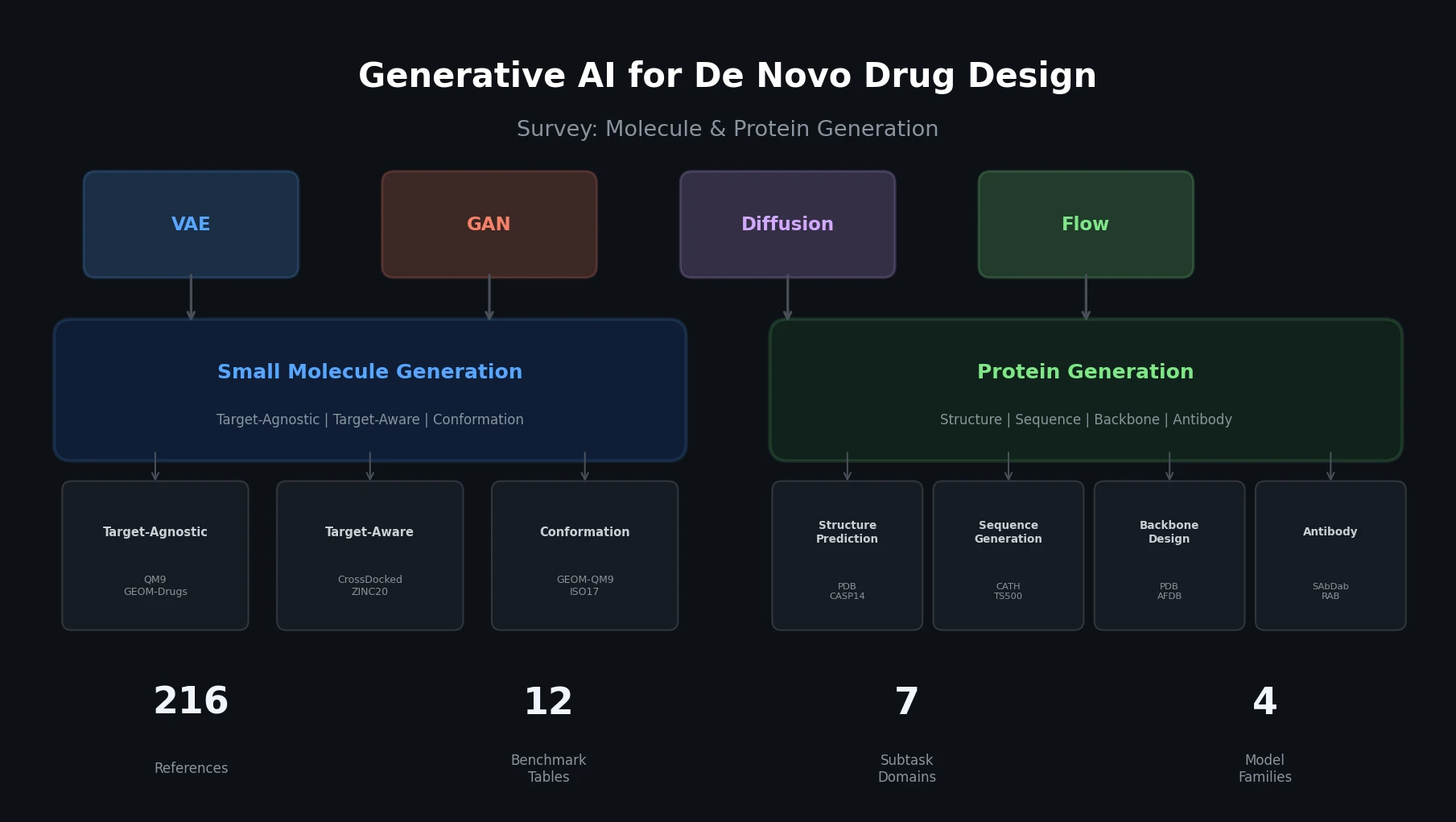
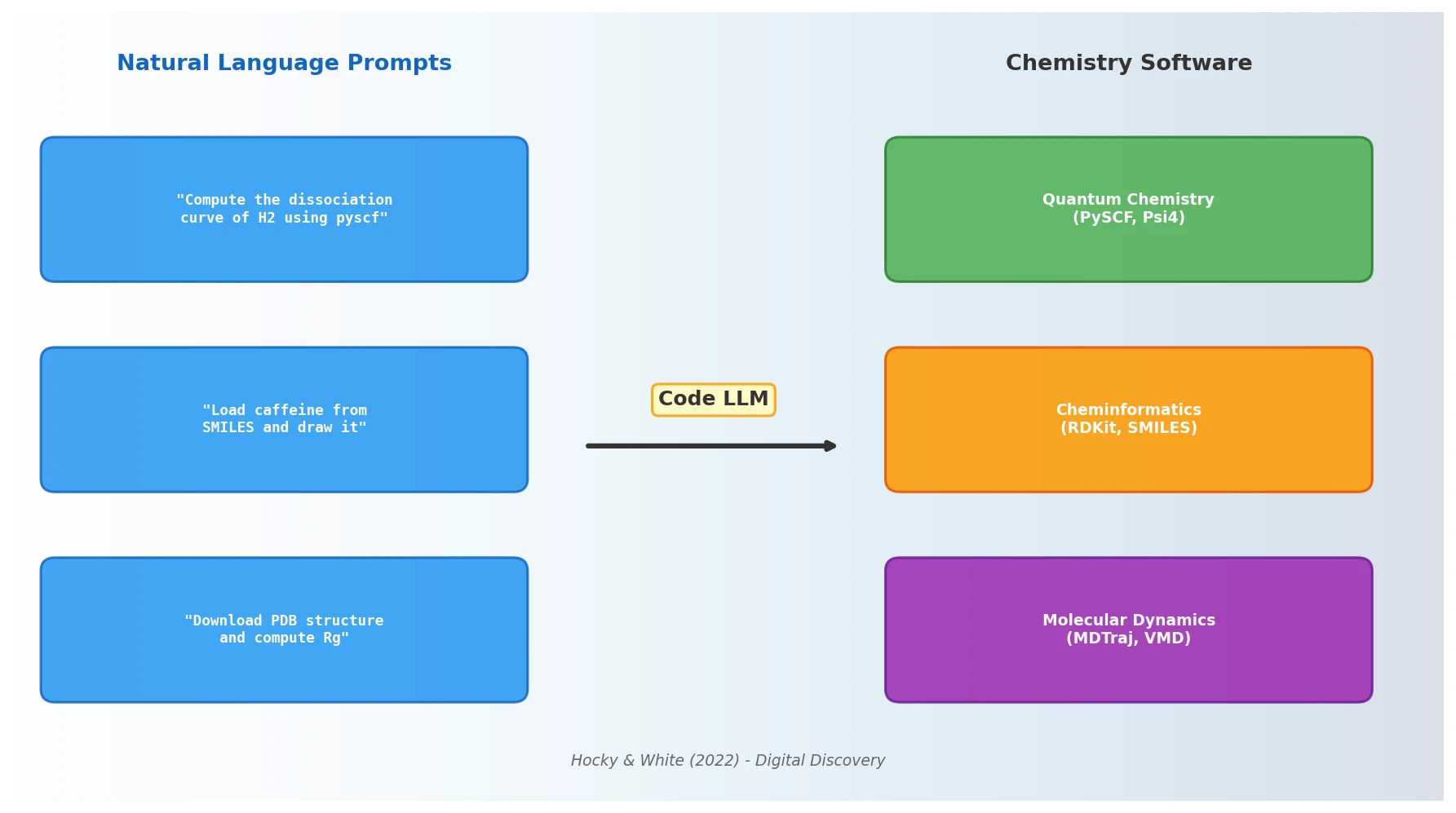
A comprehensive review of how solid-state materials can be numerically represented for machine learning, spanning structural features, graph neural networks, compositional descriptors, transfer learning, and generative models for inverse design.