Liquid Argon: LAMMPS Simulation
Explore the molecular dynamics of liquid argon in this fundamental LAMMPS simulation. This classic system demonstrates liquid-state behavior and serves as a benchmark for molecular dynamics methods.

Explore the molecular dynamics of liquid argon in this fundamental LAMMPS simulation. This classic system demonstrates liquid-state behavior and serves as a benchmark for molecular dynamics methods.
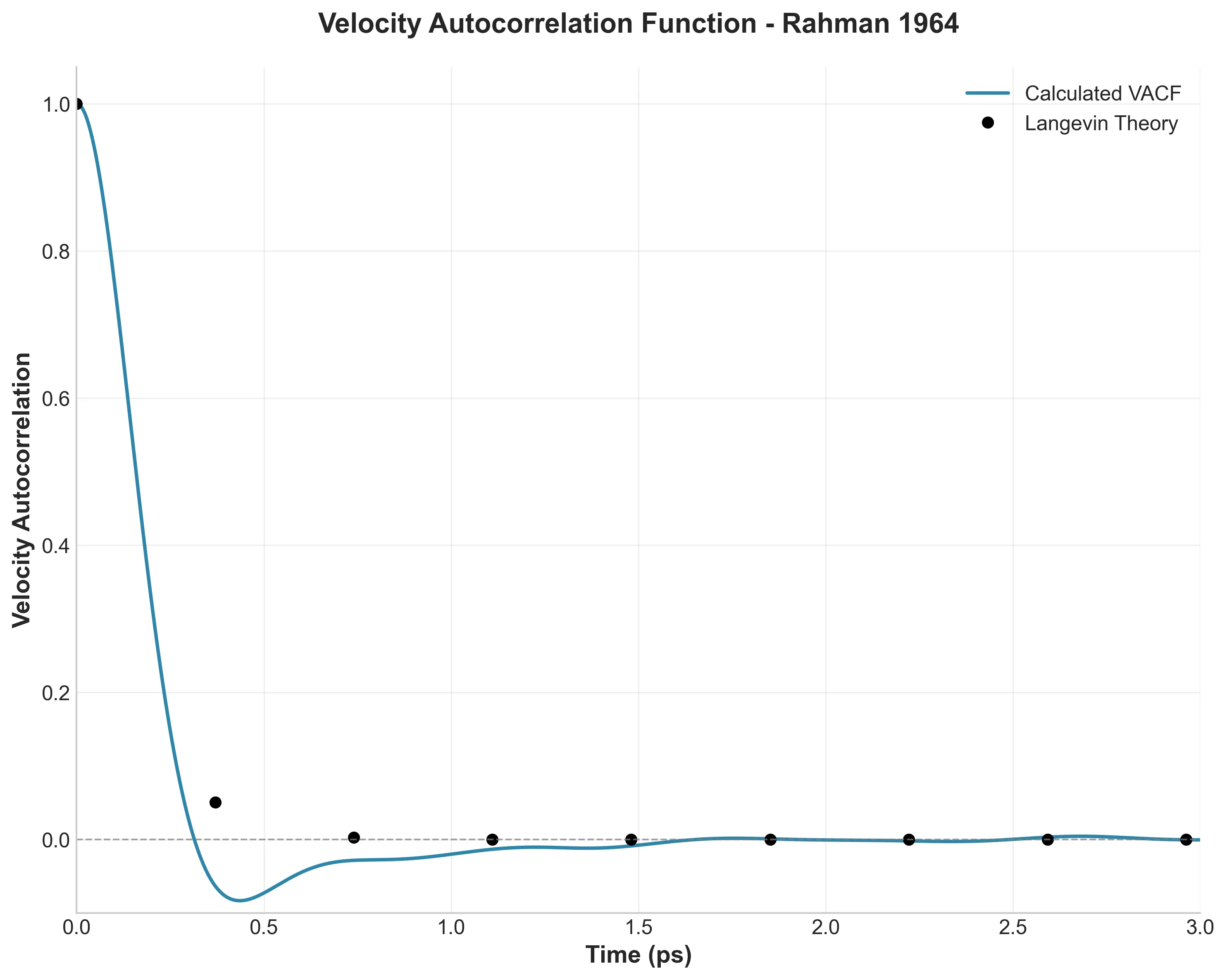
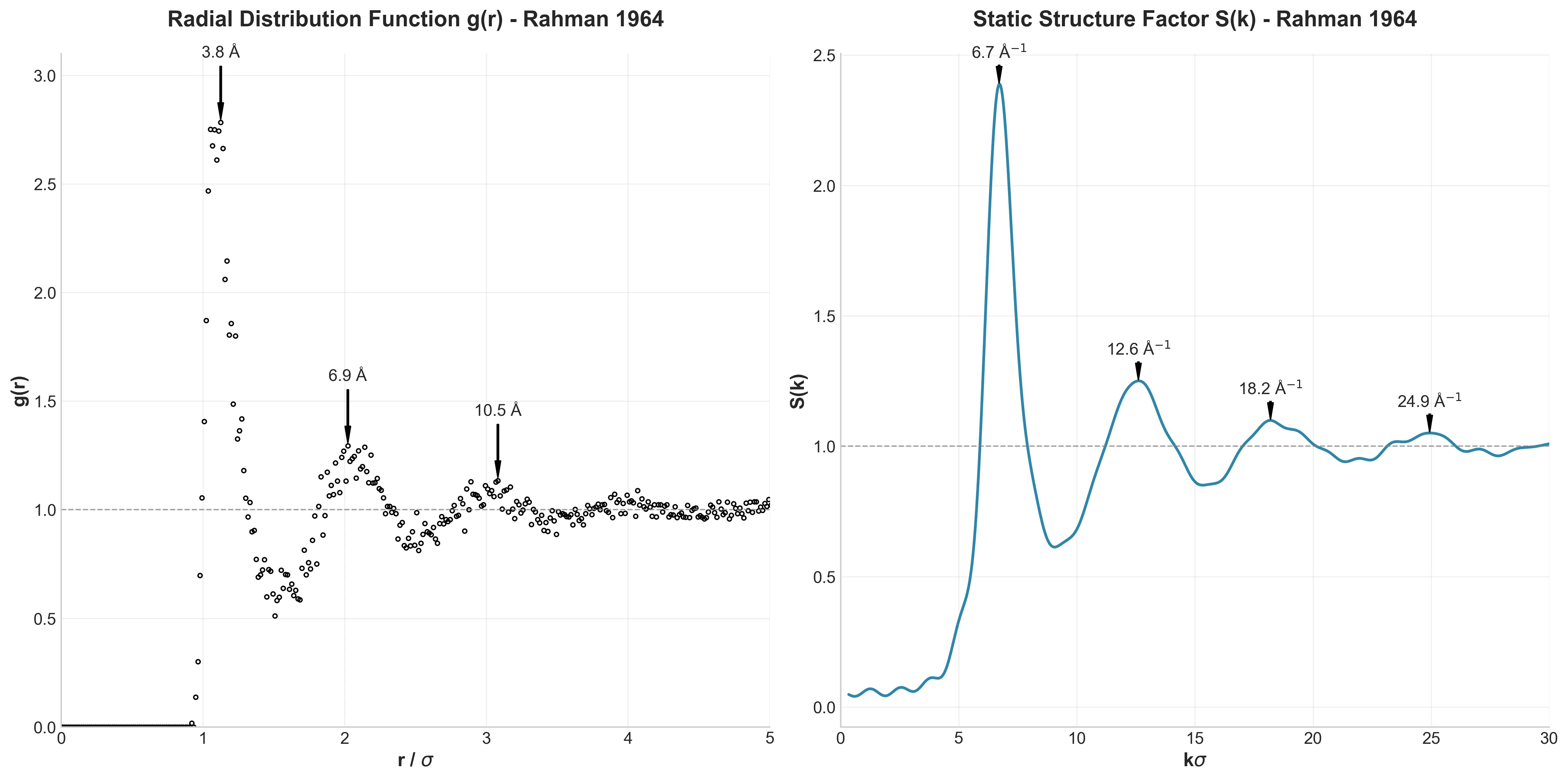
A digital restoration of Rahman’s seminal 1964 molecular dynamics paper using LAMMPS and a production-grade Python analysis pipeline featuring intelligent decorator-based caching, fully vectorized NumPy computations for O(N^2) operations, and modern tooling (uv, type hints, Makefile automation) transforming academic scripts into reproducible research toolkit.

I replicated Rahman’s landmark 1964 liquid argon molecular dynamics simulation using modern tools, building a production-grade Python analysis pipeline with intelligent caching, vectorization, and type safety to bridge vintage science with contemporary software engineering.
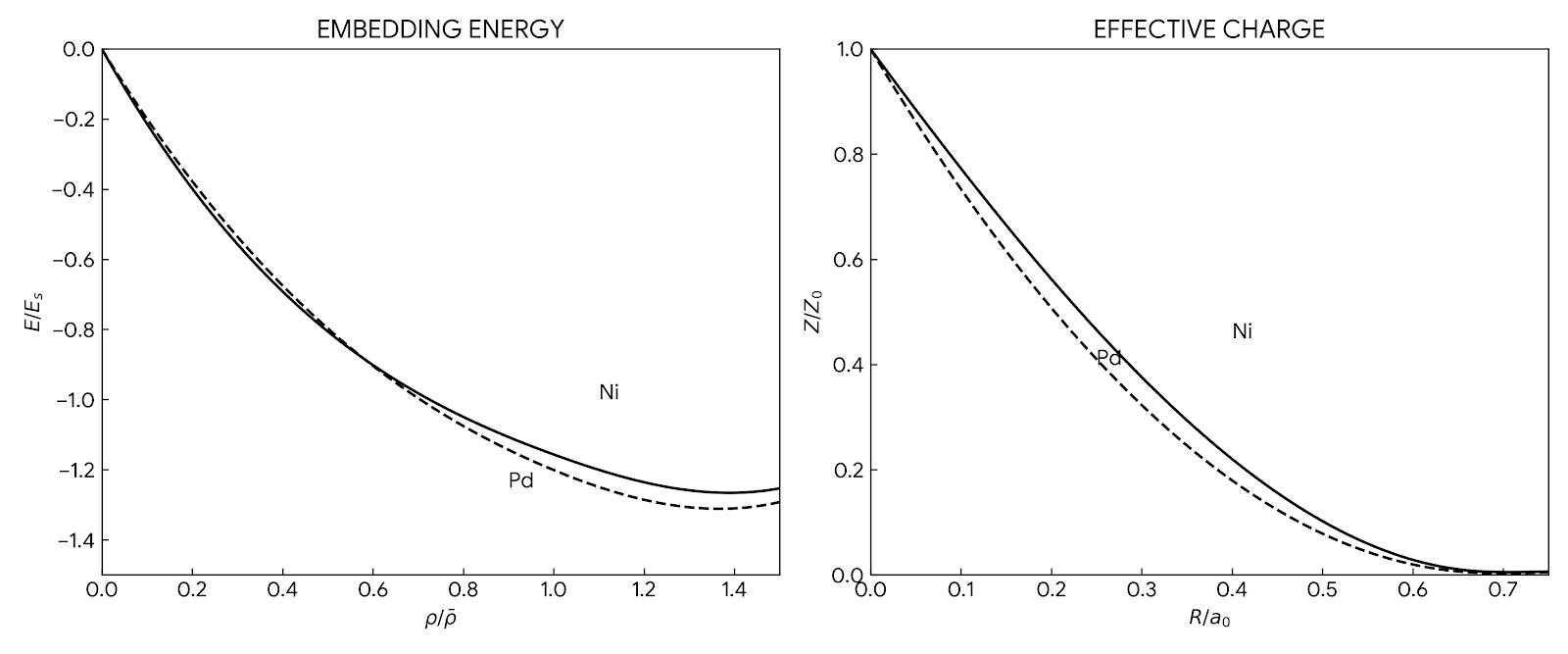
The foundational 1984 paper introducing EAM, a semi-empirical many-body interatomic potential that incorporates density functional theory concepts to accurately simulate metallic systems while maintaining computational efficiency comparable to pair potentials.
Torrie and Valleau’s 1977 paper introducing importance sampling with non-physical distributions to overcome the sampling gap problem in Monte Carlo free-energy calculations, particularly for phase transitions.
Watch copper atoms move across a crystal surface in this molecular dynamics simulation. This video demonstrates surface diffusion mechanisms important for understanding catalysis and crystal growth processes.

Visualize platinum atom diffusion on crystal surfaces in this LAMMPS molecular dynamics simulation. Understand surface mobility mechanisms crucial for catalysis and materials design.
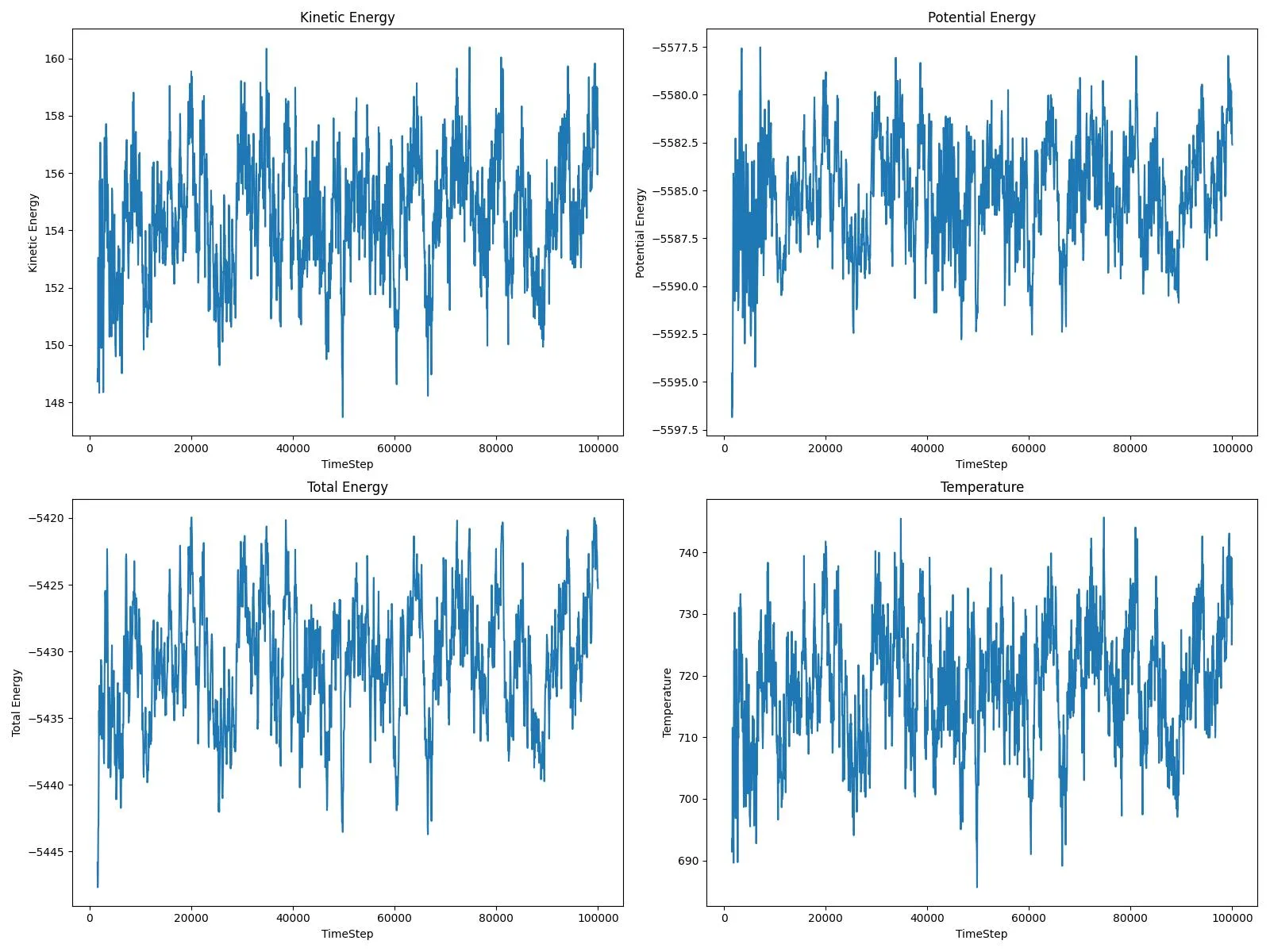
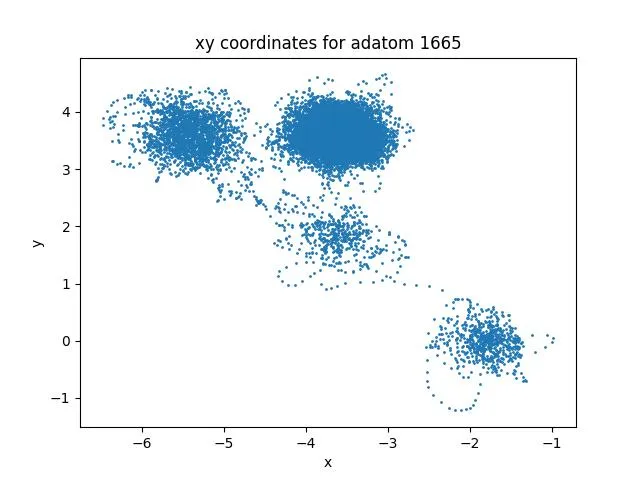
A complete input-to-analysis workflow for simulating adatom diffusion on FCC metal surfaces using LAMMPS and EAM potentials, providing comparative datasets for copper and platinum that demonstrate how atomic mass and bonding strength affect surface dynamics, with automated Python analysis generating publication-ready visualizations.
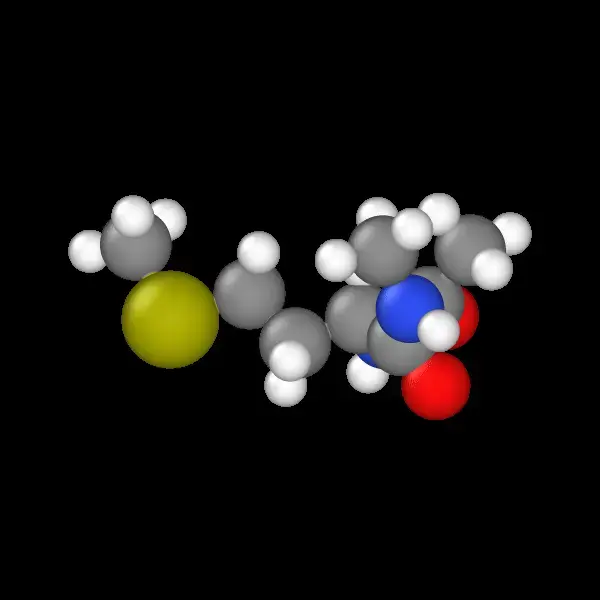
A practical guide to simulating mini-proteins using GROMACS; from alanine dipeptide to tryptophan systems for ML training data generation.

An automated GROMACS pipeline for generating high-fidelity molecular dynamics datasets suitable for machine learning, simulating capped dipeptides across nine residue types with 0.1 ps resolution and atomic force extraction optimized for training Neural Network Potentials.
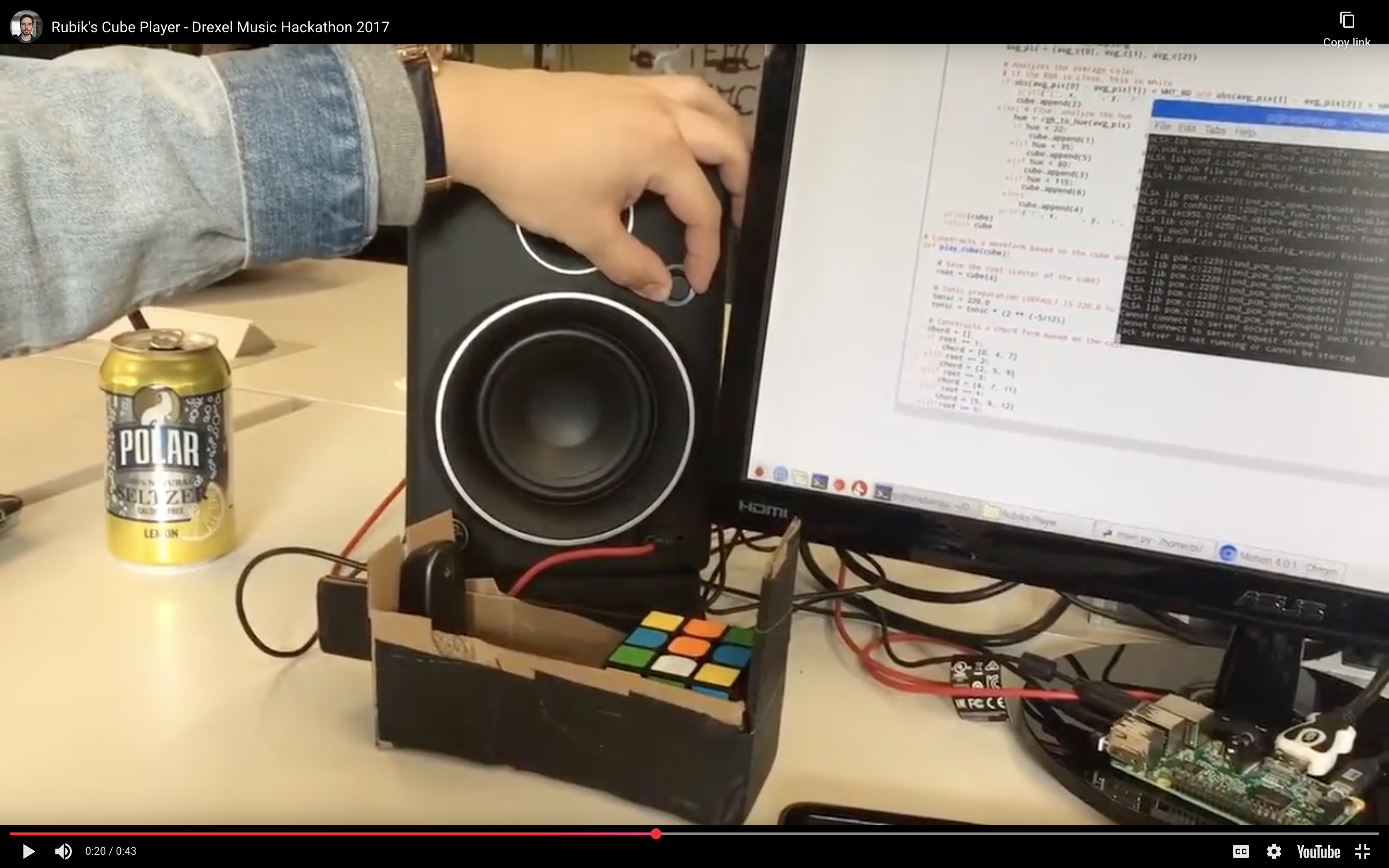
A project I built with Emmanuel Espino and Jason Zogheb at the 2017 Drexel Music Hackathon. It uses computer vision to read a Rubik’s cube and generates music based on how solved each face is.

An audio reactive music video created using Processing, where the visuals are generated from audio information in real-time.