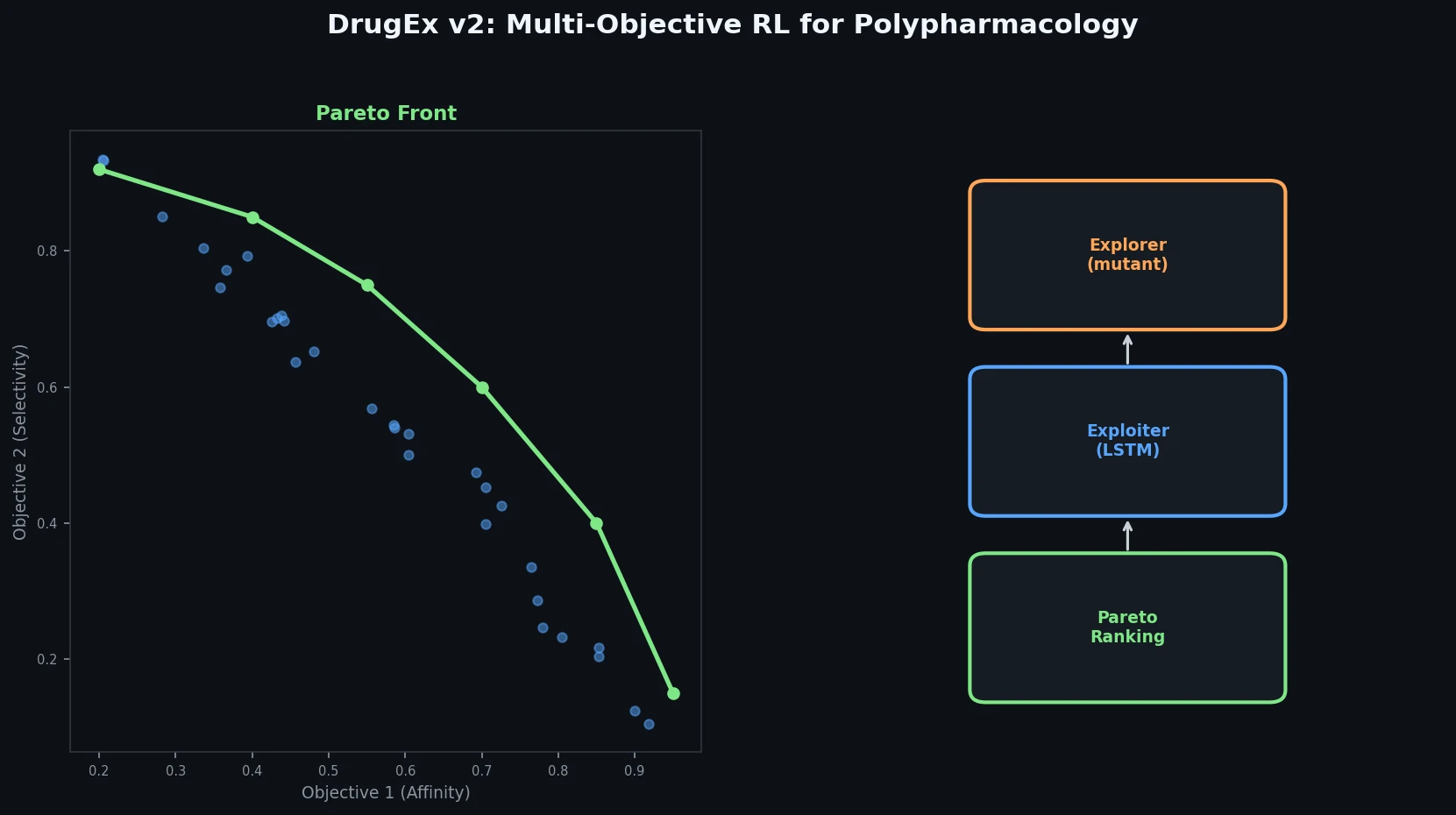
DrugEx v2: Pareto Multi-Objective RL for Drug Design
DrugEx v2 introduces Pareto-based multi-objective optimization and evolutionary exploration strategies into an RNN reinforcement learning framework for de novo drug design toward multiple protein targets.