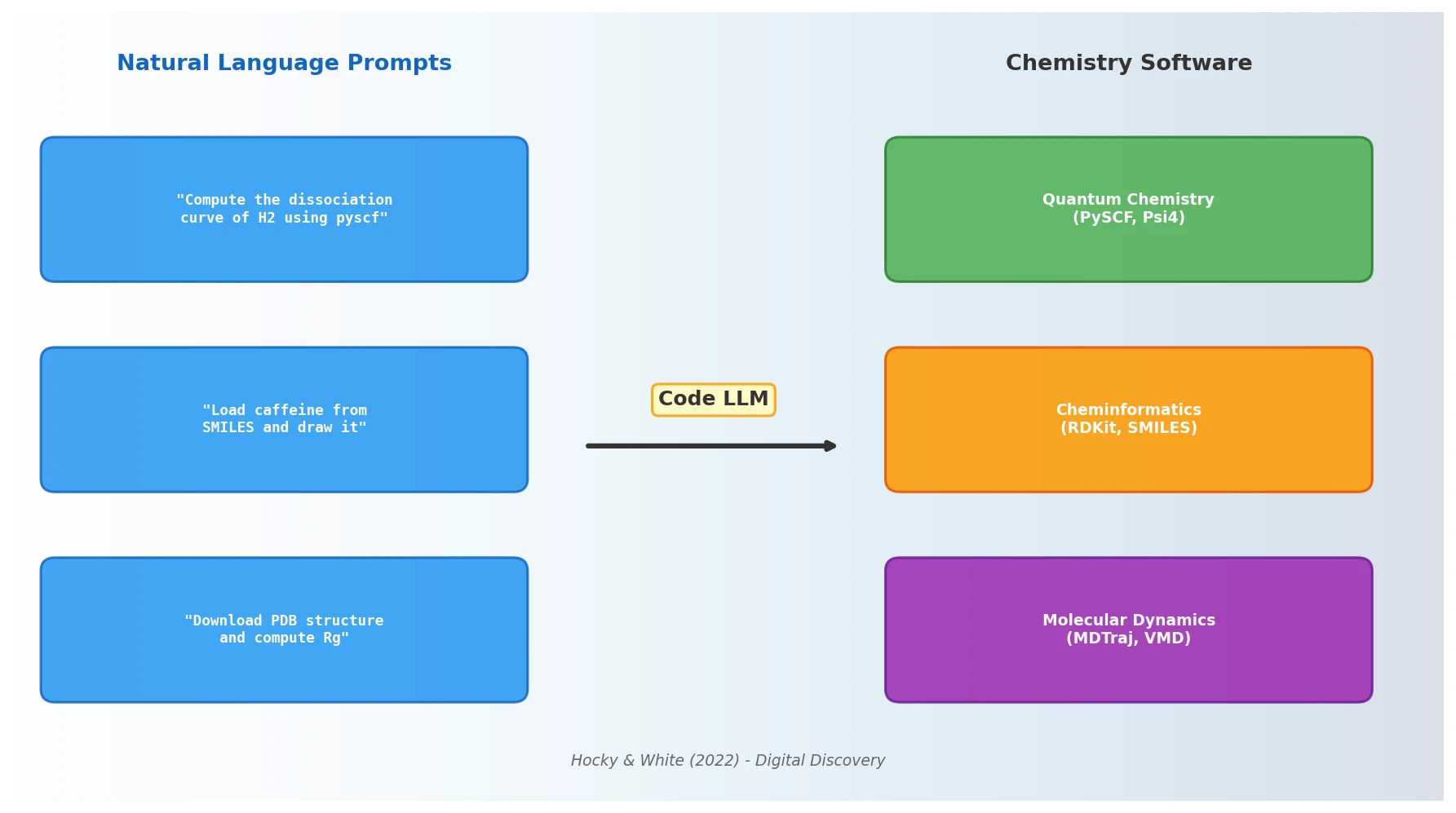
NLP Models That Automate Programming for Chemistry
Hocky and White argue that NLP models capable of generating code from natural language prompts will fundamentally alter how chemists interact with scientific software, reducing barriers to computational research and reshaping programming pedagogy.