SMINA Docking Benchmark for De Novo Drug Design Models
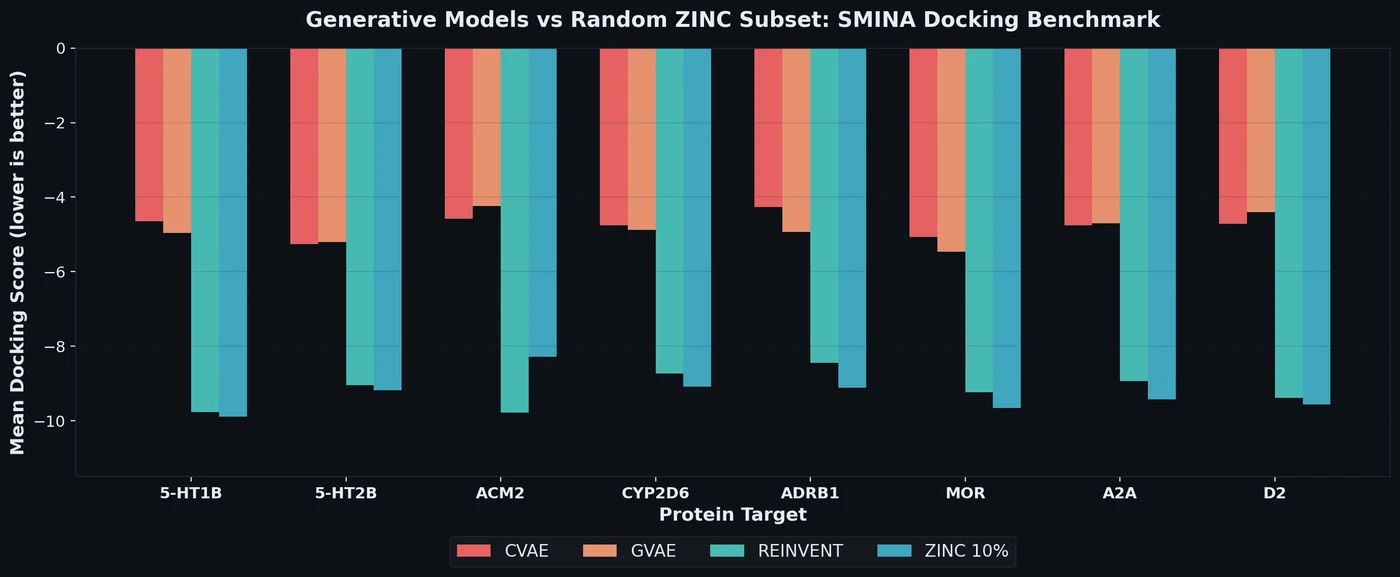
Proposes a benchmark for de novo drug design using SMINA docking scores across eight drug targets, revealing that popular generative models fail to outperform random ZINC subsets.

Proposes a benchmark for de novo drug design using SMINA docking scores across eight drug targets, revealing that popular generative models fail to outperform random ZINC subsets.
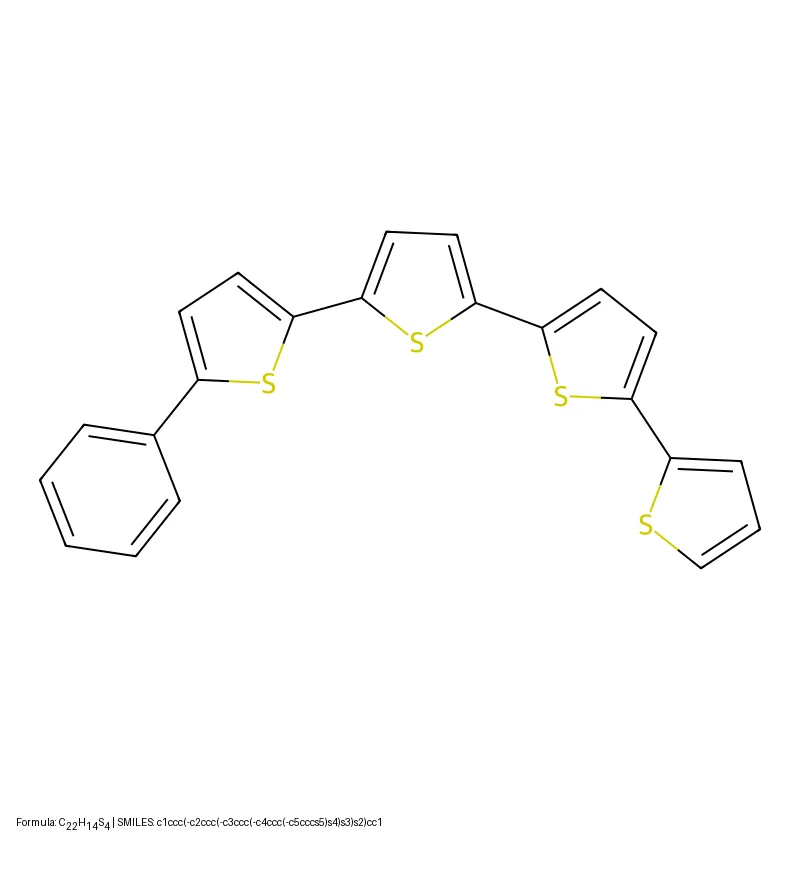
Tartarus introduces a modular suite of realistic molecular design benchmarks grounded in computational chemistry simulations. Benchmarking eight generative models reveals that no single algorithm dominates all tasks, and simple genetic algorithms often outperform deep generative models.
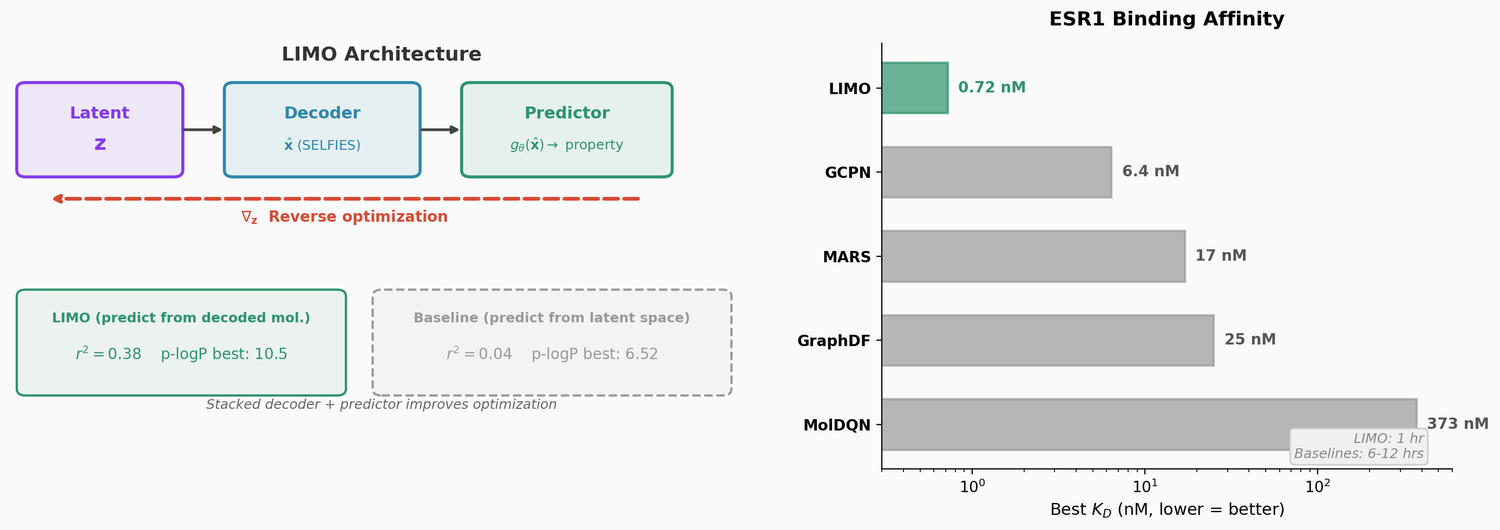
LIMO combines a SELFIES-based VAE with a novel stacked property predictor architecture (decoder output as predictor input) and gradient-based reverse optimization on the latent space. It is 6-8x faster than RL baselines and 12x faster than sampling methods while generating molecules with nanomolar binding affinities, including a predicted KD of 6e-14 M against the human estrogen receptor.
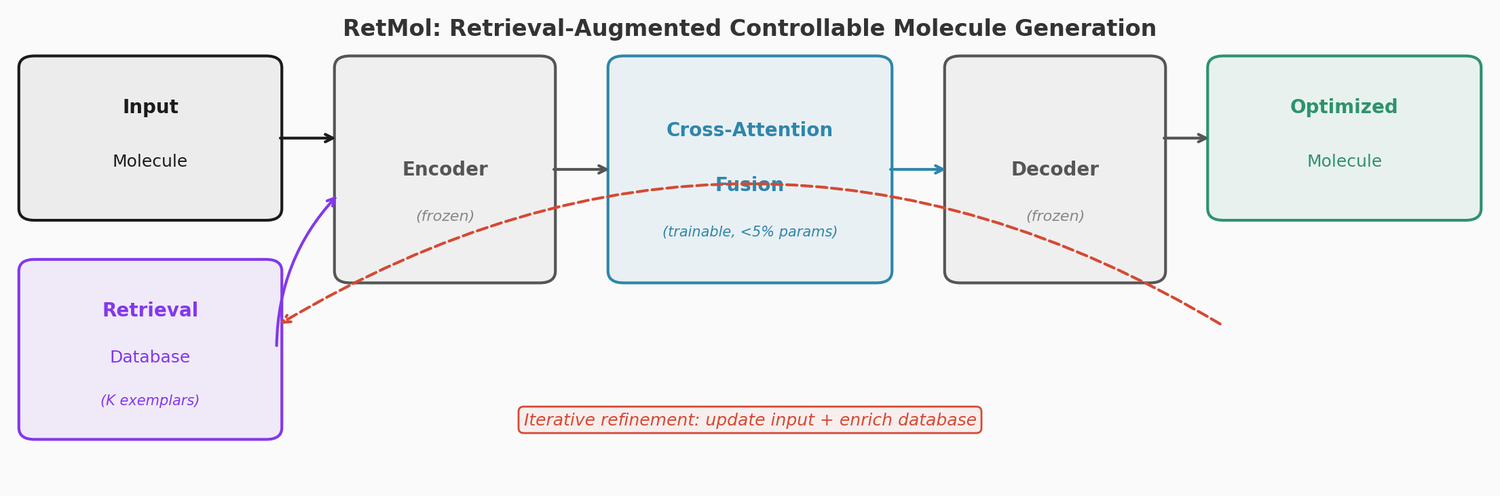
RetMol plugs a lightweight cross-attention retrieval module into a pre-trained Chemformer backbone to guide molecule generation toward multi-property design criteria. It requires no task-specific fine-tuning and works with as few as 23 exemplar molecules. It achieves 94.5% success on QED optimization, 96.9% on GSK3b/JNK3 dual inhibitor design, and 2.84 kcal/mol average binding affinity improvement on SARS-CoV-2 main protease inhibitor optimization.
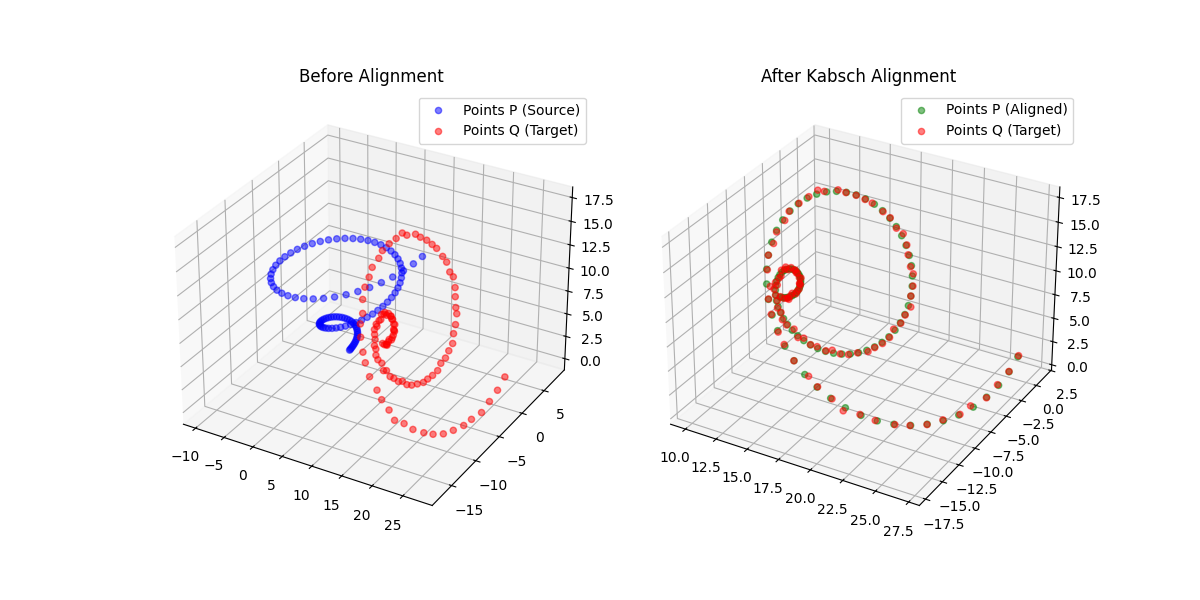
A differentiable point-set alignment library implementing N-dimensional Kabsch, Horn quaternion, and Umeyama scaling algorithms with per-point weights, batch dimensions, and custom autograd across NumPy, PyTorch, JAX, TensorFlow, and MLX.
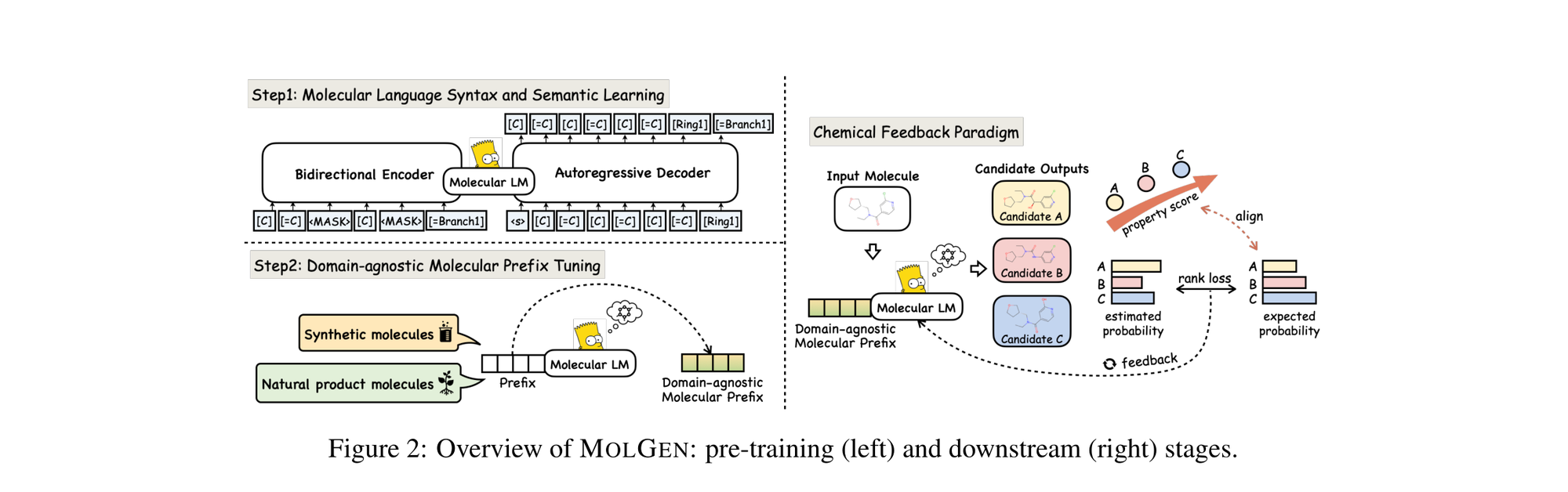
MolGen pre-trains on 100M+ SELFIES molecules, introduces domain-agnostic prefix tuning for cross-domain transfer, and applies a chemical feedback paradigm to reduce molecular hallucinations.
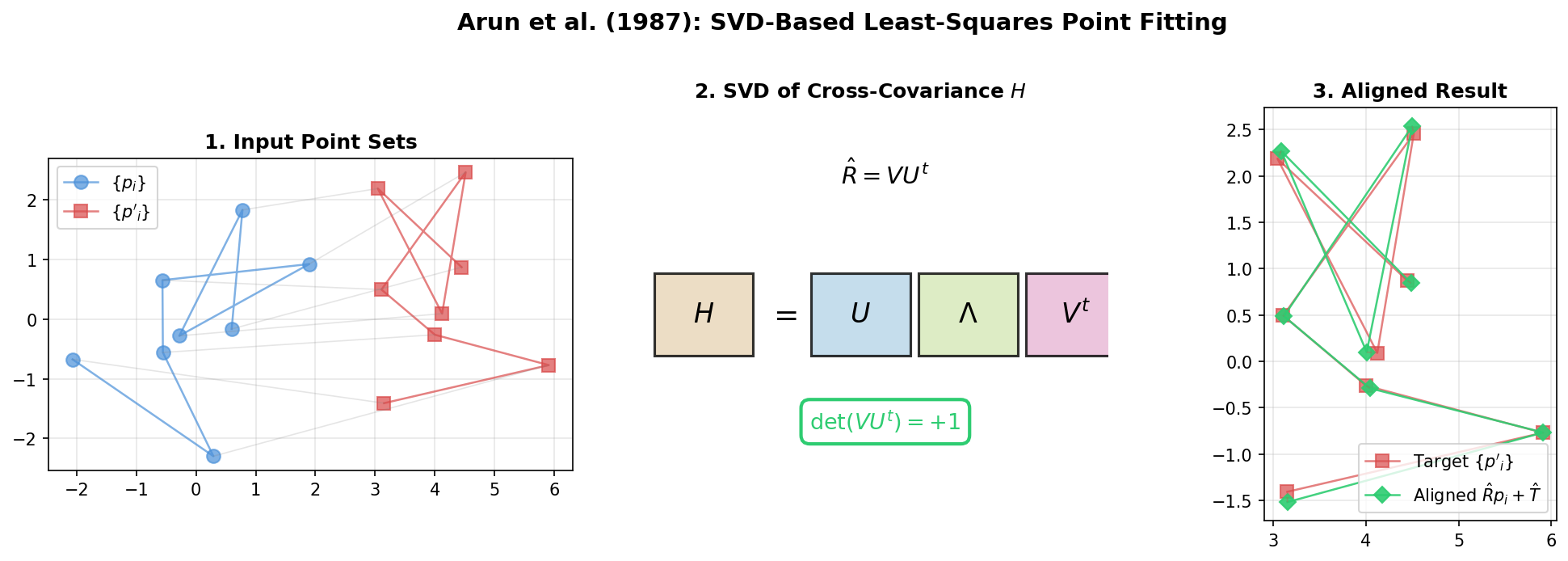
Presents a concise SVD-based algorithm for finding the optimal rotation and translation between two 3D point sets, with analysis of the degenerate reflection case that Umeyama later corrected.
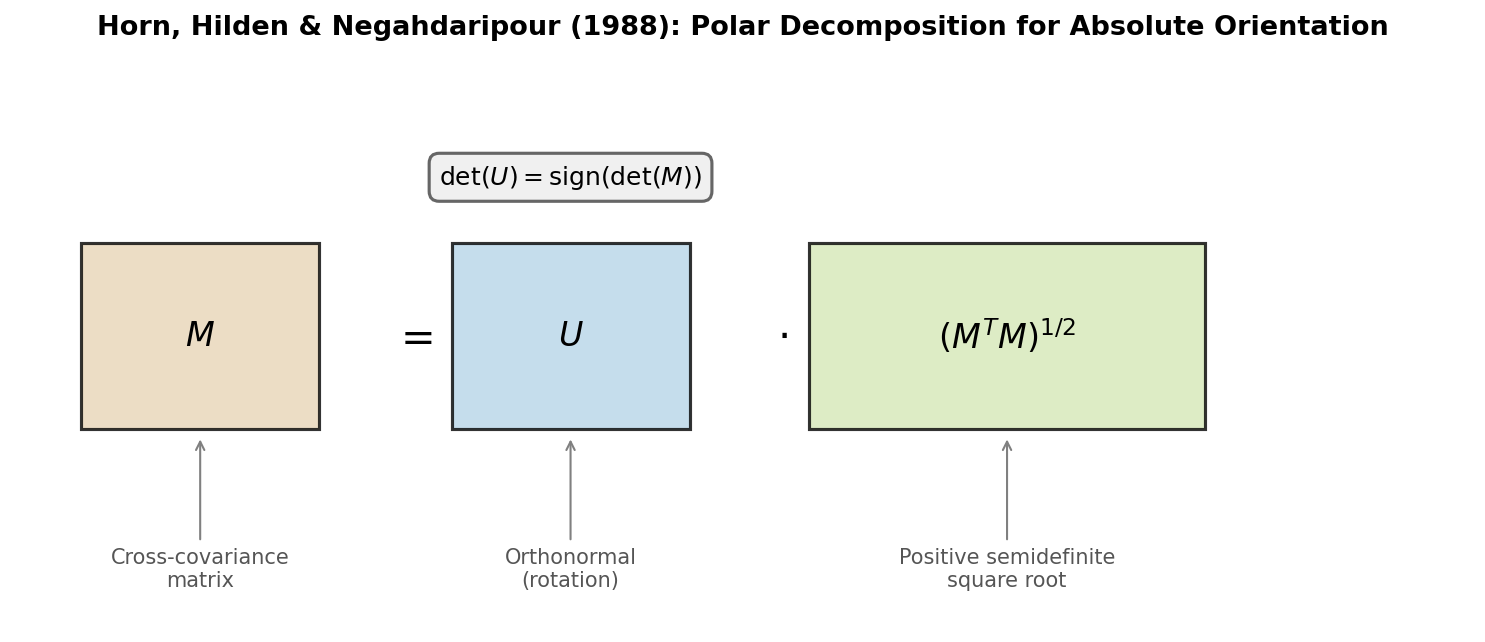
The matrix-based companion to Horn’s 1987 quaternion method, deriving the optimal rotation as the orthonormal factor in the polar decomposition of the cross-covariance matrix via eigendecomposition of a 3x3 symmetric matrix.
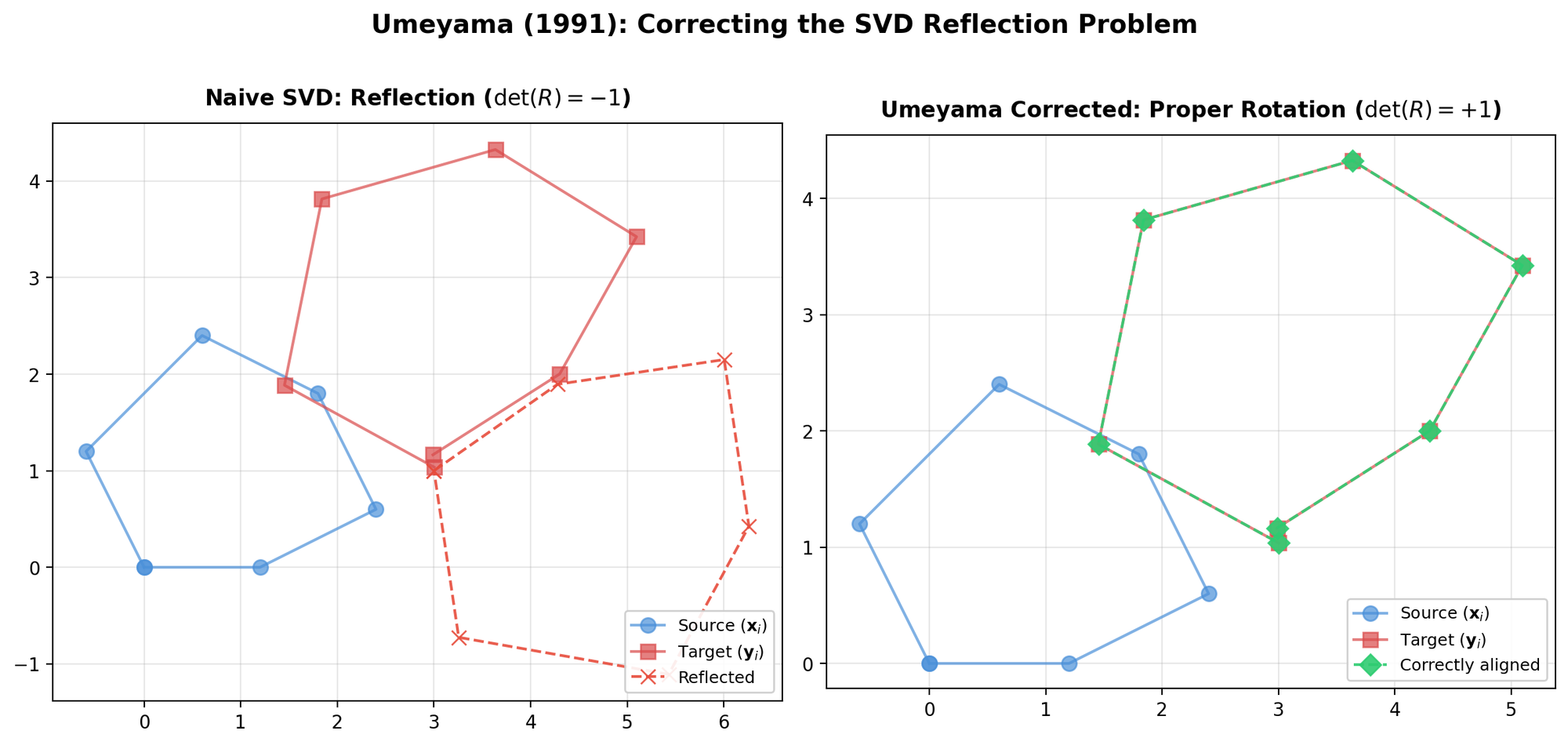
Corrects a flaw in prior SVD-based alignment methods (Arun et al., Horn et al.) that could produce reflections instead of rotations under noisy data, and provides a complete closed-form solution for similarity transformations in arbitrary dimensions.
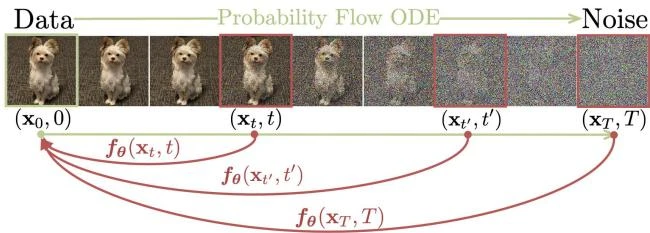
This paper introduces consistency models, a new family of generative models that map any point on a Probability Flow ODE trajectory to its origin. They support fast one-step generation by design, while allowing multi-step sampling for improved quality and zero-shot editing tasks like inpainting and colorization.
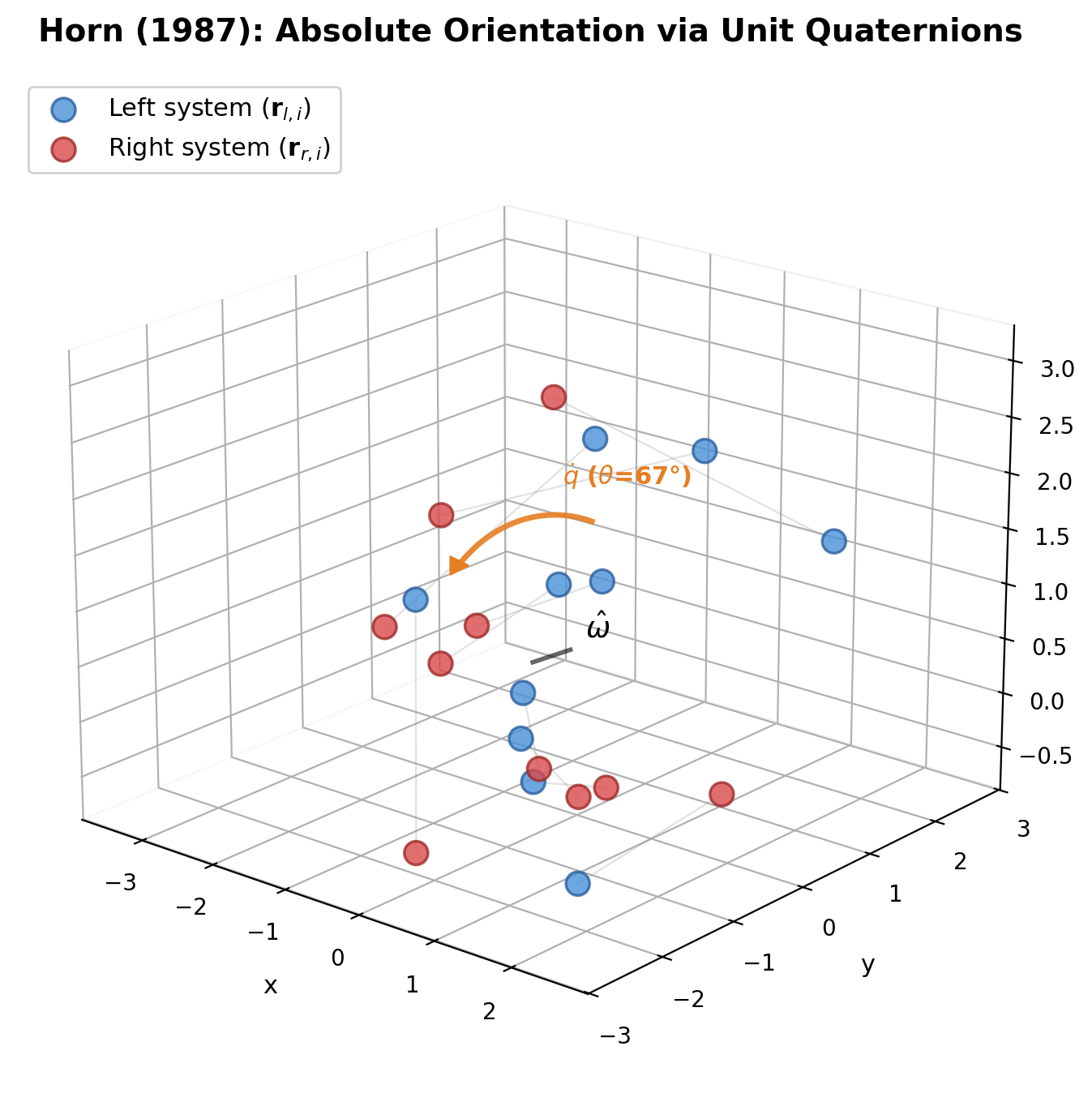
Derives the optimal rotation between two 3D point sets as the eigenvector of a 4x4 symmetric matrix built from cross-covariance sums, using unit quaternions to enforce the orthogonality constraint.
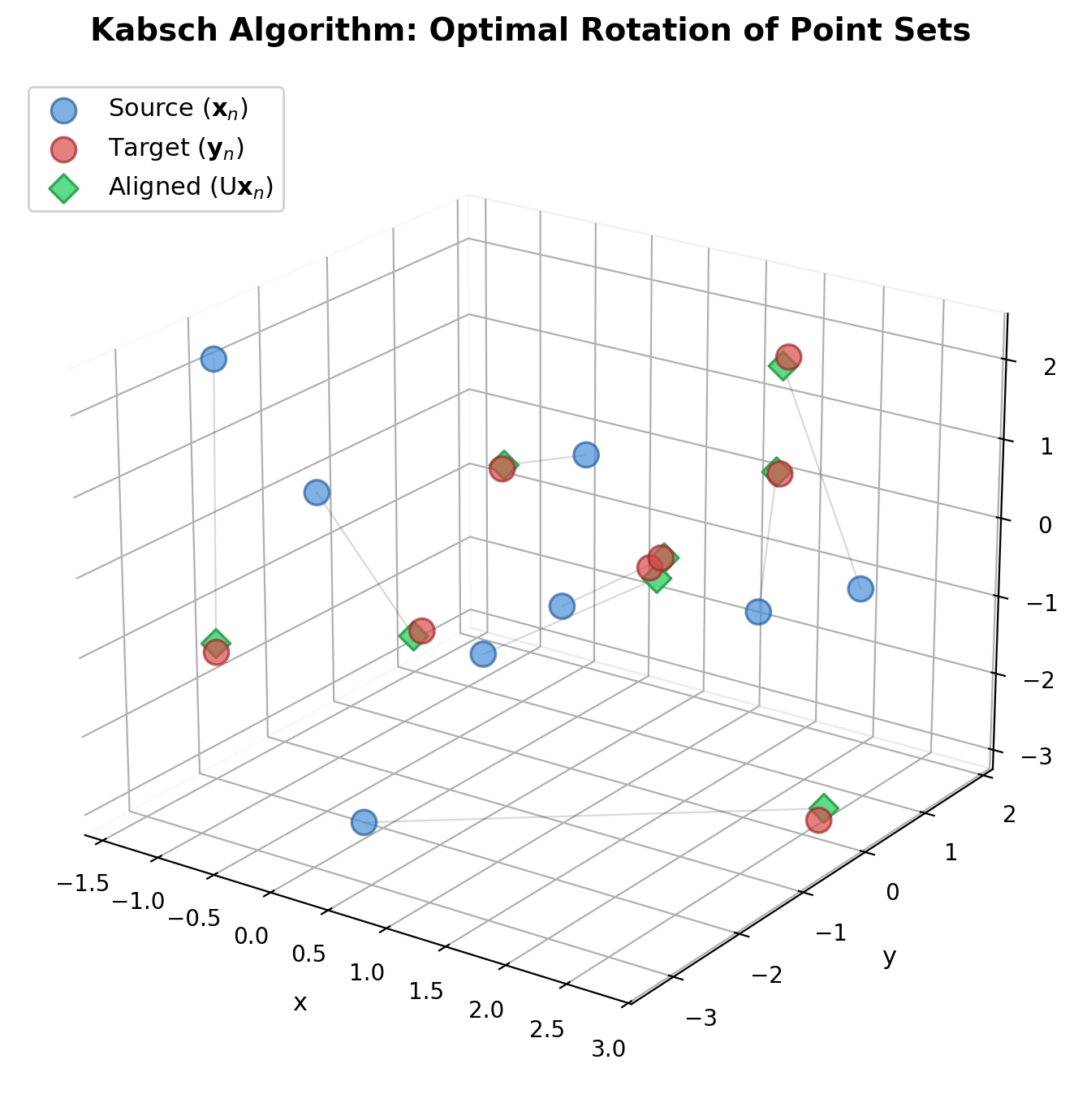
A foundational 1976 short communication presenting a direct, non-iterative method for finding the best rotation matrix between two point sets via eigendecomposition of a cross-covariance matrix.