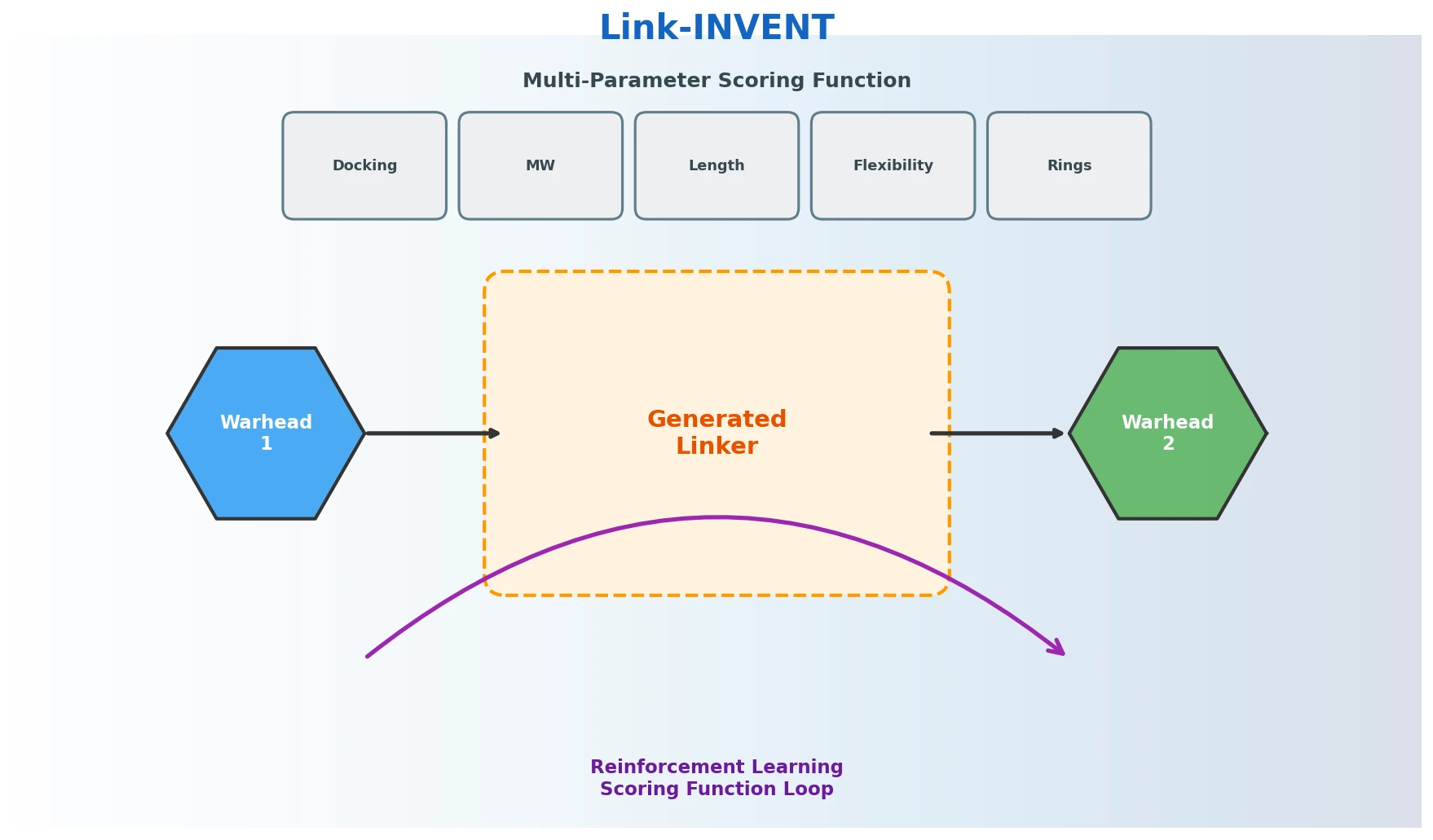
Link-INVENT: RL-Driven Molecular Linker Generation
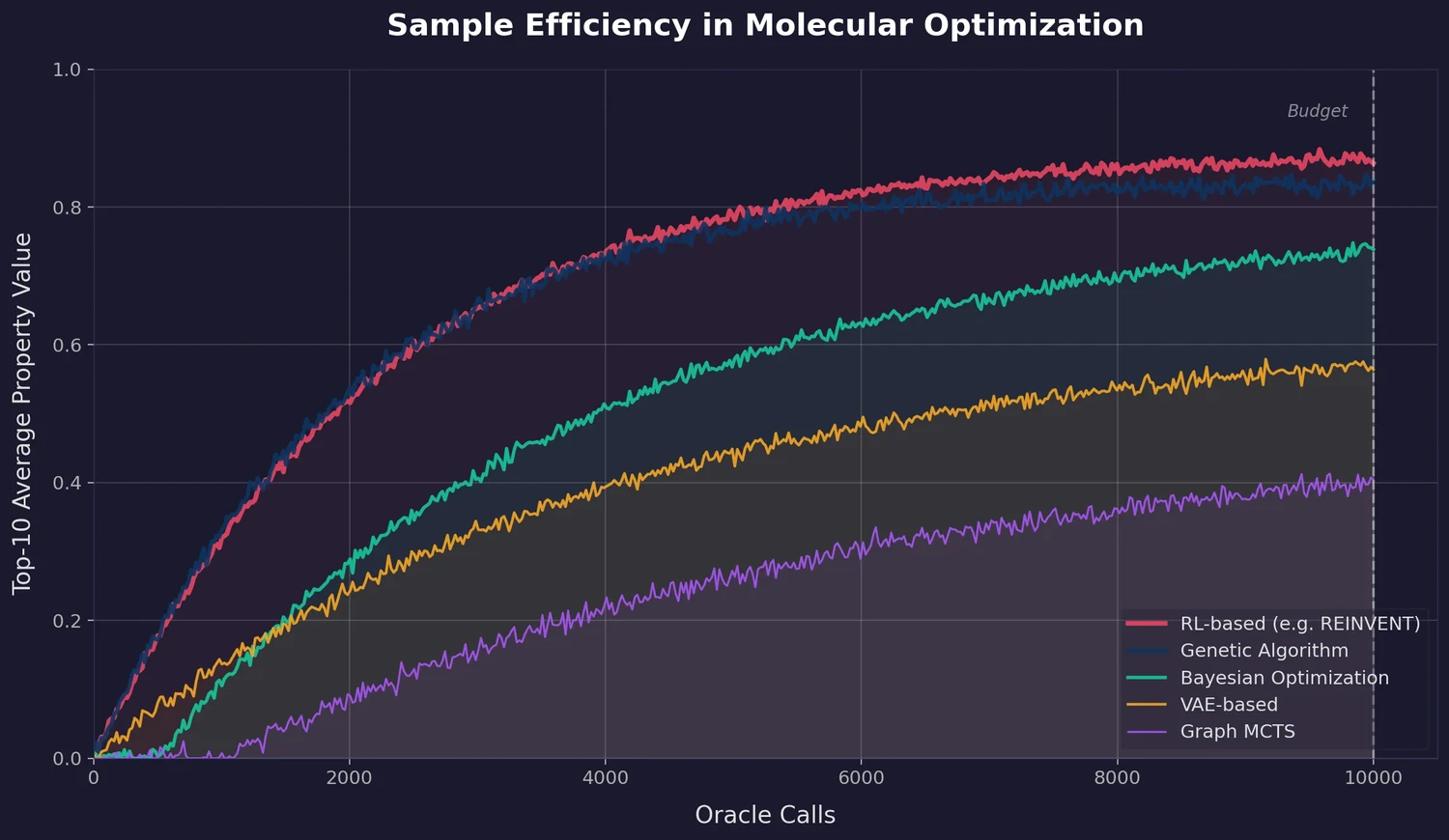
Link-INVENT is an RNN-based generative model for molecular linker design that uses reinforcement learning with a flexible scoring function, demonstrated on fragment linking, scaffold hopping, and PROTAC design.