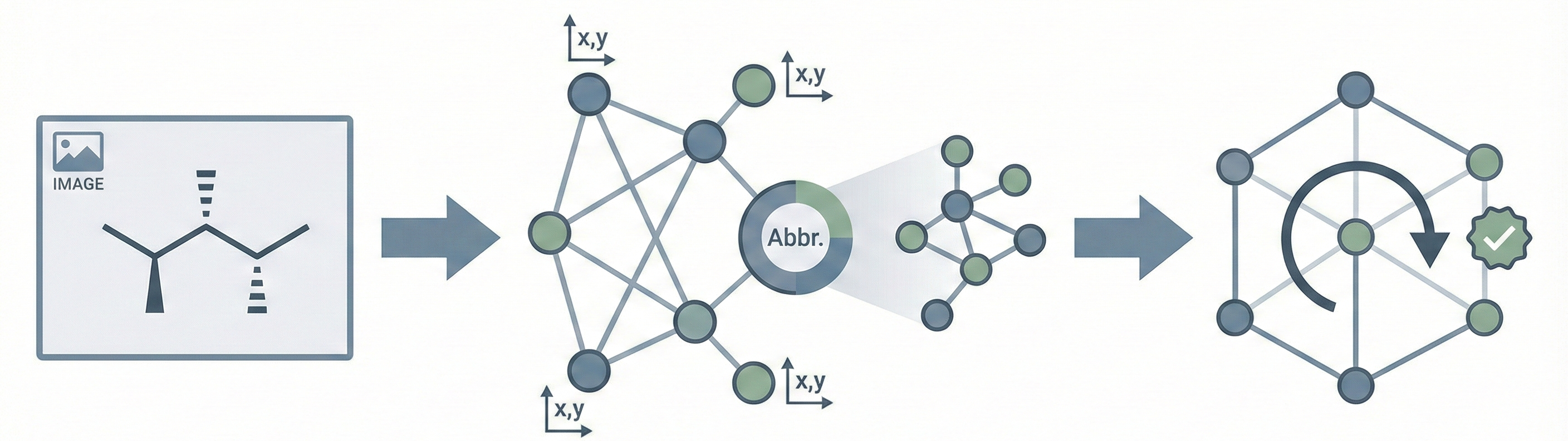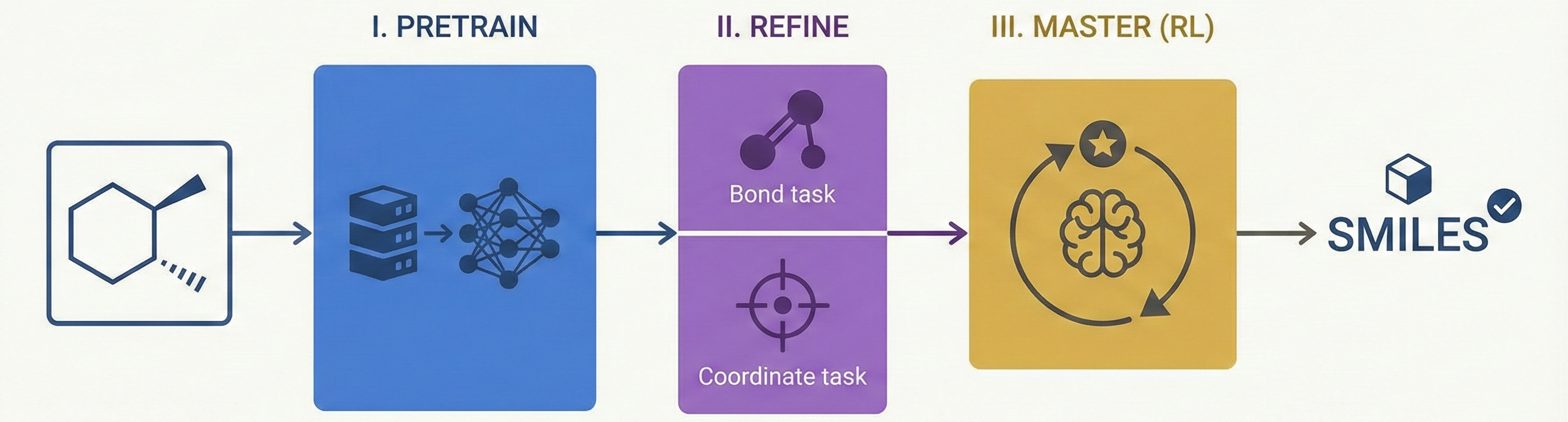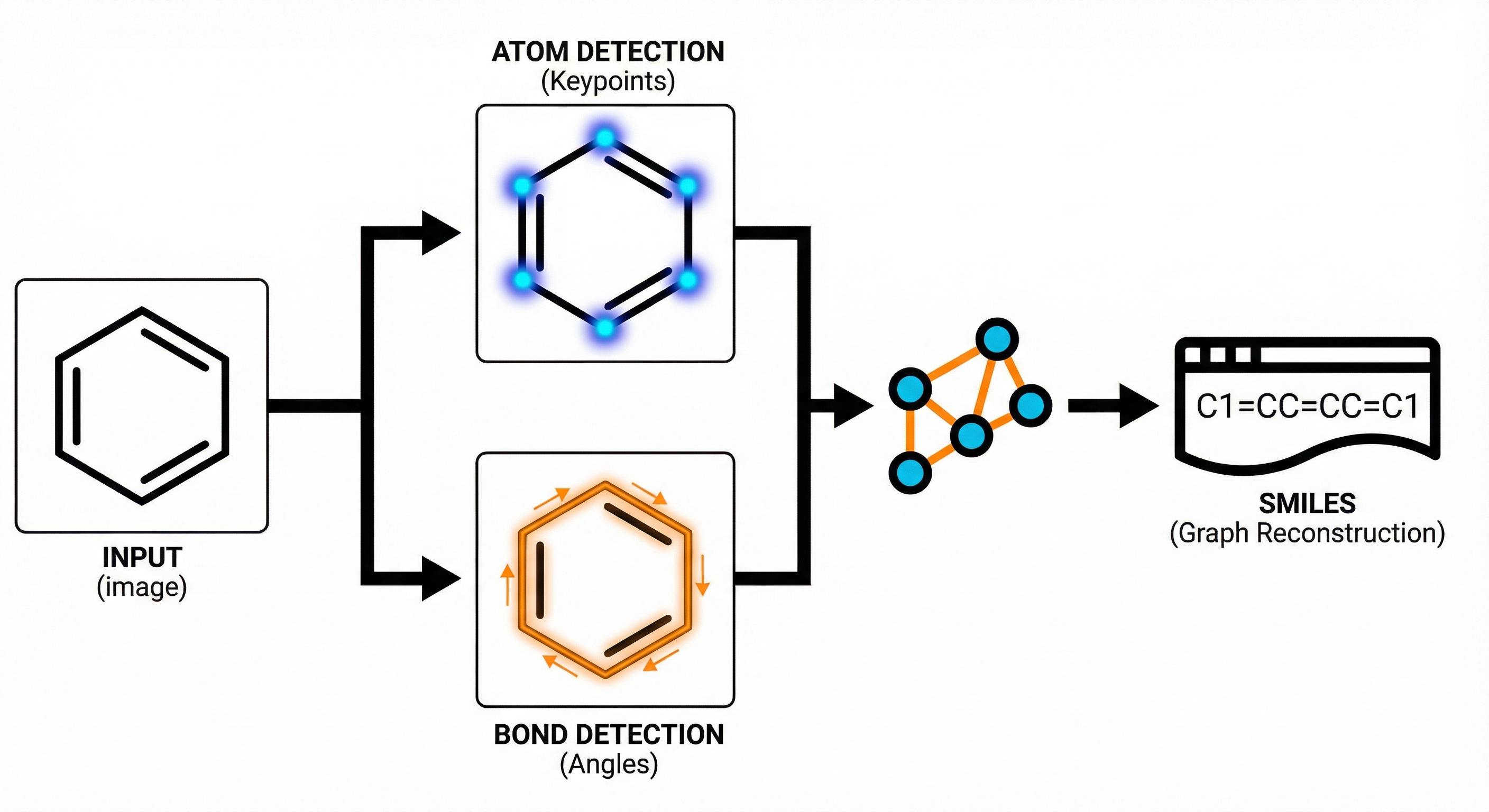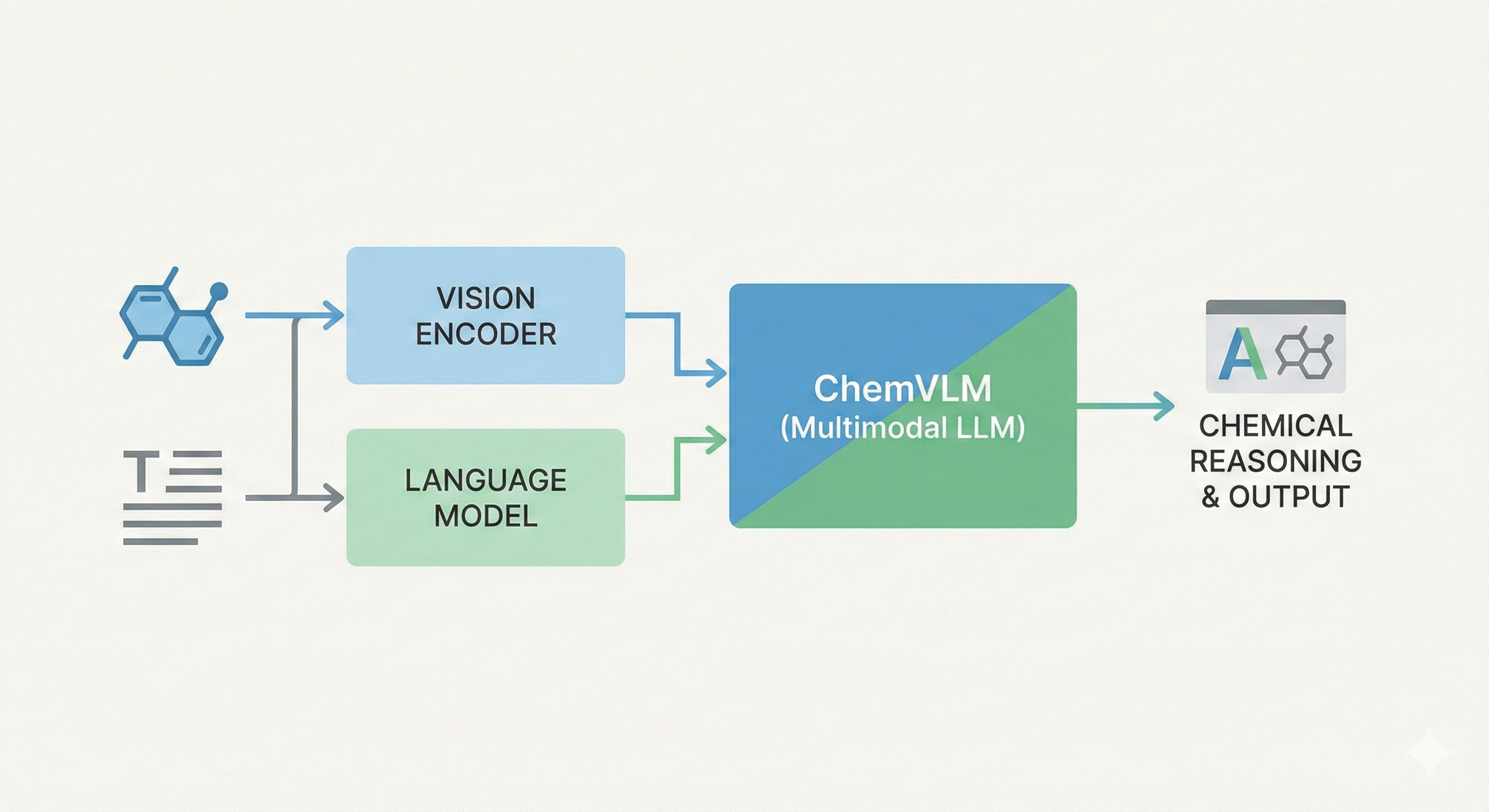
ChemVLM: A Multimodal Large Language Model for Chemistry
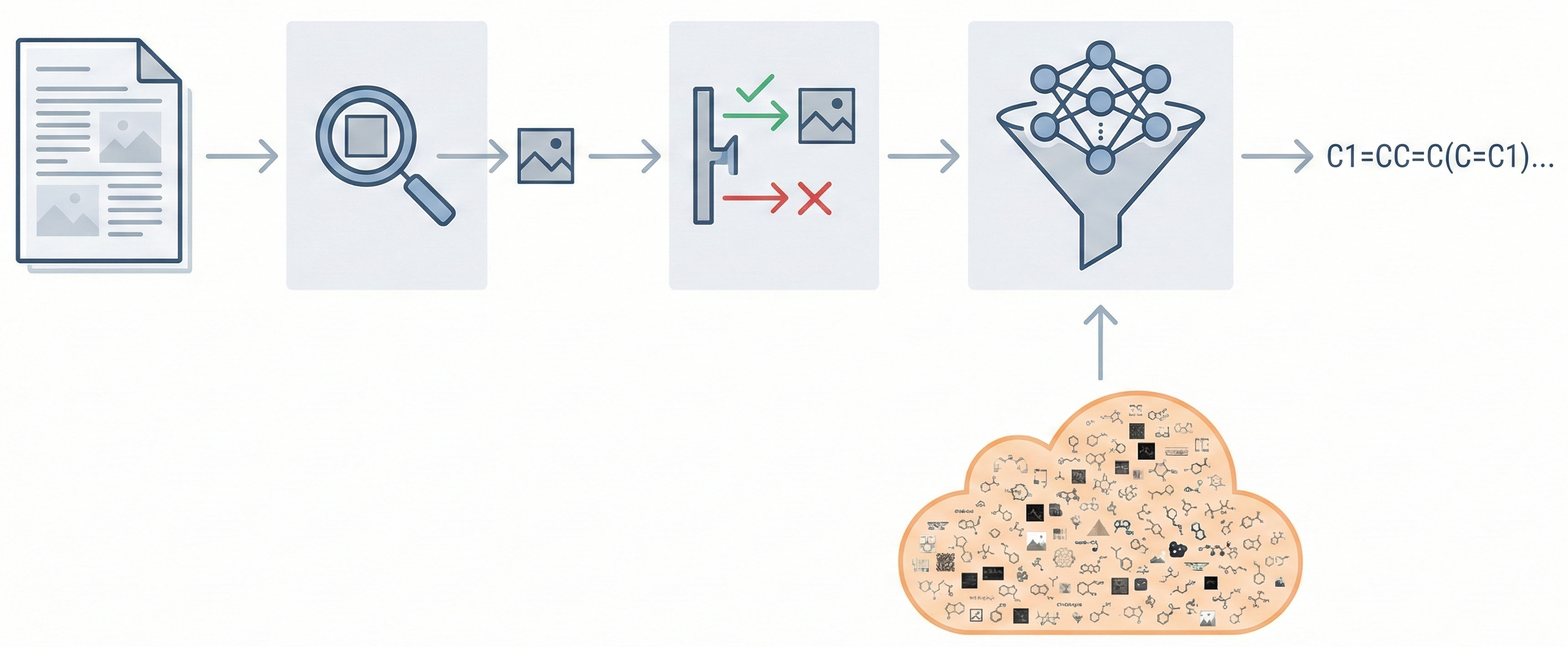
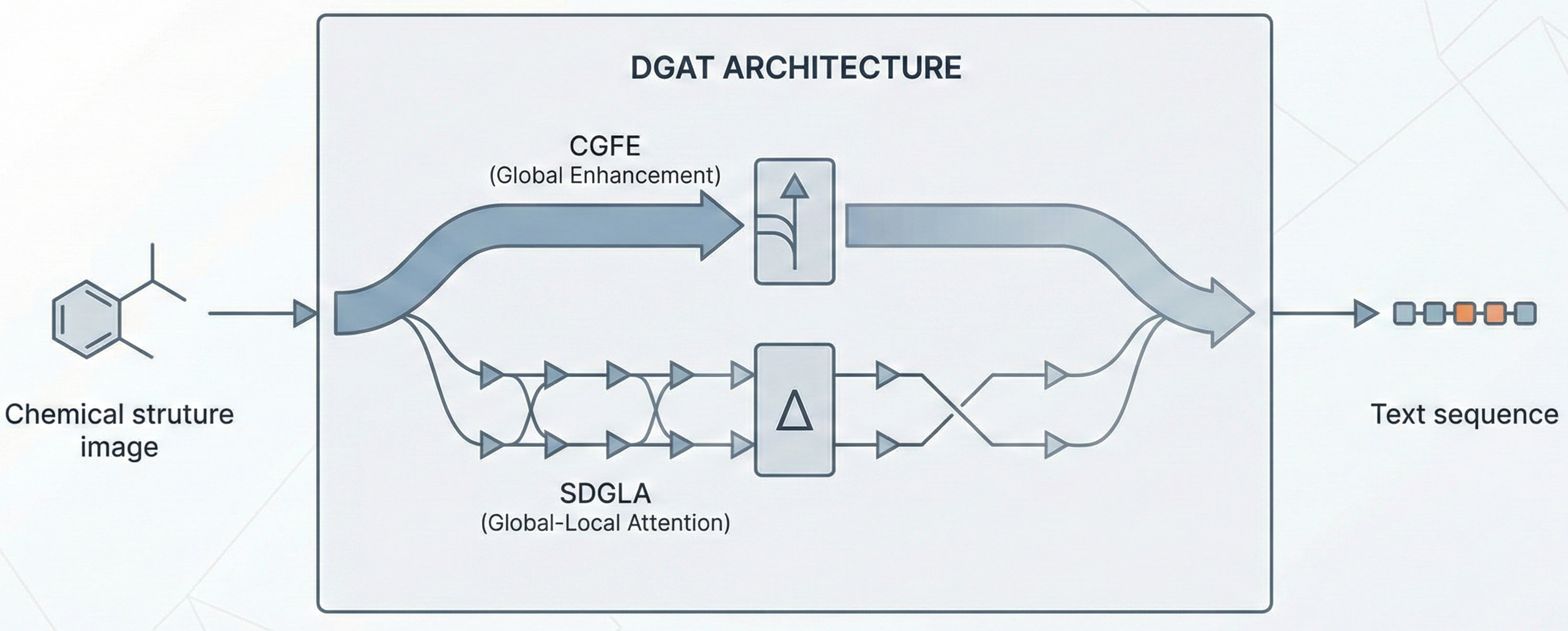
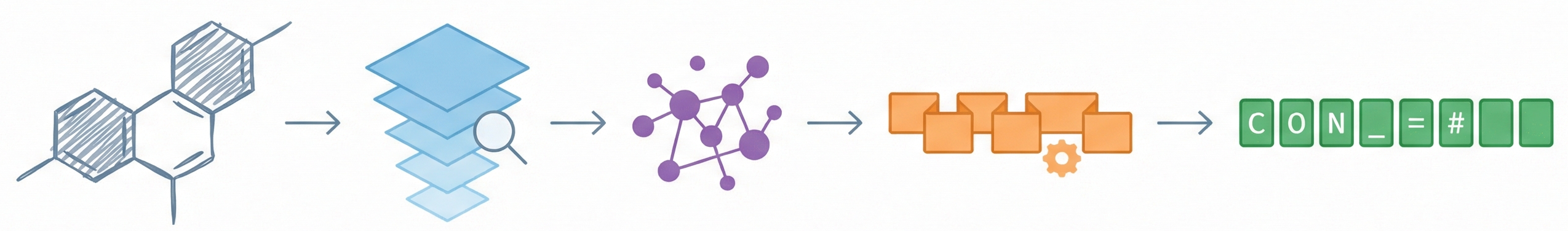
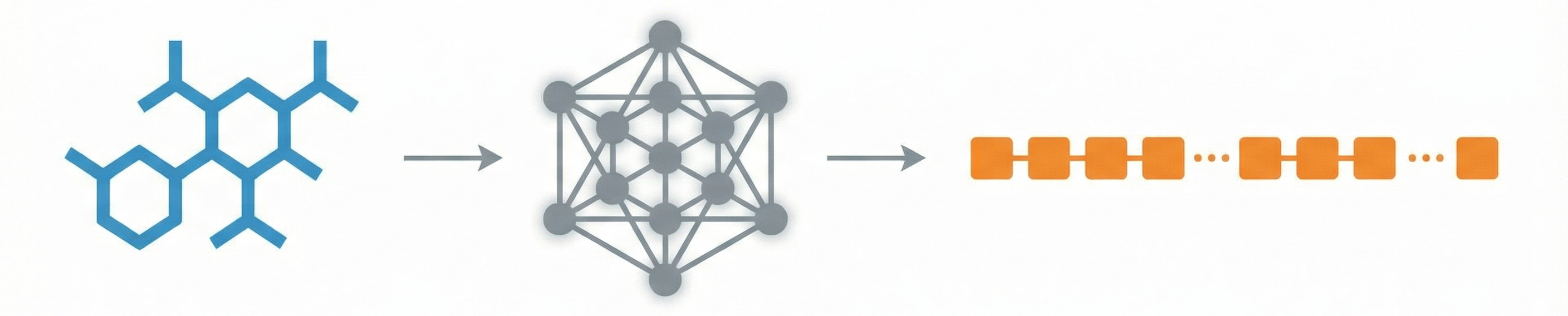
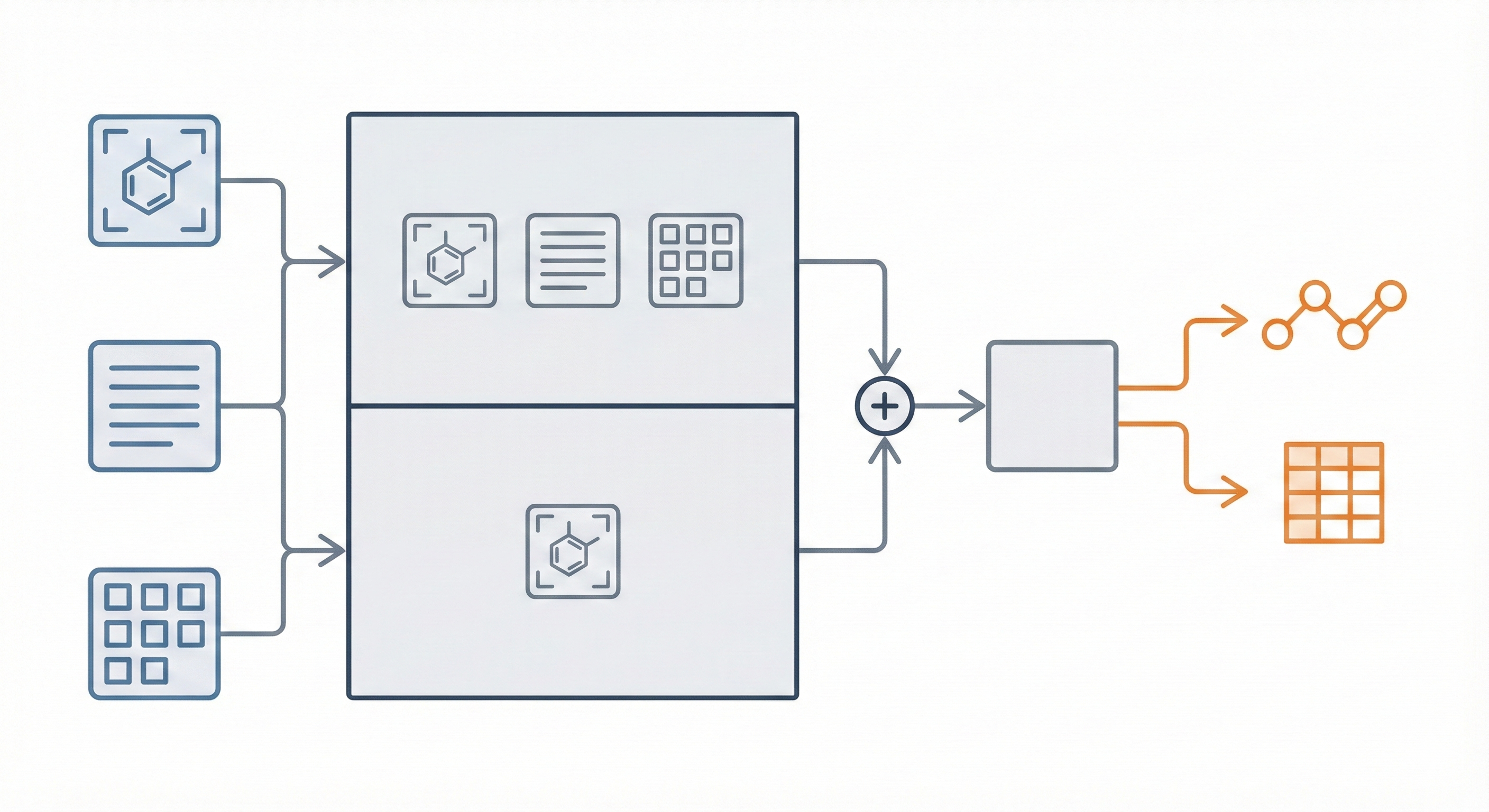
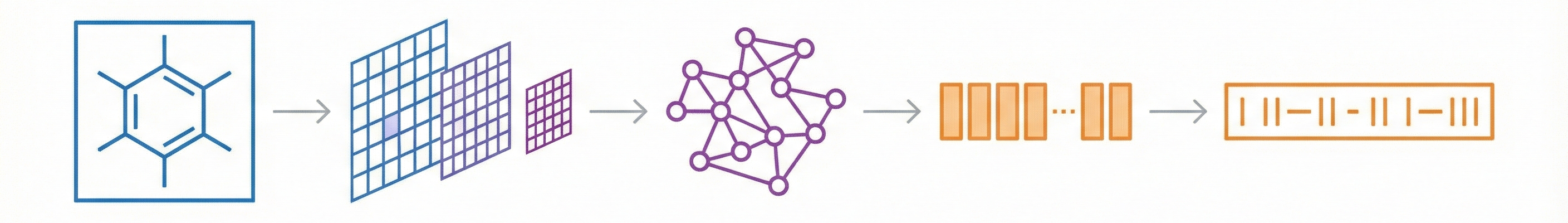
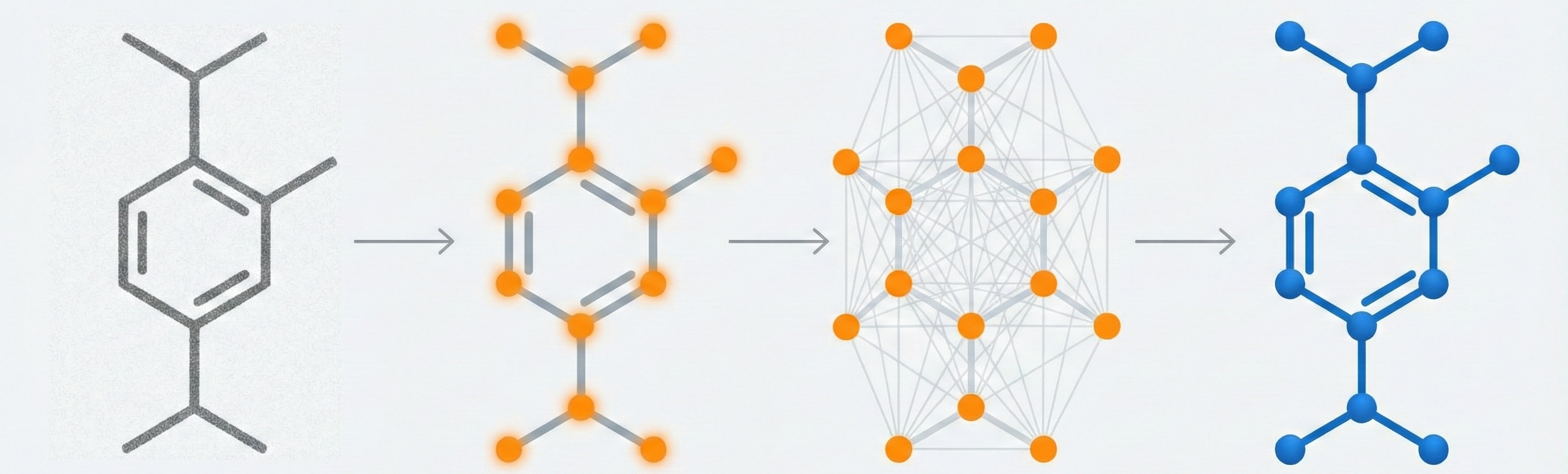
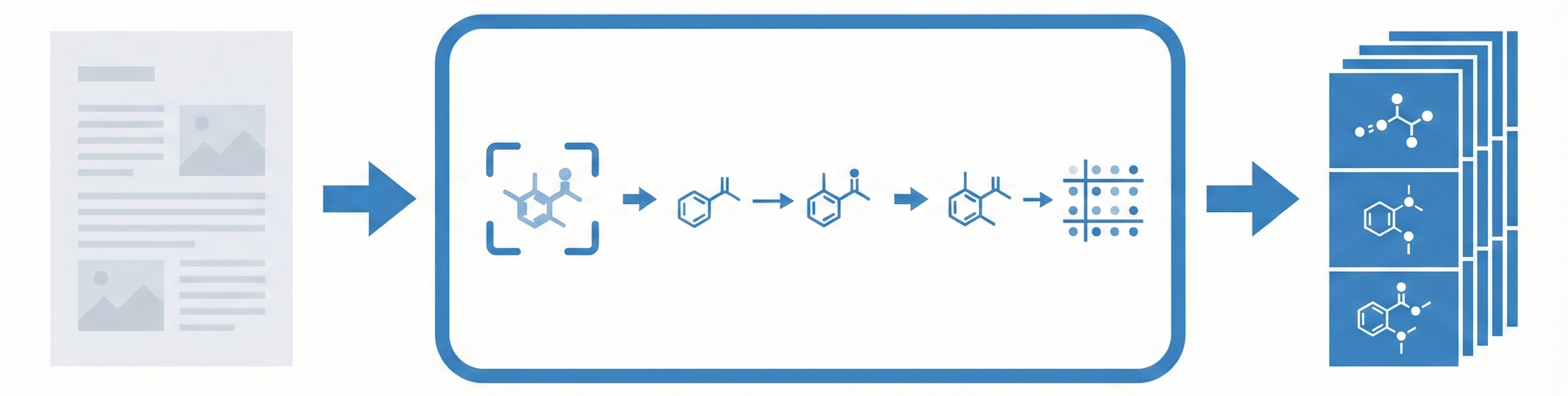
A 2025 AAAI paper introducing ChemVLM, a domain-specific multimodal LLM (26B parameters). It achieves state-of-the-art performance on chemical OCR, reasoning benchmarks, and molecular understanding tasks by combining vision and language models trained on curated chemistry data.