
Importance Weighted Autoencoders: Beyond the Standard VAE
Discover how Importance Weighted Autoencoders (IWAEs) use the same architecture as VAEs with a fundamentally more powerful objective to leverage multiple samples effectively.

Discover how Importance Weighted Autoencoders (IWAEs) use the same architecture as VAEs with a fundamentally more powerful objective to leverage multiple samples effectively.
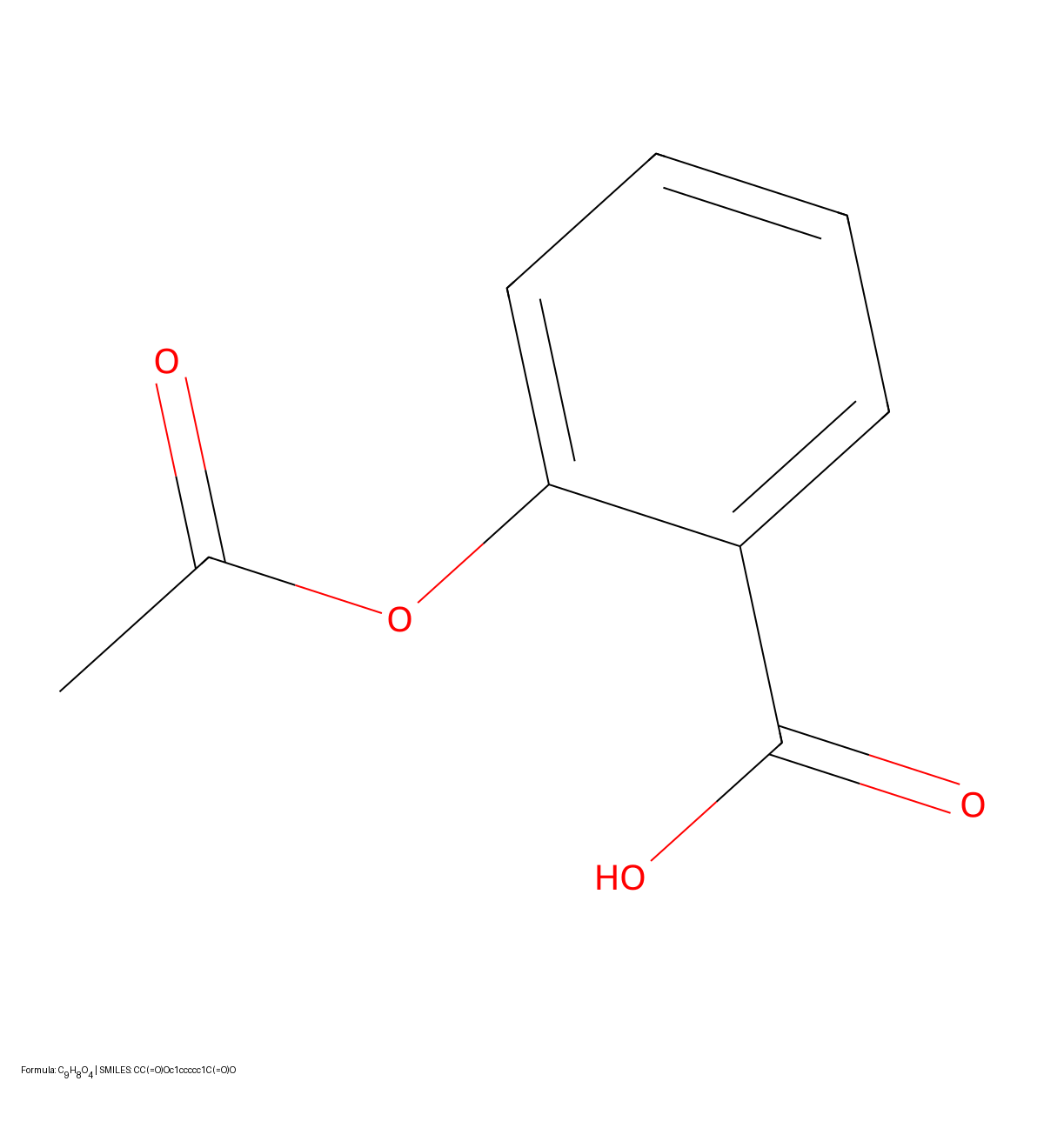
A 2024 deep learning system for optical chemical structure recognition designed specifically for biomedical literature mining, using ResNet-Transformer architecture to handle challenging conditions including low-resolution images, noise, distortions, and even hand-drawn molecular diagrams from scientific documents.
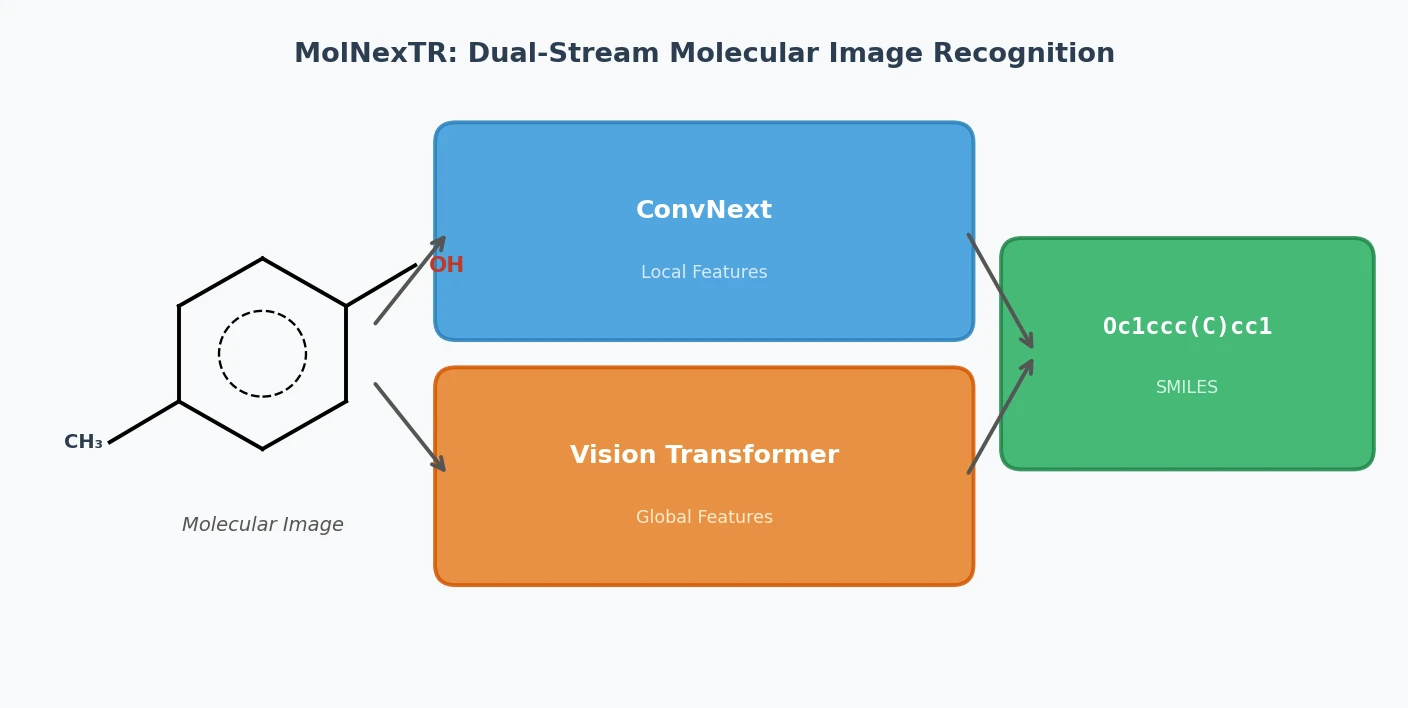
MolNexTR proposes a dual-stream architecture combining ConvNext and Vision Transformers to improve molecular image recognition (OCSR). It achieves 81-97% accuracy across diverse benchmarks utilizing simultaneous local and global feature extraction alongside specialized image contamination augmentations.
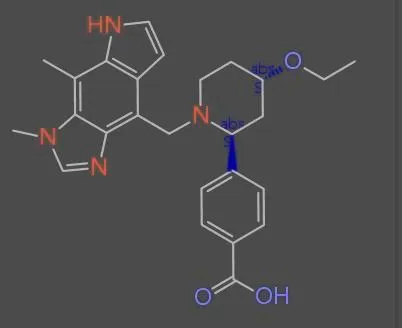
The MolParser project introduces two key datasets: MolParser-7M, the largest training dataset for Optical Chemical Structure Recognition (OCSR) with 7.7M pairs of images and E-SMILES strings, and WildMol, a new 20k-sample benchmark for evaluating models on challenging real-world data. The training data uniquely combines millions of diverse synthetic molecules with 400,000 manually annotated in-the-wild samples.
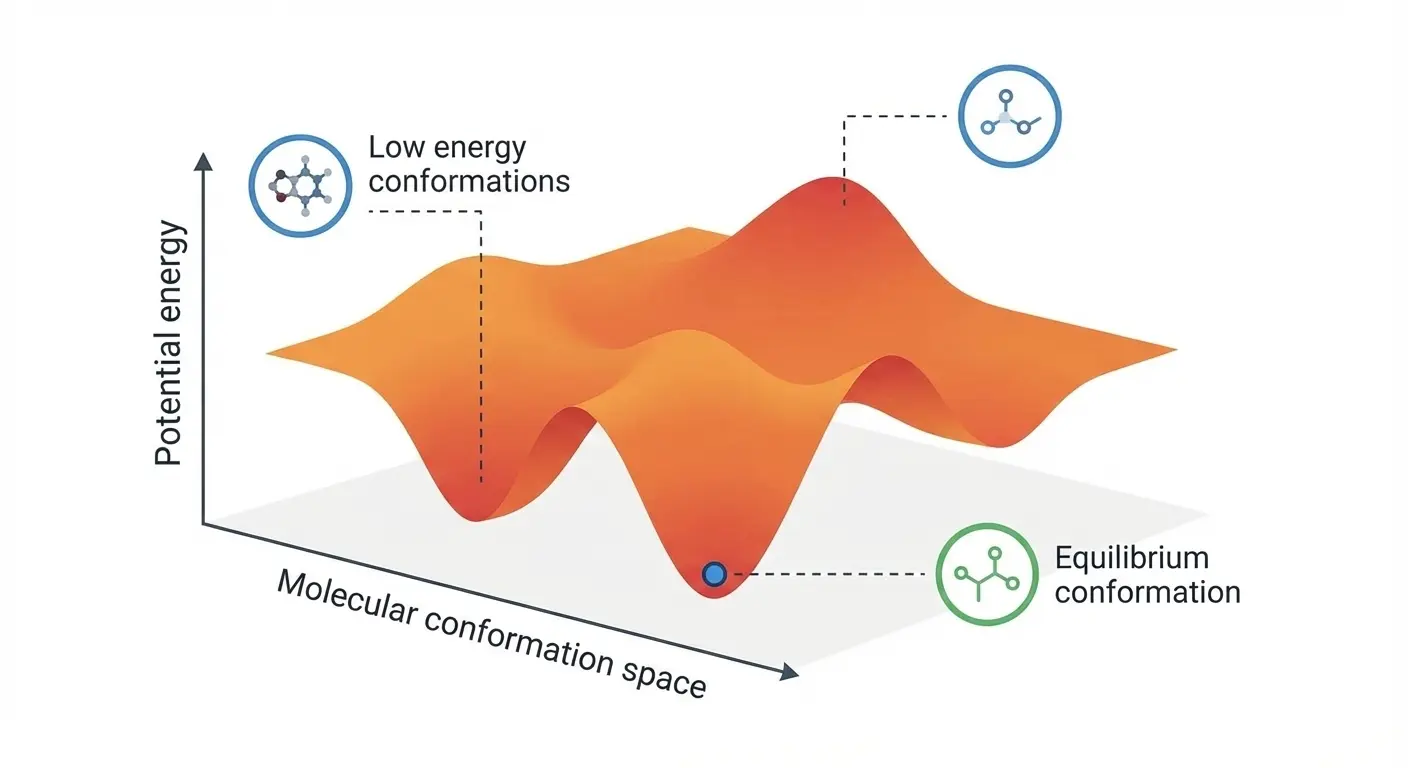
ICLR 2025 paper introducing DenoiseVAE, which learns adaptive, atom-specific noise distributions through a VAE framework to improve denoising-based pre-training for molecular force field prediction, outperforming fixed Gaussian noise approaches on quantum chemistry benchmarks.
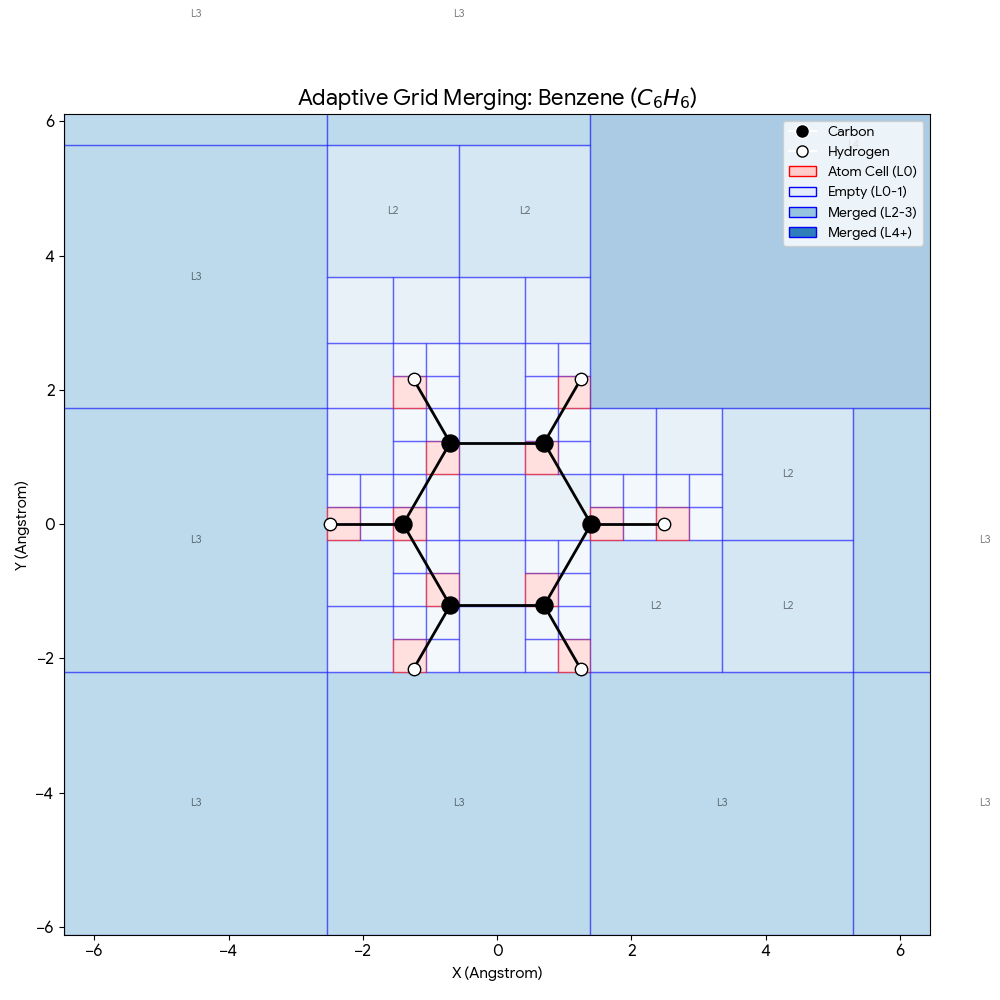
ICML 2025 paper introducing SpaceFormer, a Transformer architecture that challenges the atom-centric paradigm by modeling the continuous 3D space surrounding molecules using adaptive multi-resolution grids, ranking first in 10 of 15 molecular property prediction tasks.

ICML 2025 analysis rigorously quantifying when non-conservative force models (which predict forces directly) fail in molecular dynamics, demonstrating simulation instabilities and proposing hybrid architectures that capture speed benefits without sacrificing physical correctness.
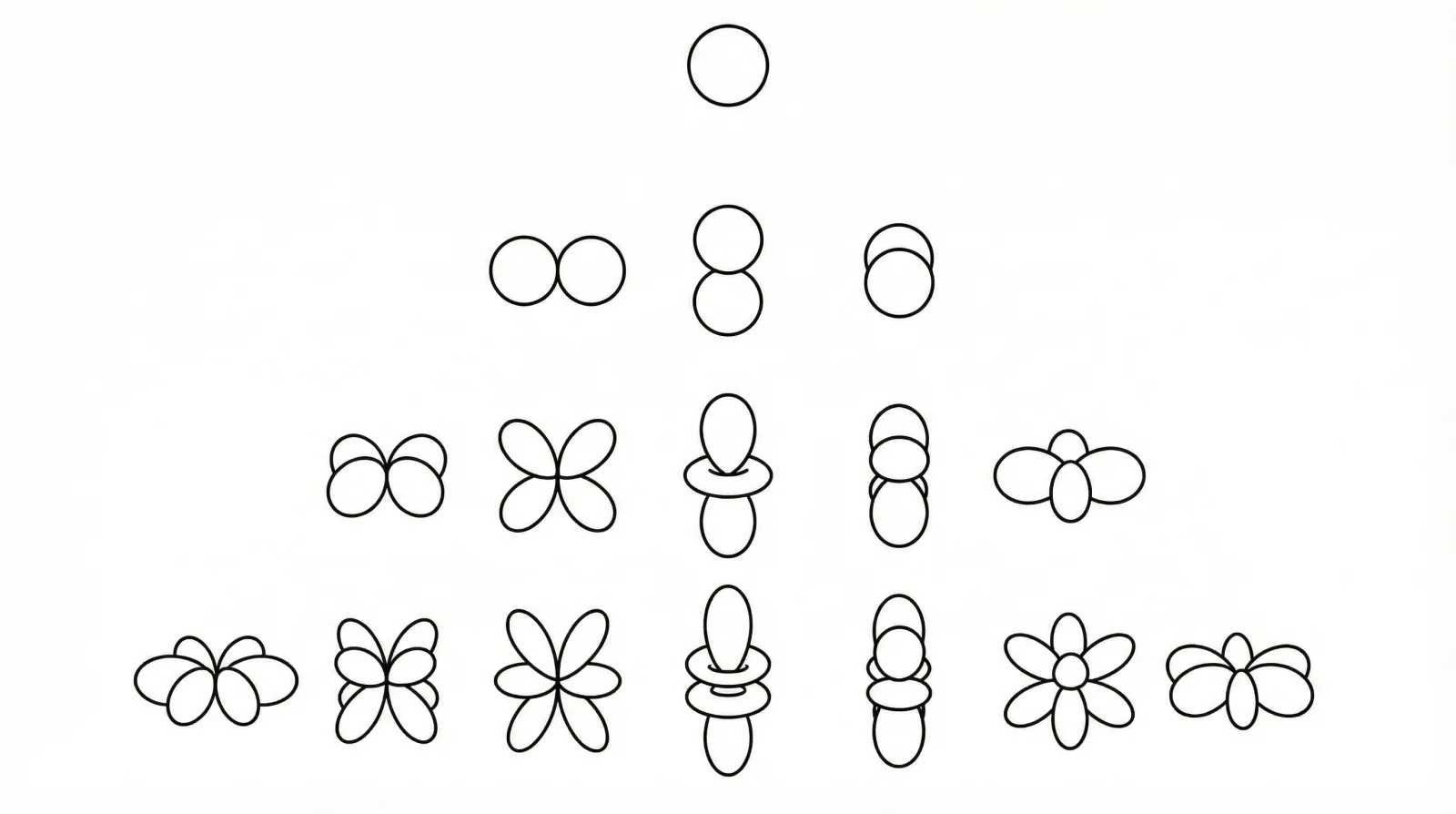
ICML 2025 methodological paper introducing SPHNet, which uses adaptive network sparsification to overcome the computational bottleneck of tensor products in SE(3)-equivariant networks, achieving up to 7x speedup and 75% memory reduction on DFT Hamiltonian prediction tasks.
ICML 2025 paper proposing energy conservation metrics as critical diagnostics for machine learning interatomic potentials and introducing eSEN, a novel architecture designed to bridge the gap between test-set accuracy and real simulation performance on materials property prediction.
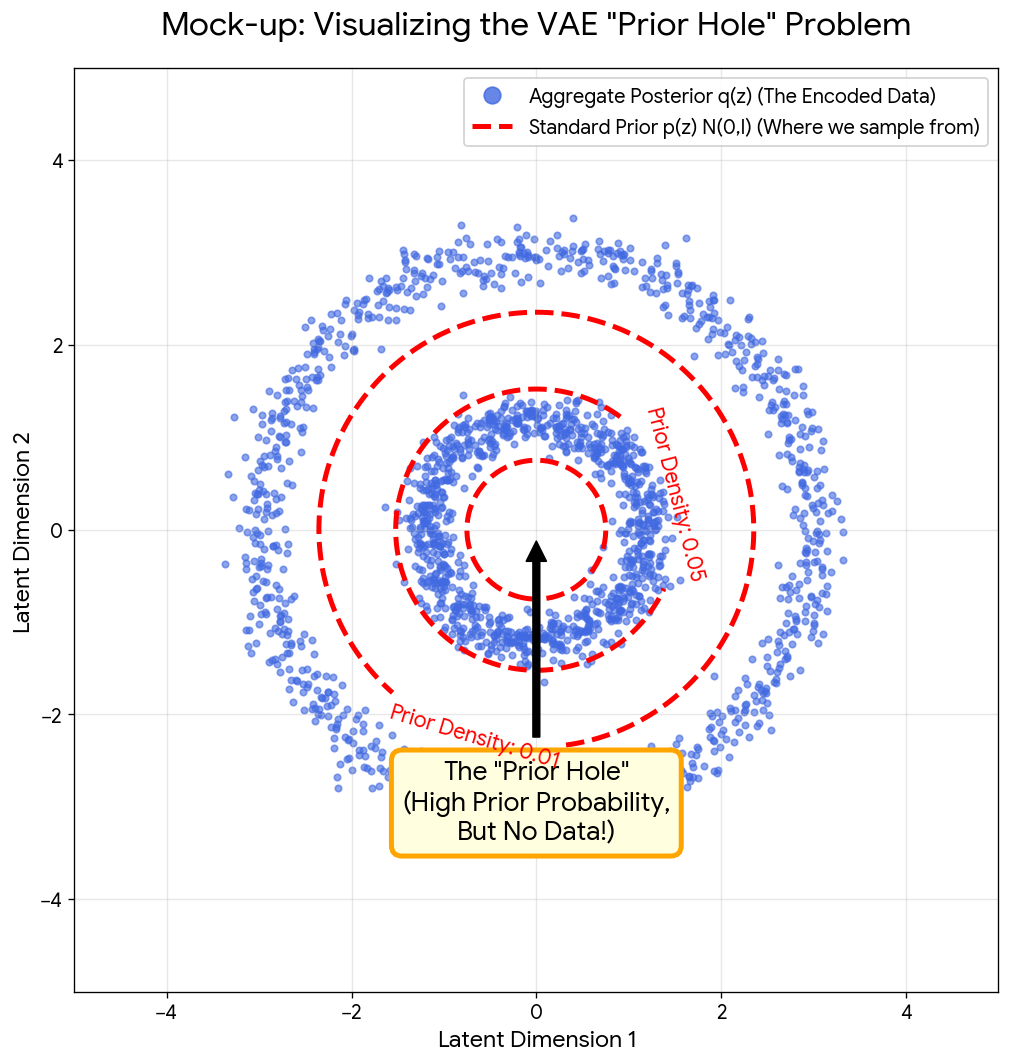
A NeurIPS 2021 method paper introducing Noise Contrastive Priors to address the VAE ‘prior hole’ problem, where standard Gaussian priors assign high density to regions of latent space that don’t correspond to realistic data, using energy-based models trained with contrastive learning to match the aggregate posterior.
A 2025 Vision-Language Model for OCSR that uses graph traversal chain-of-thought reasoning and a two-stage SFT plus GRPO training scheme to handle both printed molecules (including chemical abbreviations like Ph and Et) and hand-drawn structures, achieving strong performance on the new MolRec-Bench benchmark.
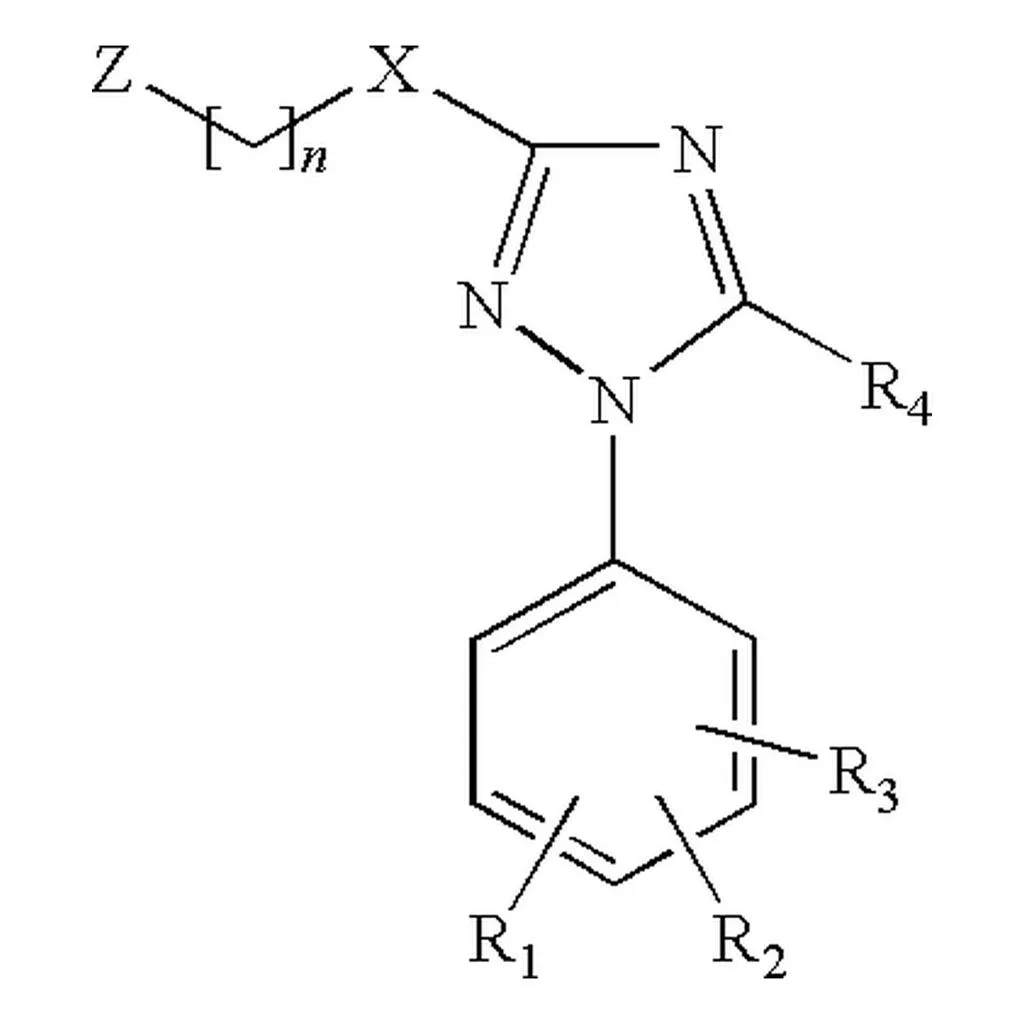
SubGrapher introduces a visual fingerprinting approach to Optical Chemical Structure Recognition that detects functional groups directly from images, enabling chemical database searches without full structure reconstruction and handling complex patent images including Markush structures.