PASITHEA: Gradient-Based Molecular Design via Dreaming
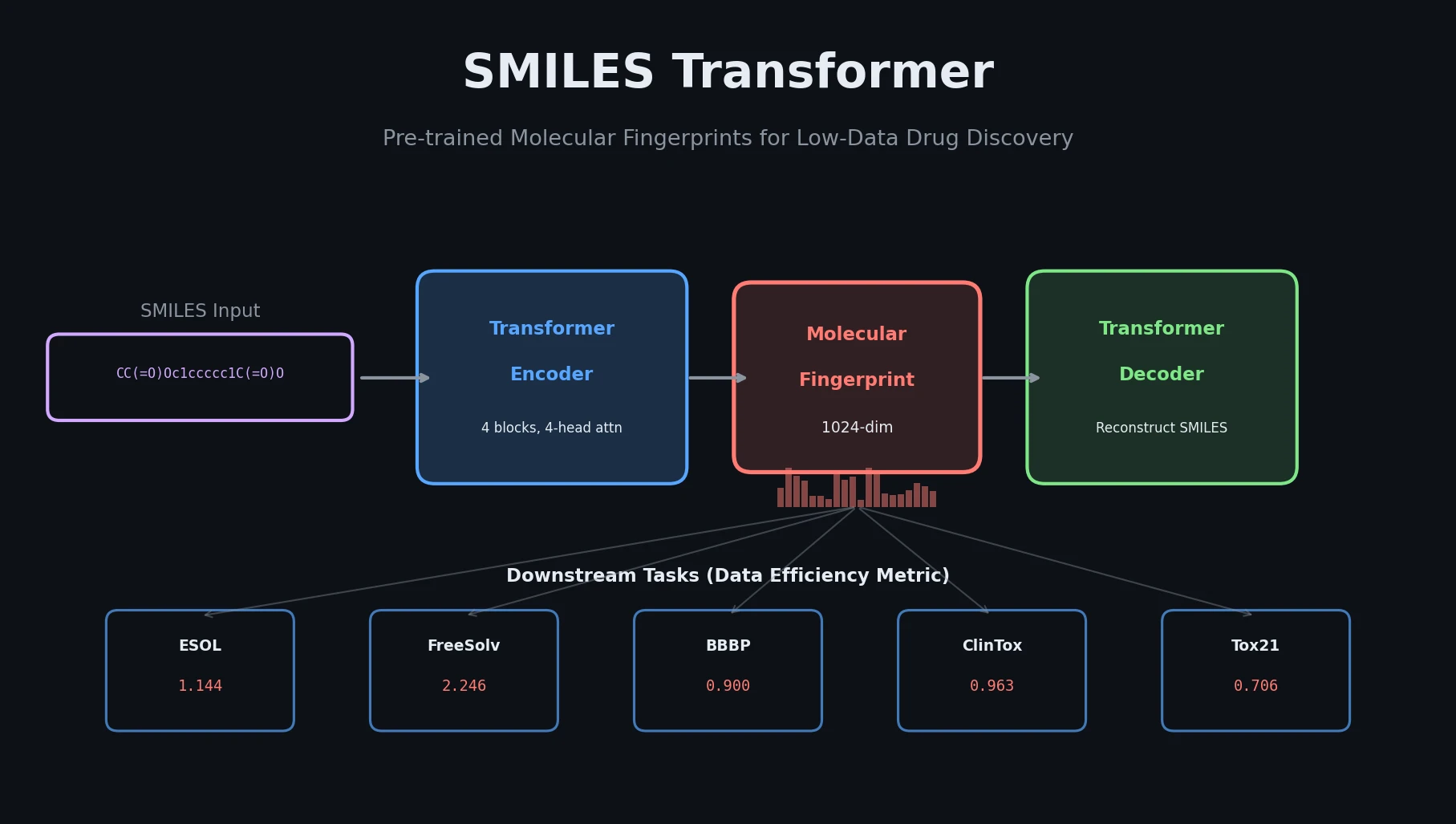
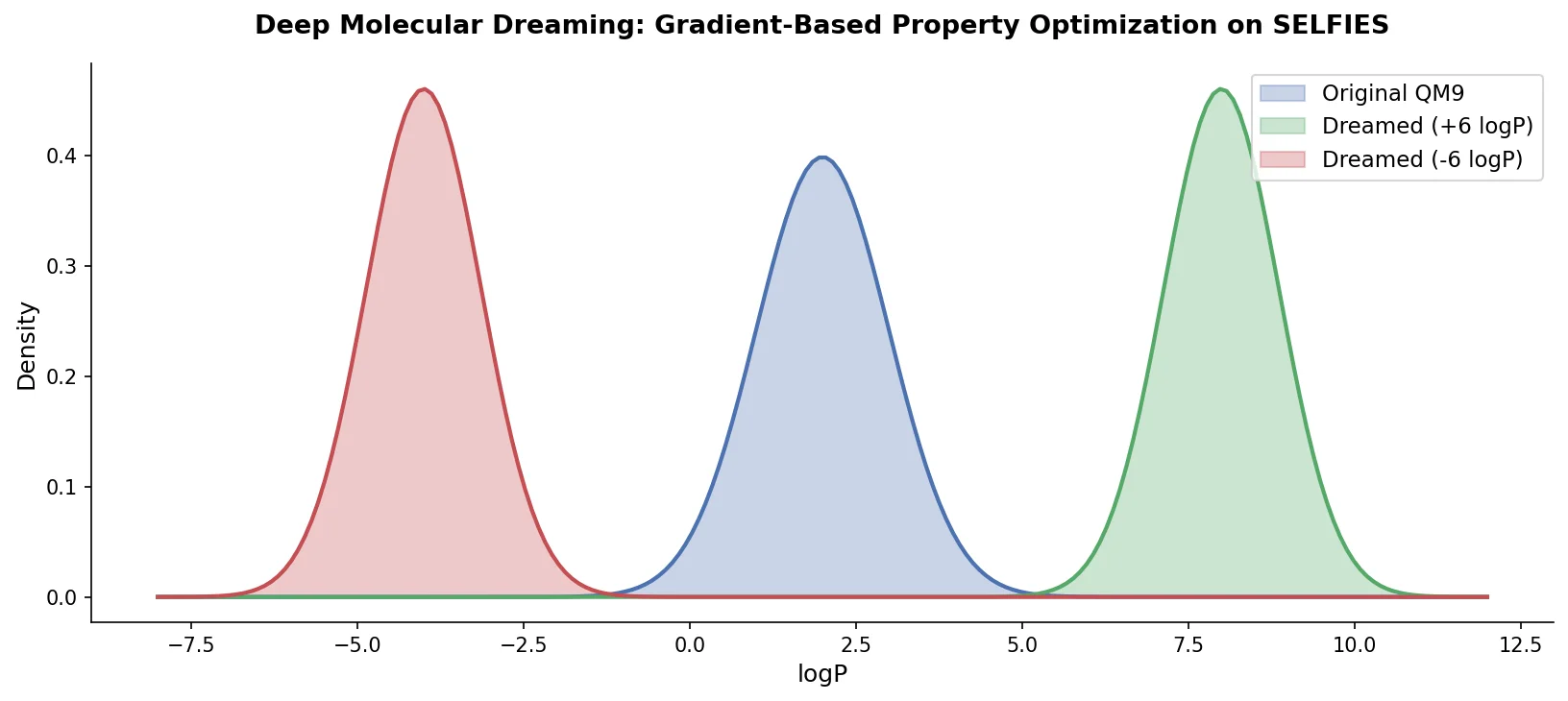
PASITHEA adapts deep dreaming from computer vision to molecular design, directly optimizing SELFIES-encoded molecules for target chemical properties via gradient-based inversion of a trained regression network.