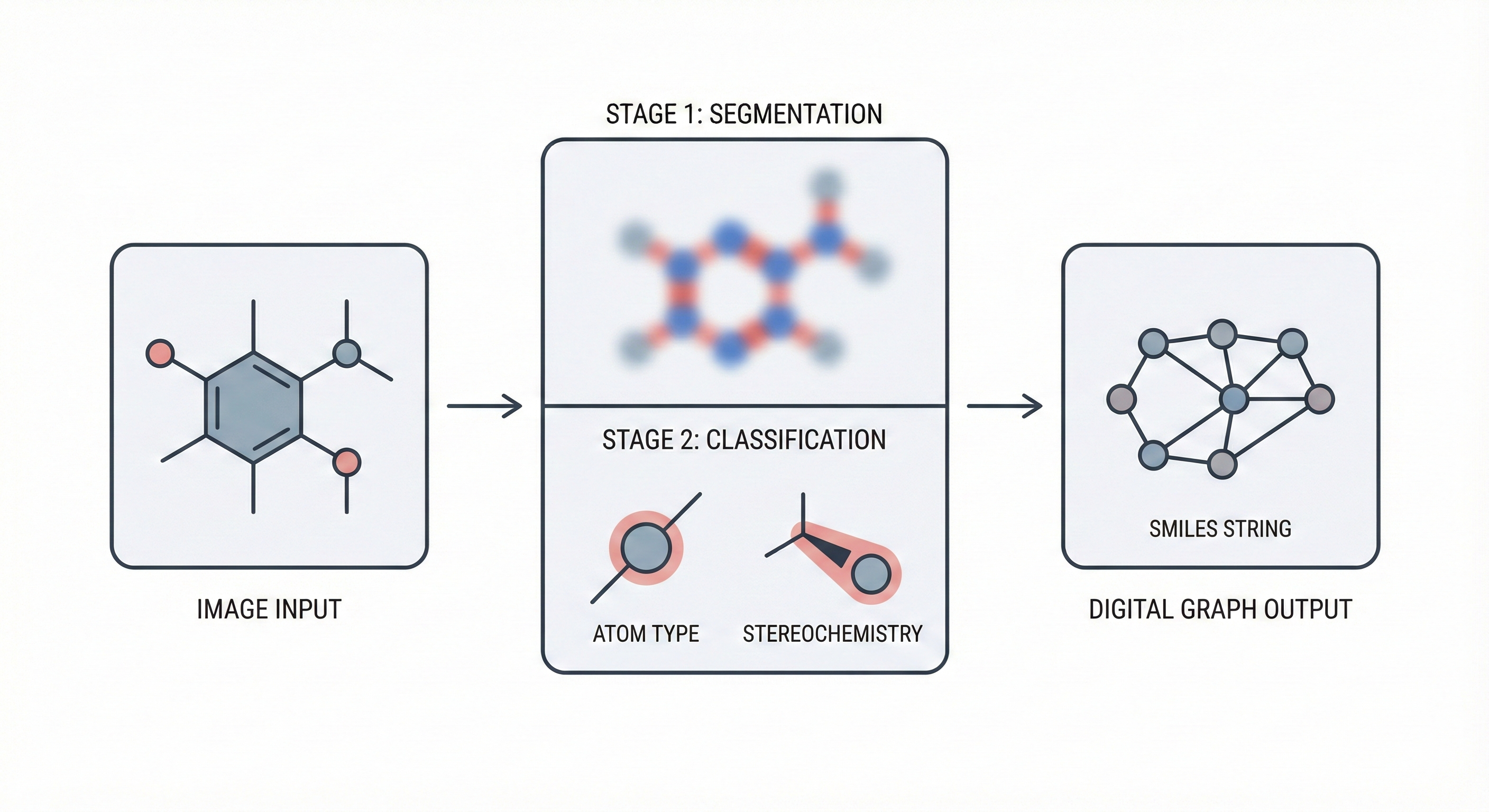
DECIMER 1.0: Transformers for Chemical Image Recognition
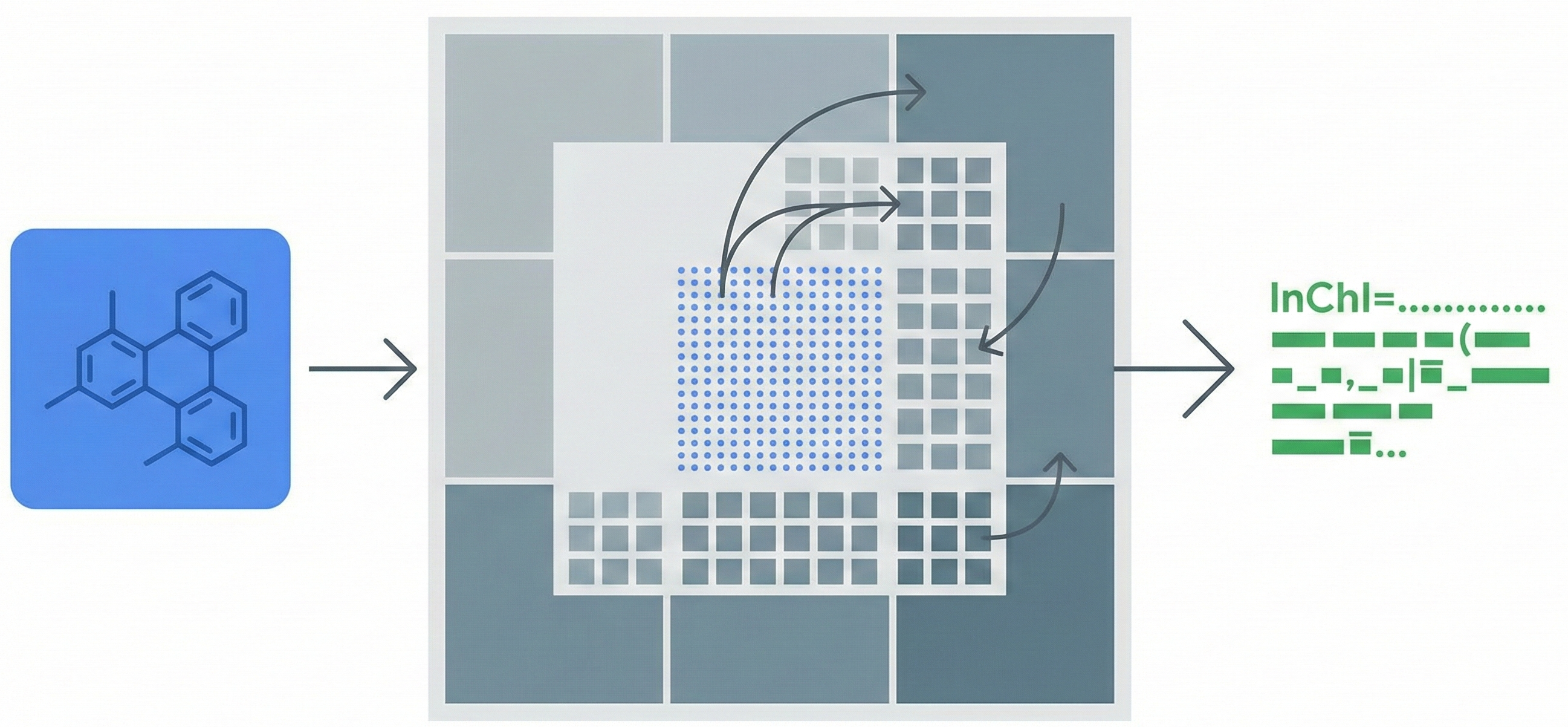
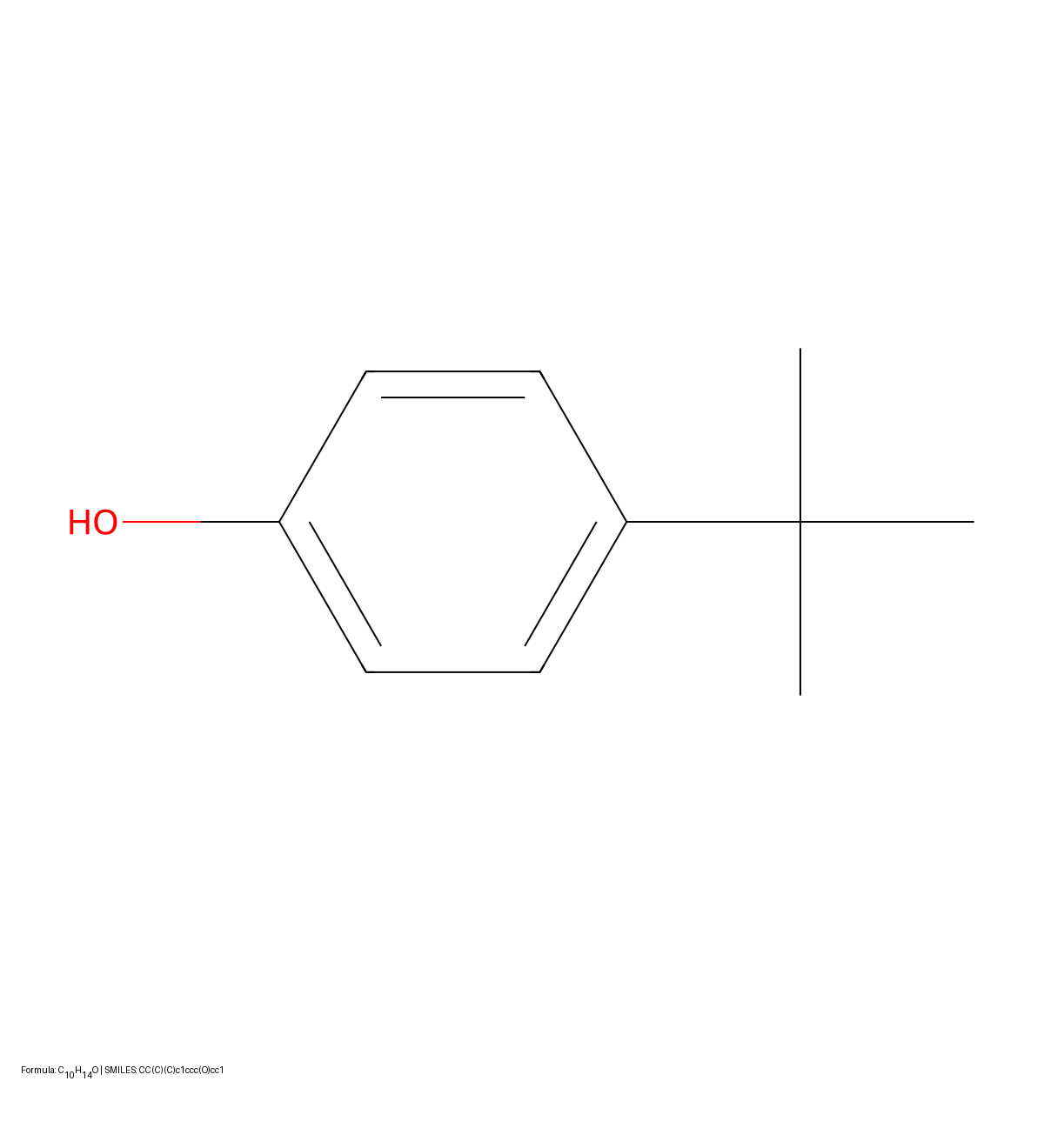
DECIMER 1.0 introduces a Transformer-based architecture coupled with EfficientNet-B3 to solve Optical Chemical Structure Recognition. By using the SELFIES representation (which guarantees 100% valid output strings) and scaling training to over 35 million molecules, it achieves 96.47% exact match accuracy on synthetic benchmarks, offering an open-source solution for mining chemical data from legacy literature.