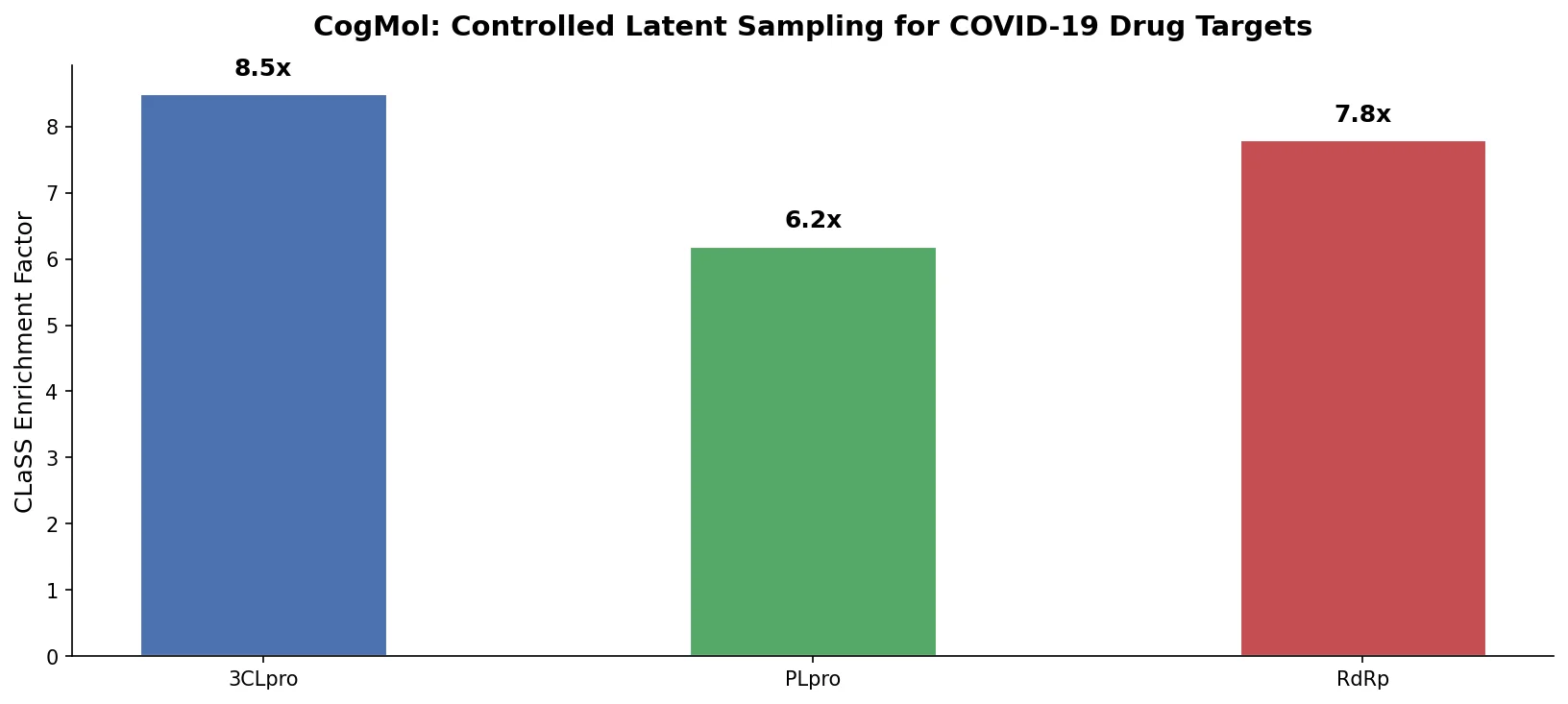
CogMol: Controlled Molecule Generation for COVID-19
CogMol uses a SMILES VAE and multi-attribute controlled sampling (CLaSS) to generate novel, target-specific drug molecules for unseen SARS-CoV-2 proteins without model retraining.

CogMol uses a SMILES VAE and multi-attribute controlled sampling (CLaSS) to generate novel, target-specific drug molecules for unseen SARS-CoV-2 proteins without model retraining.
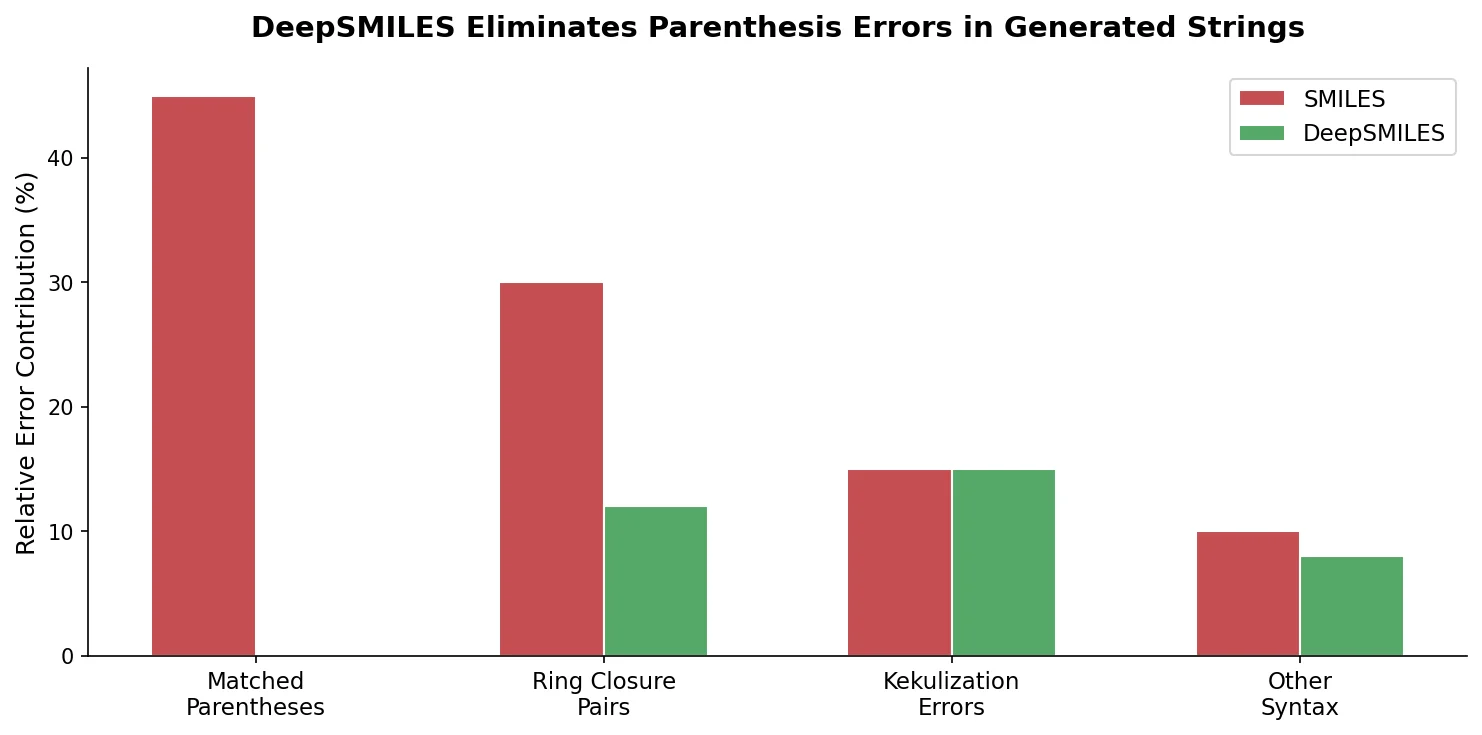
DeepSMILES replaces paired parentheses and ring closure symbols in SMILES with a postfix notation and single ring-size digits, making it easier for generative models to produce syntactically valid molecular strings.
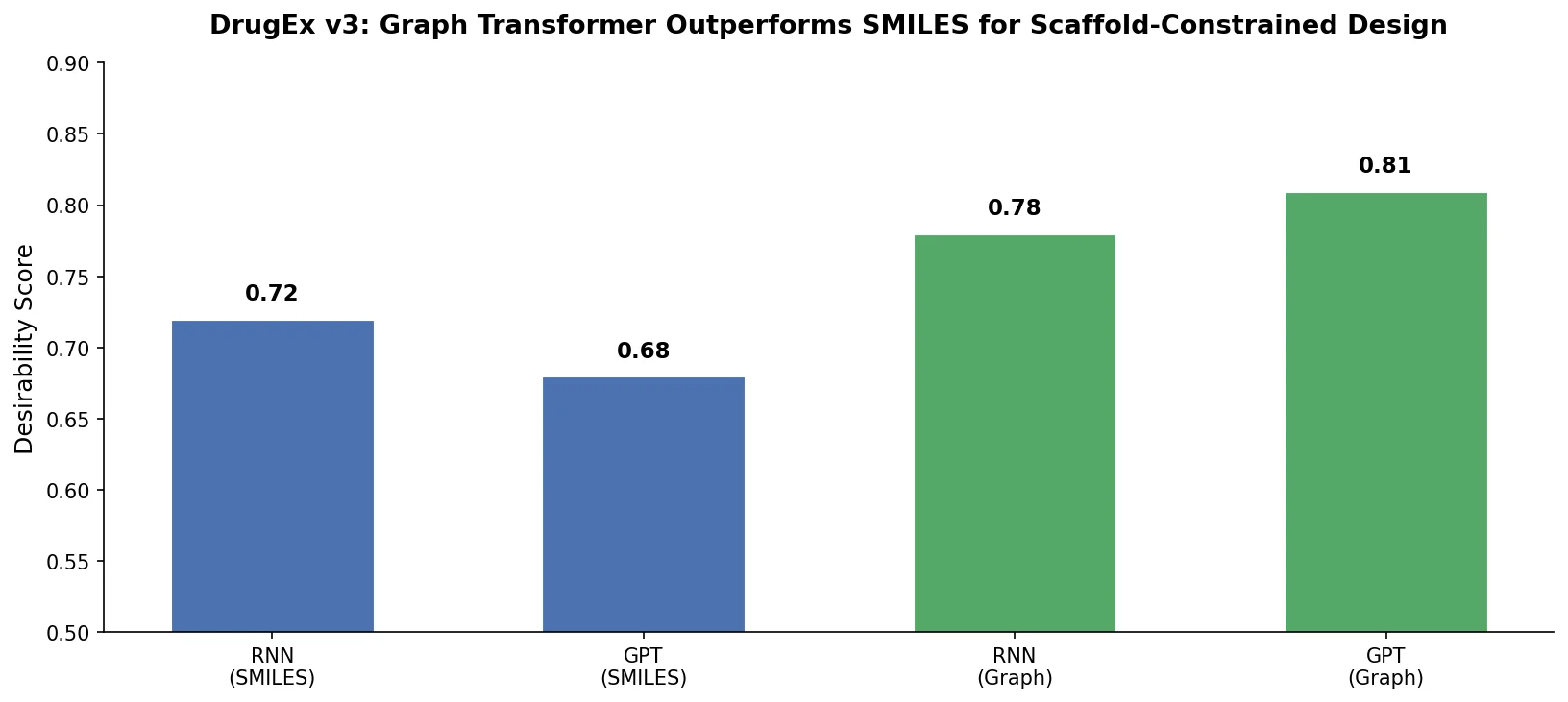
DrugEx v3 extends scaffold-constrained drug design by introducing a Graph Transformer with adjacency-matrix-based positional encoding, achieving 100% molecular validity and high predicted affinity for adenosine A2A receptor ligands.
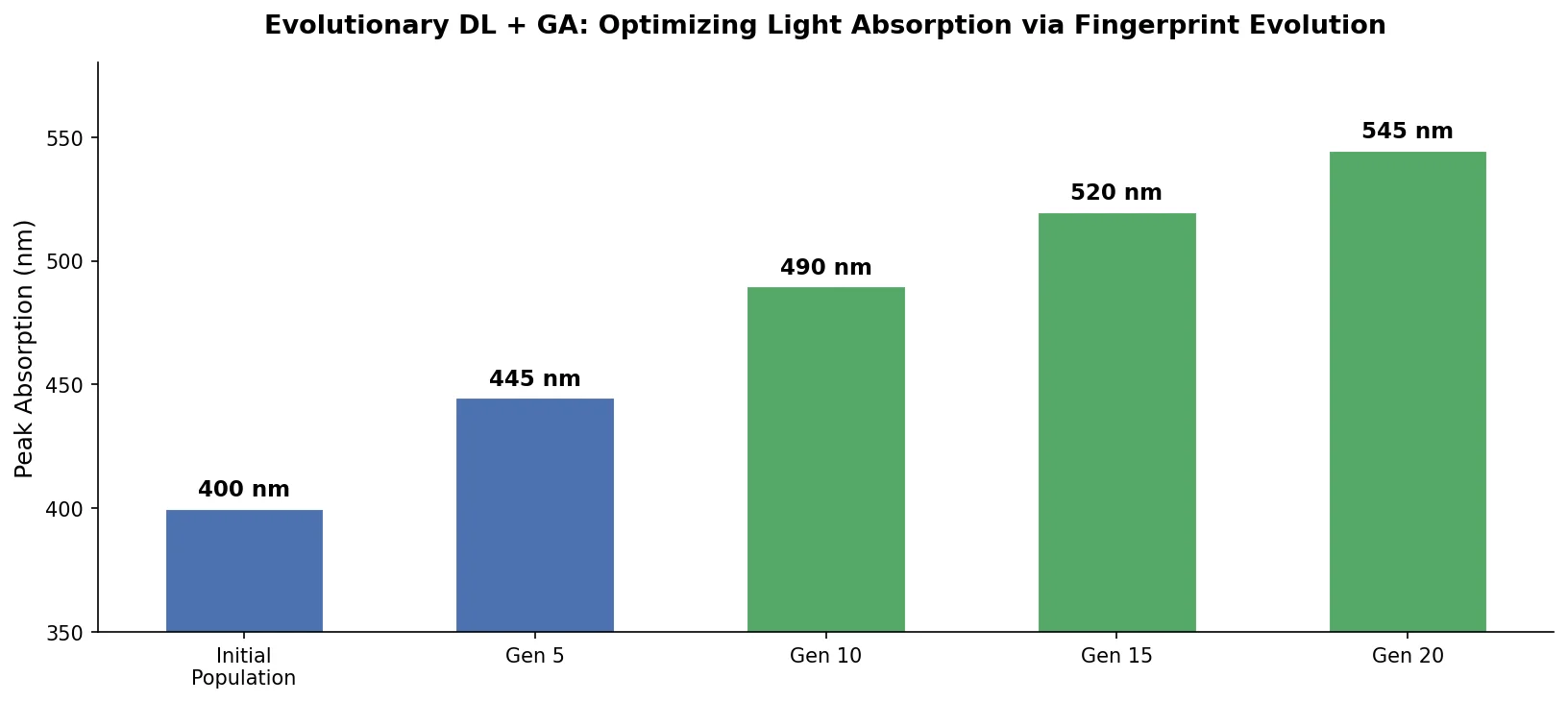
An evolutionary molecular design framework that evolves ECFP fingerprint vectors using a genetic algorithm, reconstructs valid SMILES via an RNN decoder, and evaluates fitness with a DNN property predictor.
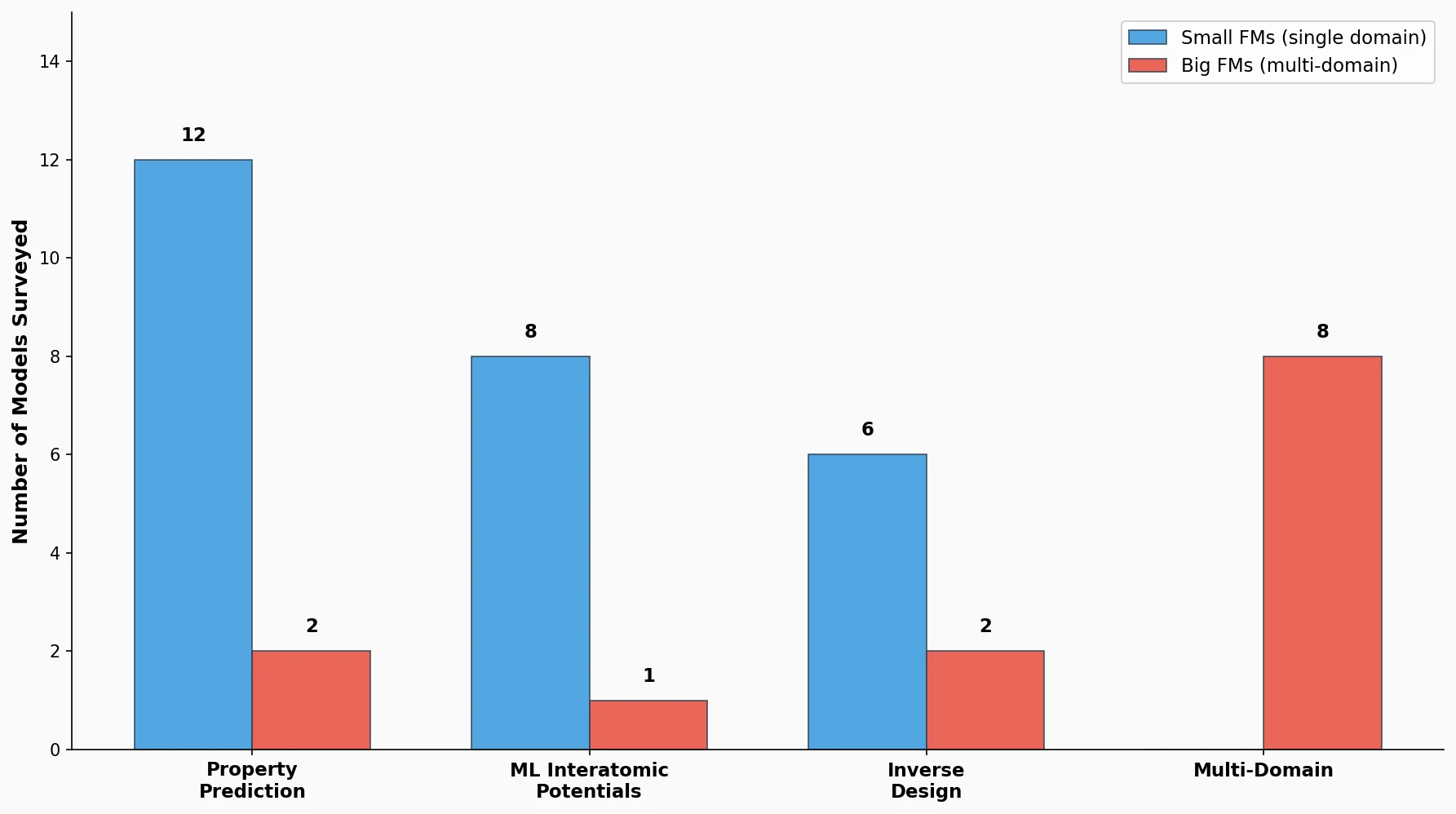
This perspective from Choi et al. reviews foundation models in chemistry, categorizing them as ‘small’ (domain-specific, e.g., property prediction, MLIPs, inverse design) and ‘big’ (multi-domain, e.g., multimodal and LLM-based). It surveys pretraining strategies, key architectures (GNNs and language models), and outlines future directions for scaling, efficiency, and interpretability.
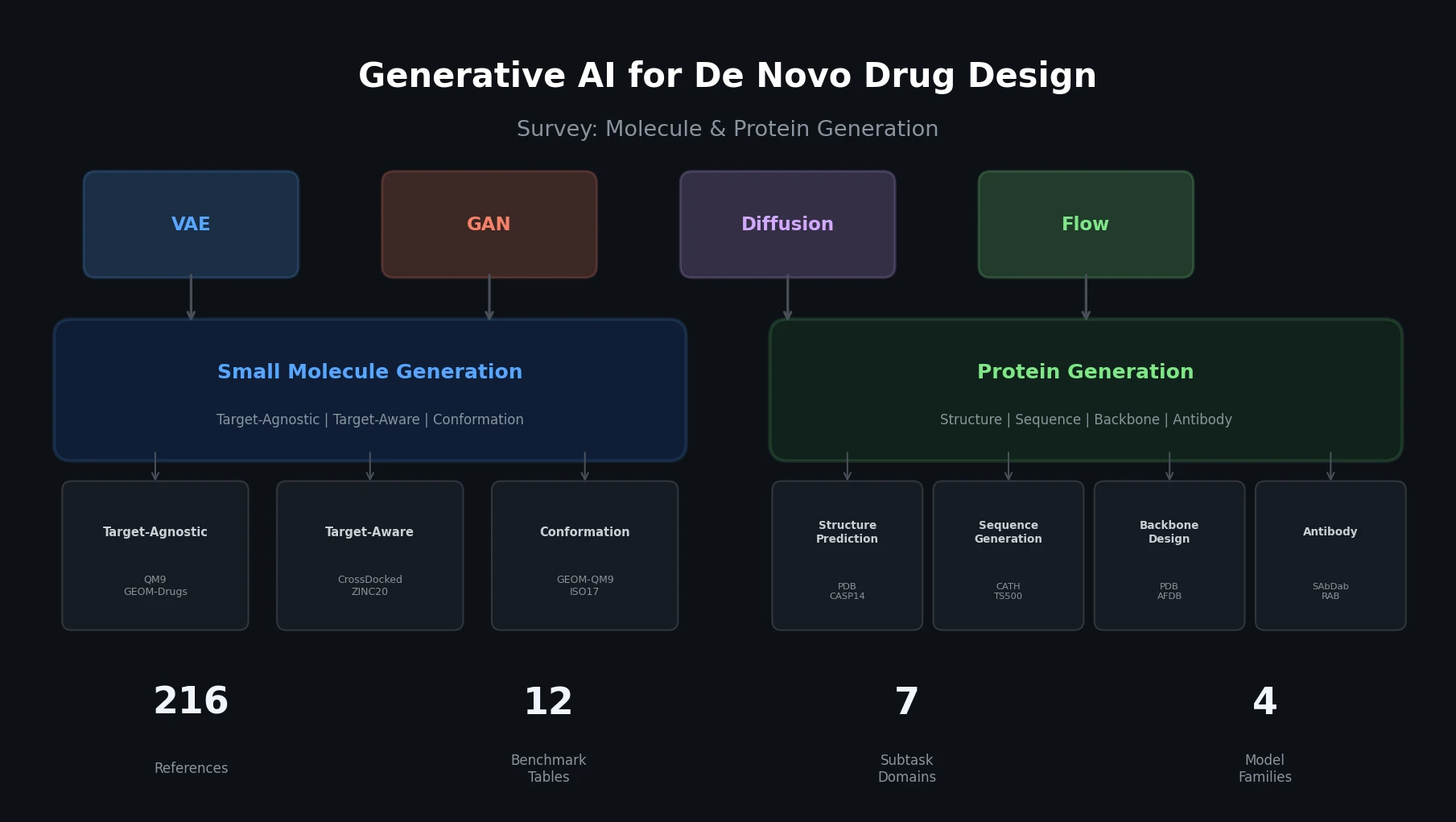
This survey organizes generative AI for de novo drug design into two themes: small molecule generation (target-agnostic, target-aware, conformation) and protein generation (structure prediction, sequence generation, backbone design, antibody, peptide). It covers four generative model families (VAEs, GANs, diffusion, flow-based), catalogs key datasets and benchmarks, and provides 12 comparative benchmark tables across all subtasks.
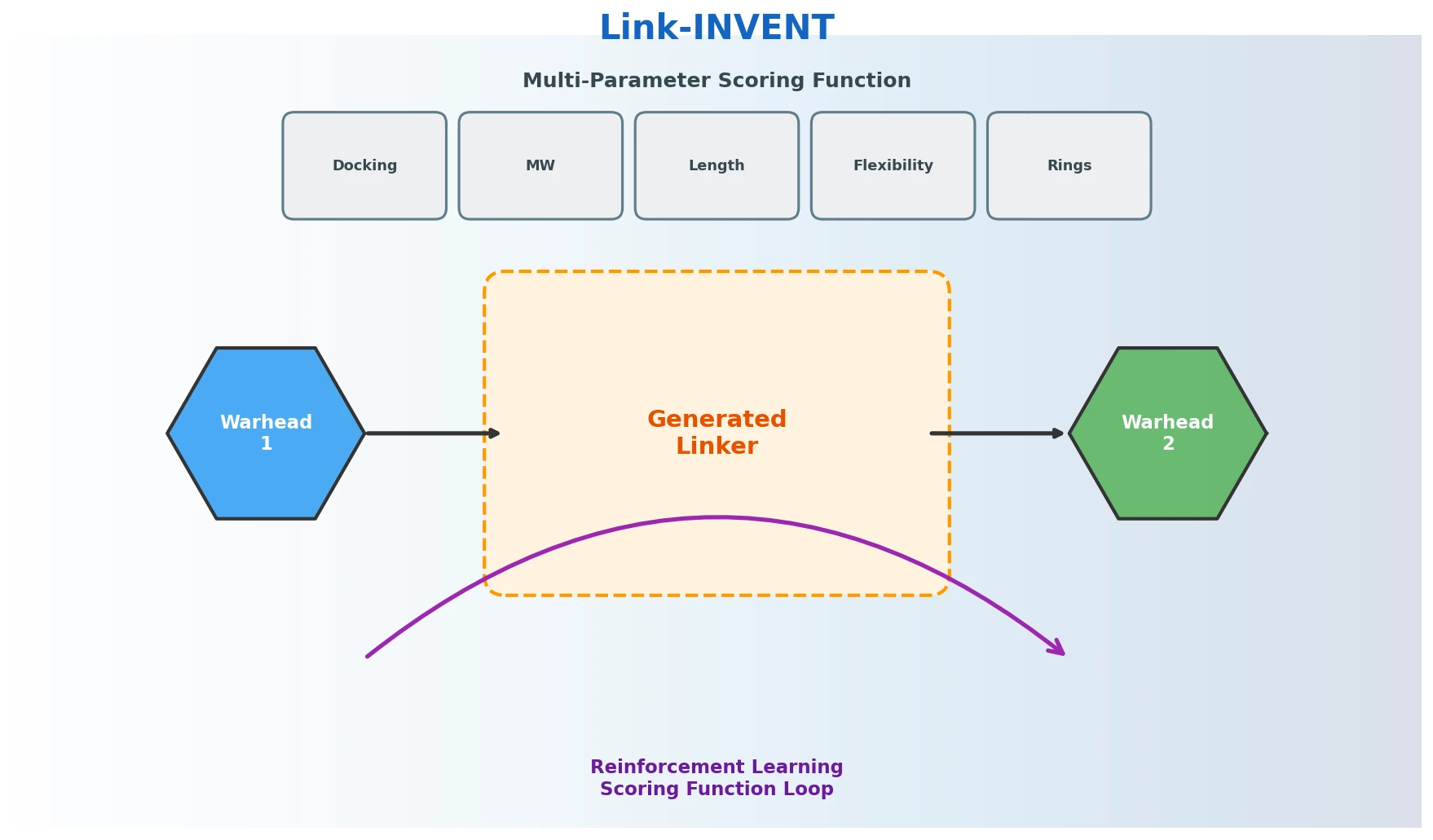
Link-INVENT is an RNN-based generative model for molecular linker design that uses reinforcement learning with a flexible scoring function, demonstrated on fragment linking, scaffold hopping, and PROTAC design.
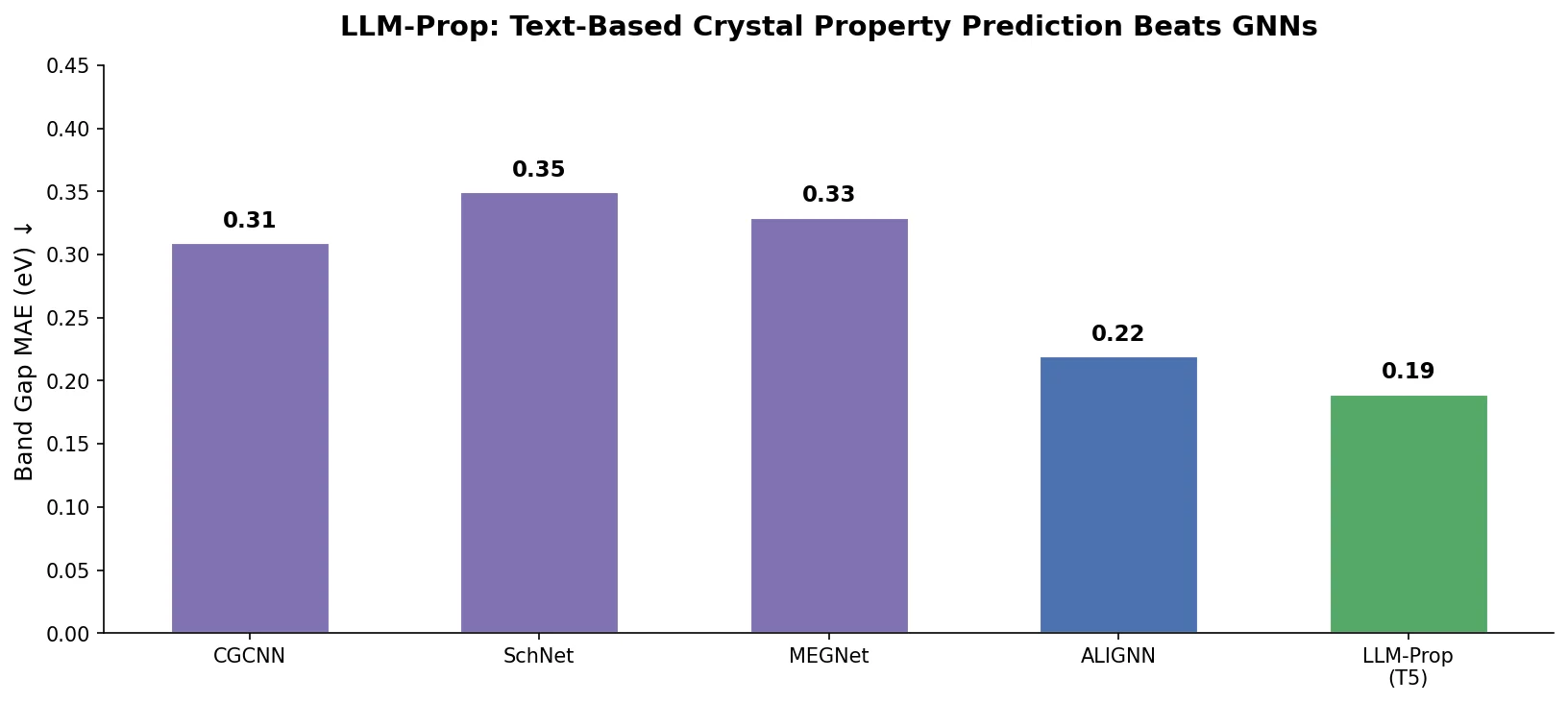
LLM-Prop uses the encoder half of T5, fine-tuned on Robocrystallographer text descriptions, to predict crystal properties. It outperforms GNN baselines like ALIGNN on band gap and volume prediction while using fewer parameters.
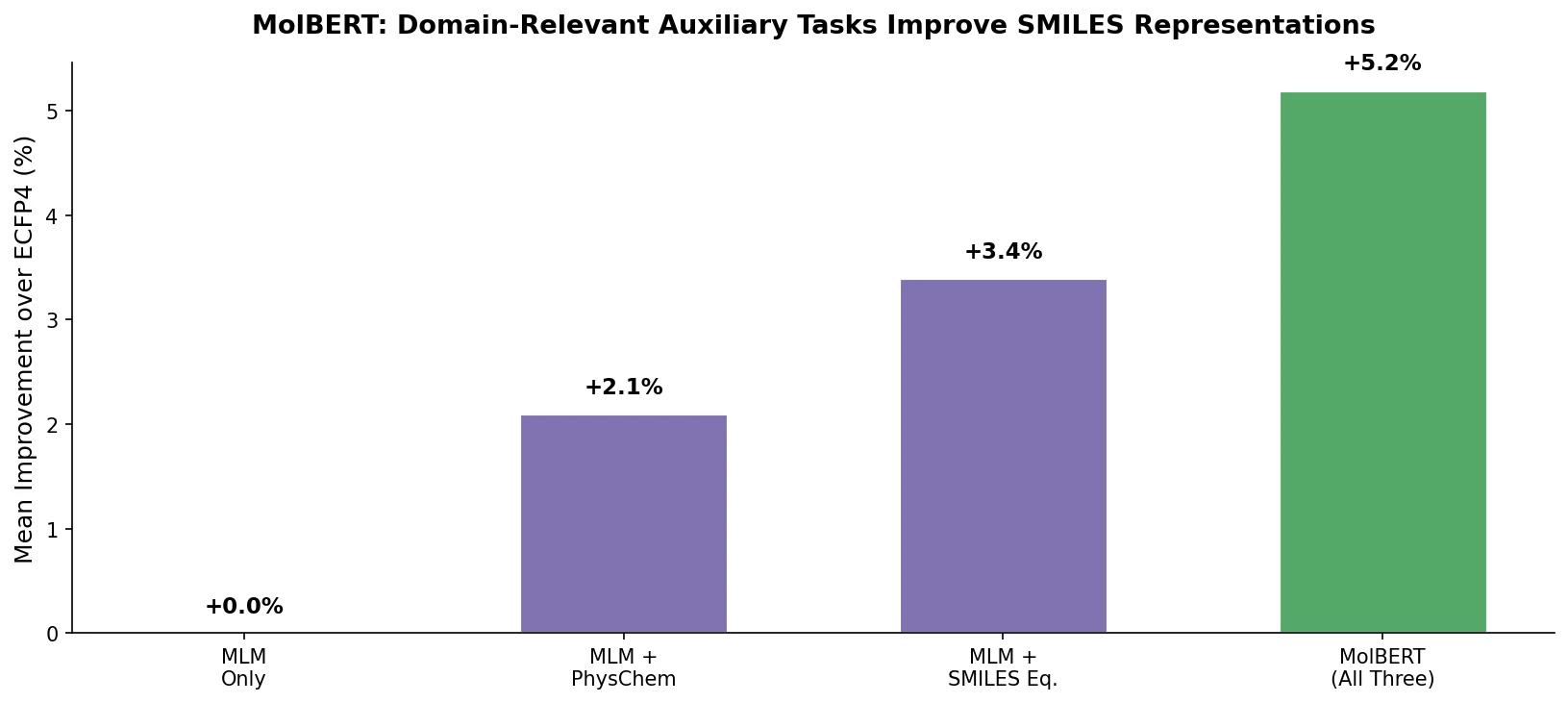
MolBERT pre-trains a BERT model on SMILES strings using masked language modeling, SMILES equivalence, and physicochemical property prediction as auxiliary tasks, achieving state-of-the-art results on virtual screening and QSAR benchmarks.
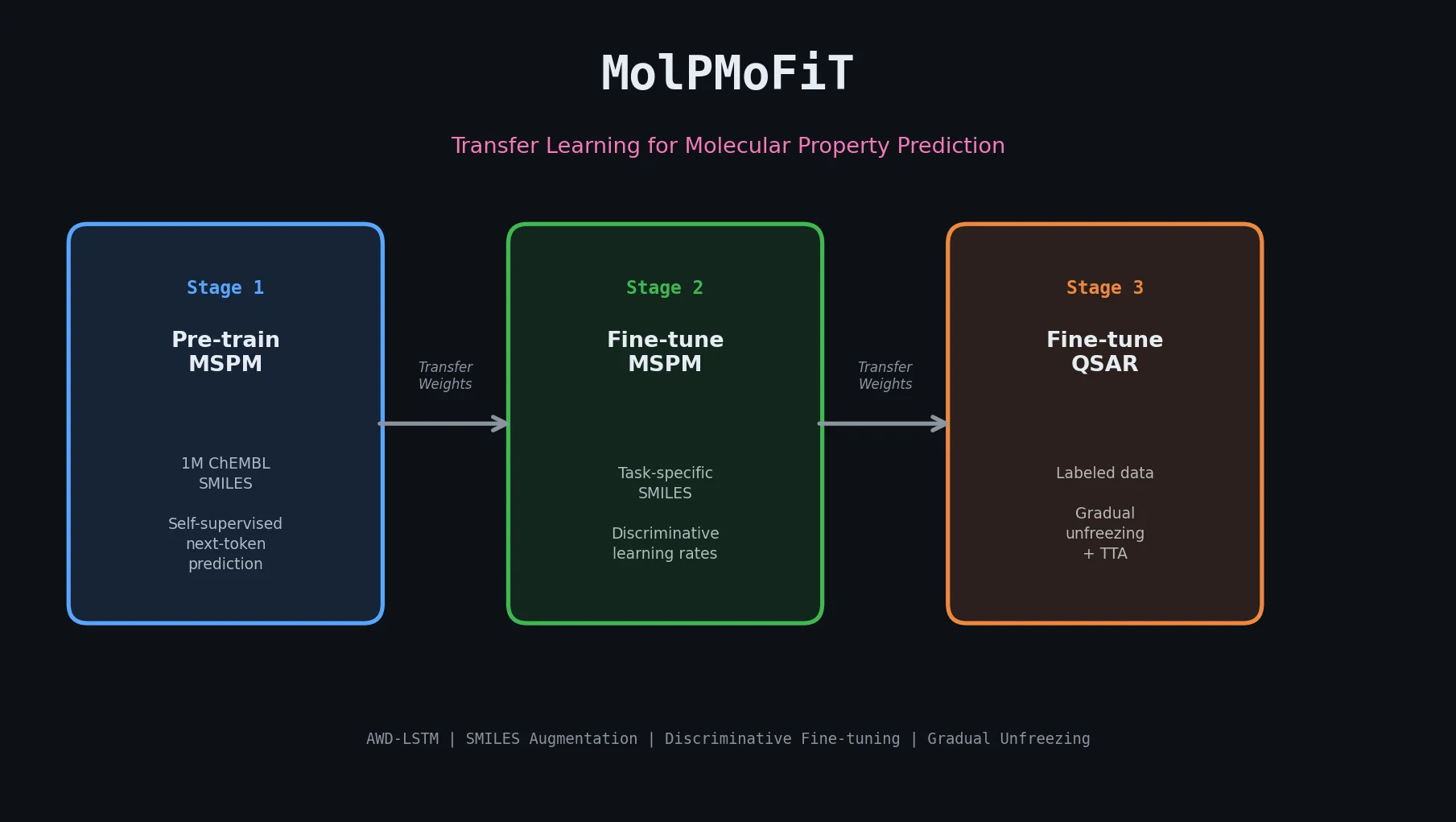
MolPMoFiT applies ULMFiT-style transfer learning to QSAR modeling, pre-training an AWD-LSTM on one million ChEMBL molecules and fine-tuning for property prediction on small datasets.
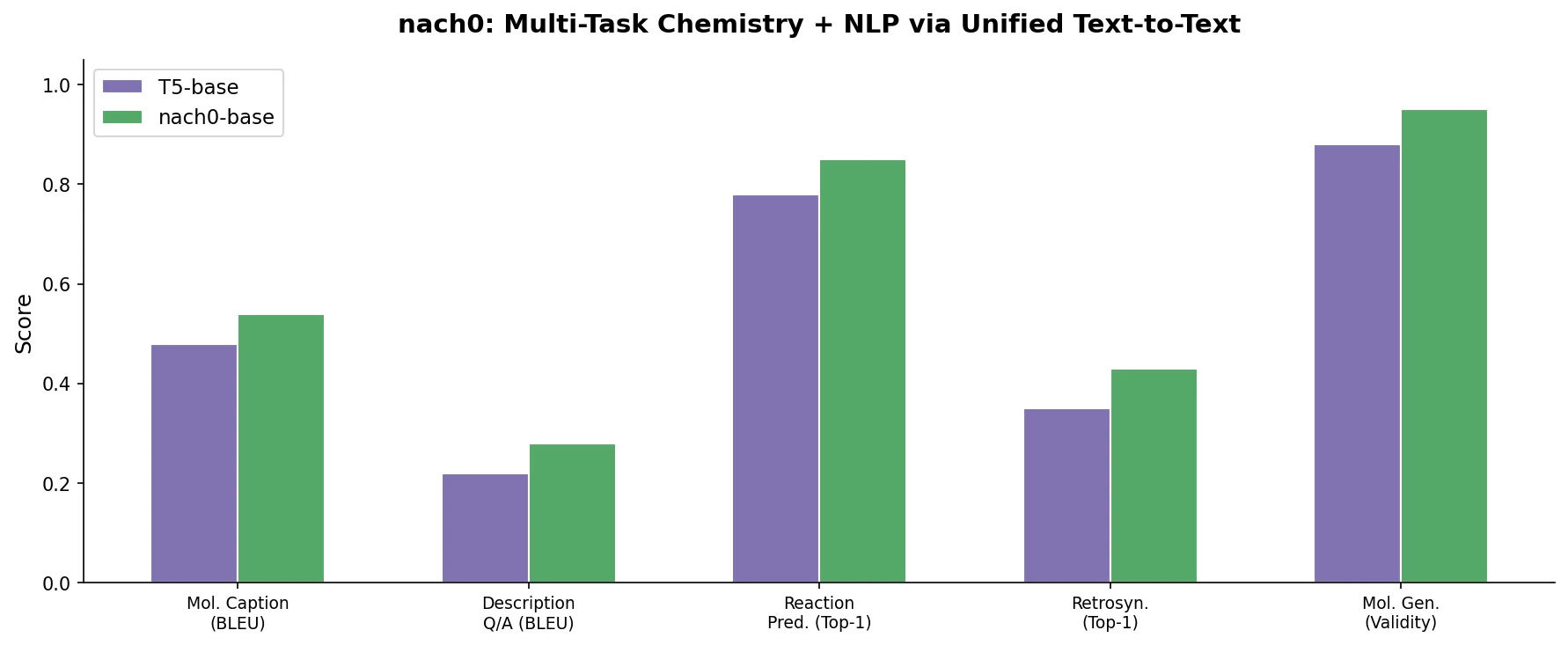
nach0 unifies natural language and SMILES-based chemical tasks in a single encoder-decoder model, achieving competitive results across molecular property prediction, reaction prediction, molecular generation, and biomedical NLP benchmarks.
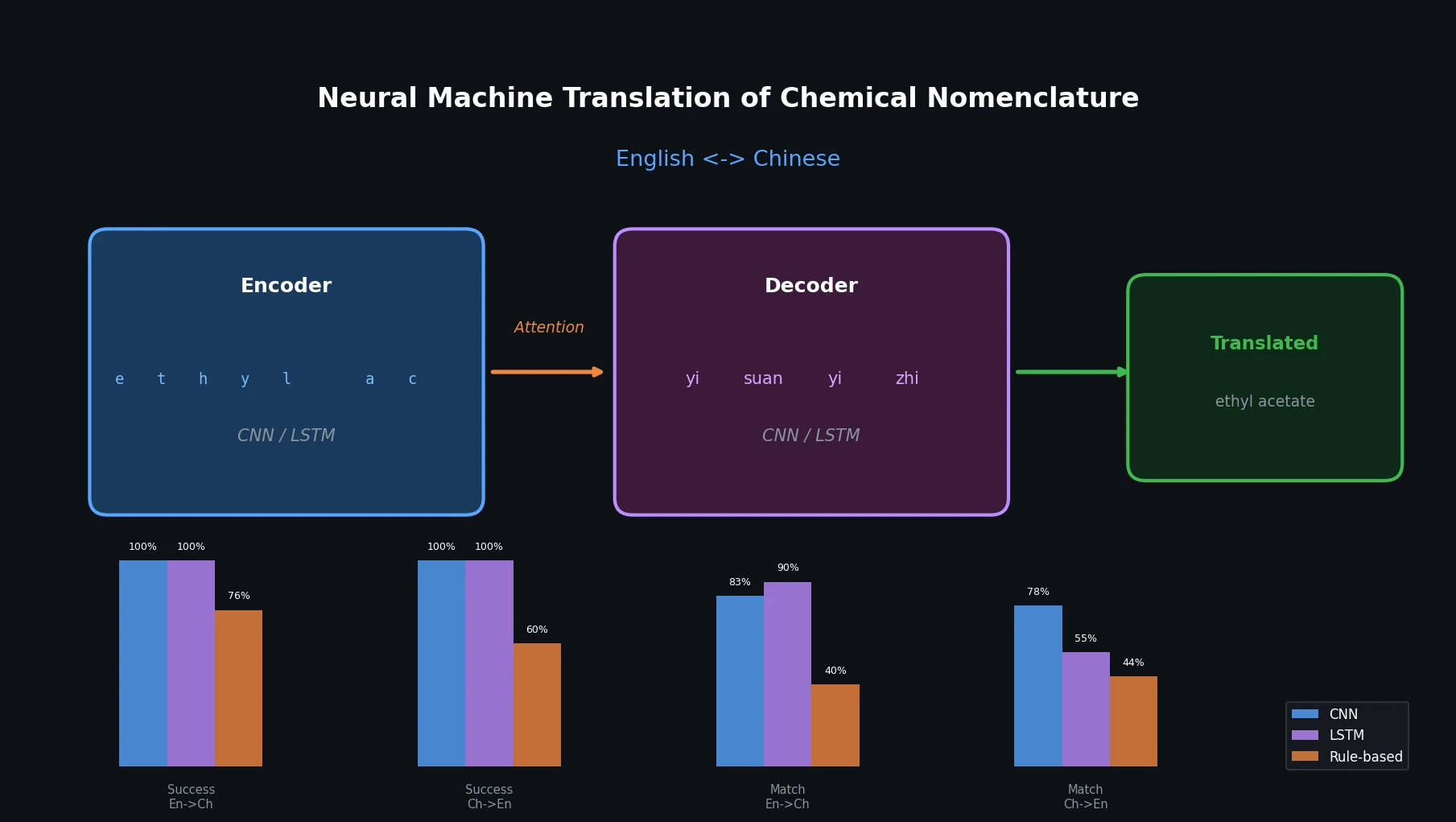
This paper applies character-level CNN and LSTM encoder-decoder networks to translate chemical names between English and Chinese, comparing them against an existing rule-based tool.