Image-to-Sequence OCSR: A Comparative Analysis
Deep dive into 24 image-to-sequence OCSR methods (2019-2025), comparing encoder-decoder architectures, molecular string representations, training scale, and hardware requirements.

Deep dive into 24 image-to-sequence OCSR methods (2019-2025), comparing encoder-decoder architectures, molecular string representations, training scale, and hardware requirements.
InstructMol integrates a pre-trained molecular graph encoder (MoleculeSTM) with a Vicuna-7B LLM using a linear projector. It employs a two-stage training process (alignment pre-training followed by task-specific instruction tuning with LoRA) to excel at property prediction, description generation, and reaction analysis.
InvMSAFold replaces autoregressive decoding with a Potts model parameter generator, enabling diverse protein sequence sampling orders of magnitude faster than ESM-IF1.
MOFFlow is the first deep generative model tailored for Metal-Organic Framework (MOF) structure prediction. It utilizes Riemannian flow matching on SE(3) to assemble rigid building blocks (metal nodes and organic linkers), achieving higher accuracy and scalability than atom-based methods on large systems.
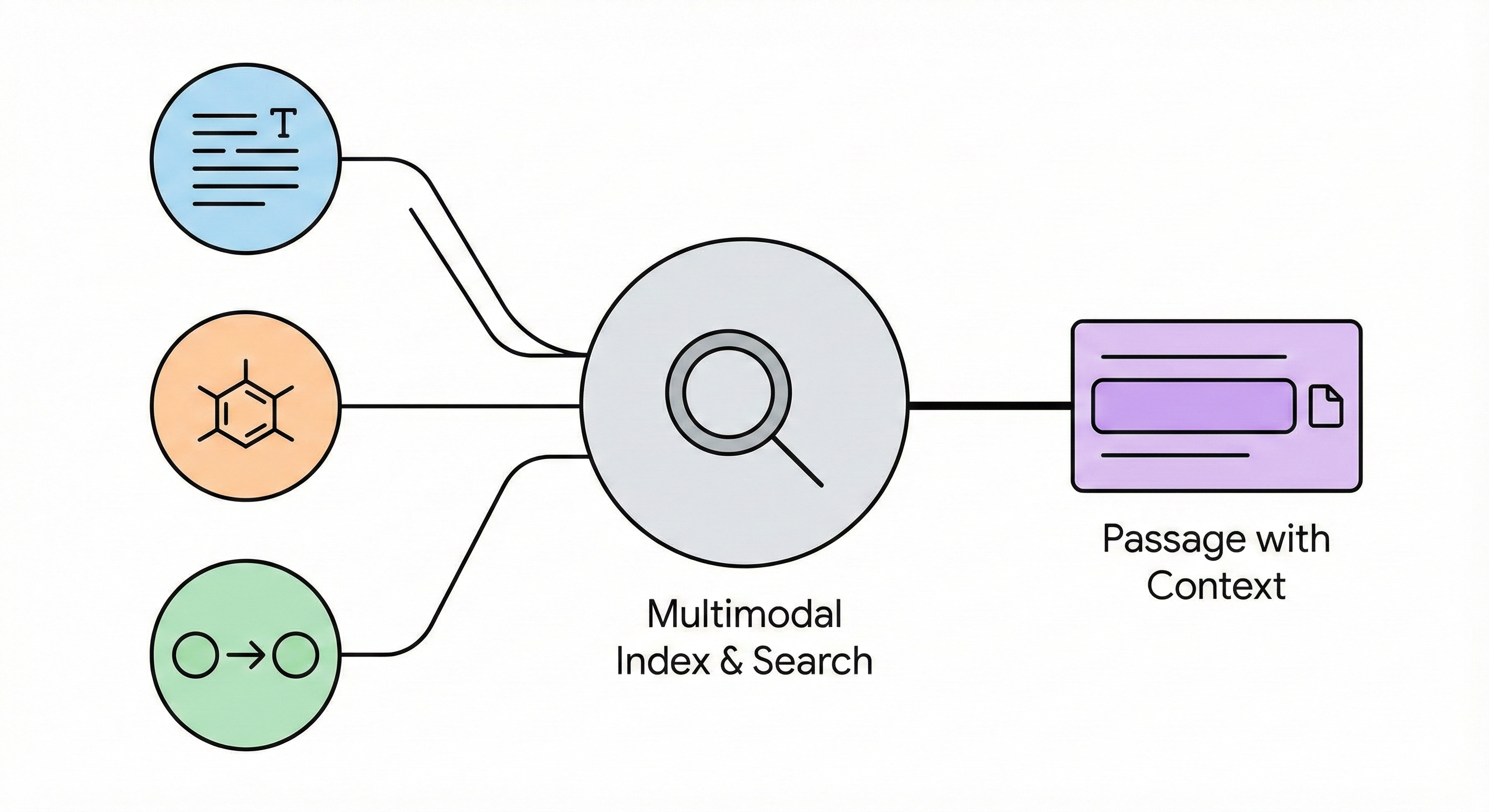
This paper presents a multimodal search system that facilitates passage-level retrieval of chemical reactions and molecular structures by linking diagrams, text, and reaction records extracted from scientific PDFs.
STOUT V2.0 uses Transformers trained on ~1 billion SMILES-IUPAC pairs to accurately translate chemical structures into systematic names (and vice-versa), outperforming its RNN predecessor.

STOUT (SMILES-TO-IUPAC-name translator) uses neural machine translation to convert chemical line notations to IUPAC names and vice versa, achieving ~90% BLEU score. It addresses the lack of open-source tools for algorithmic IUPAC naming.
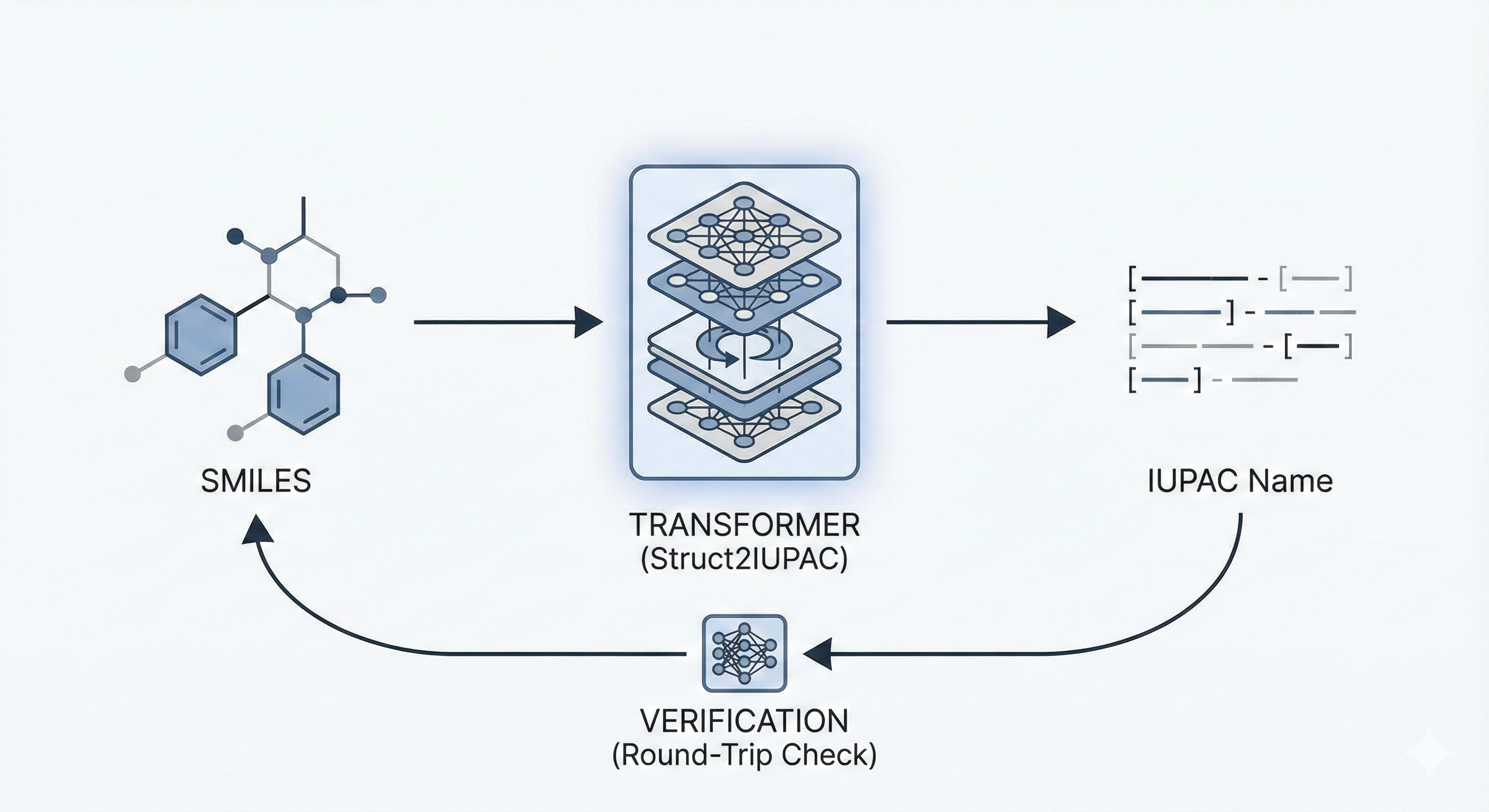
This paper proposes a Transformer-based approach (Struct2IUPAC) to convert chemical structures to IUPAC names, challenging the dominance of rule-based systems. Trained on ~47M PubChem examples, it achieves near-perfect accuracy using a round-trip verification step with OPSIN.
This study presents a sequence-to-sequence Transformer model that translates InChI identifiers into IUPAC names character-by-character. Trained on 10 million PubChem pairs, it achieves 91% accuracy on organic compounds, performing comparably to commercial software.
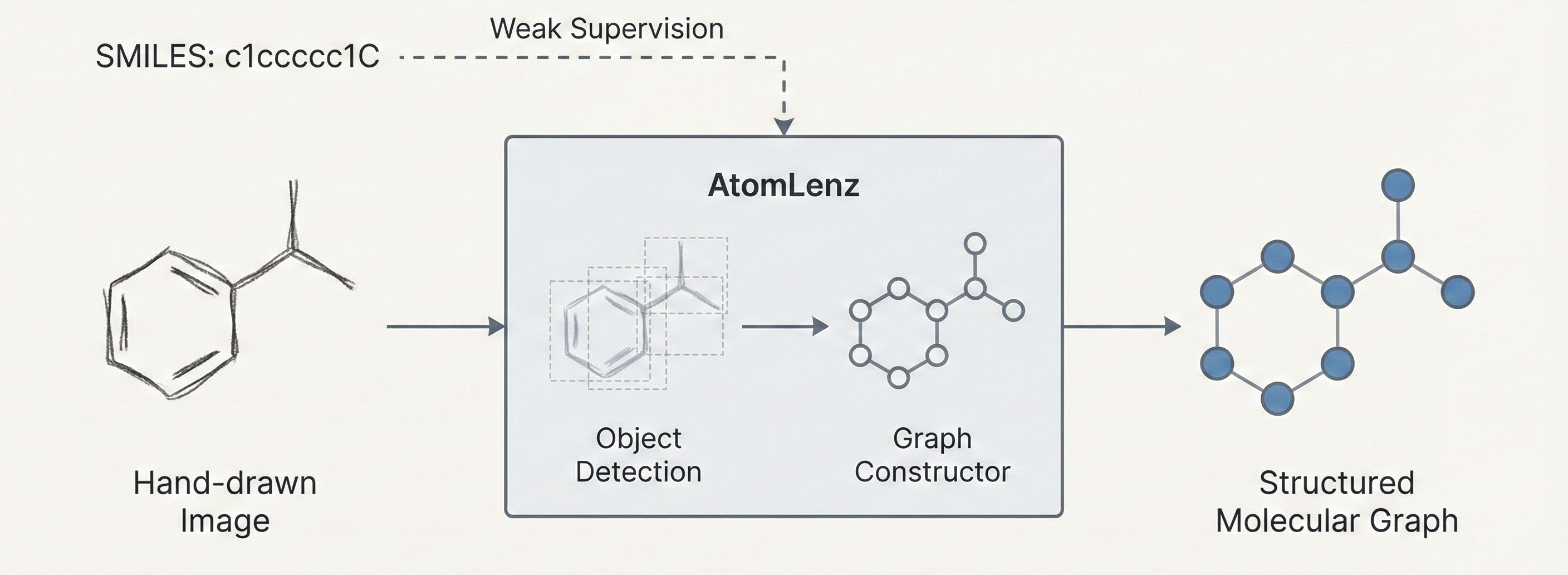
Introduces AtomLenz, an OCSR tool that combines object detection with a molecular graph constructor. Features a novel weakly supervised training scheme (ProbKT*) to learn atom-level localization from SMILES-only data, achieving state-of-the-art results on hand-drawn images.
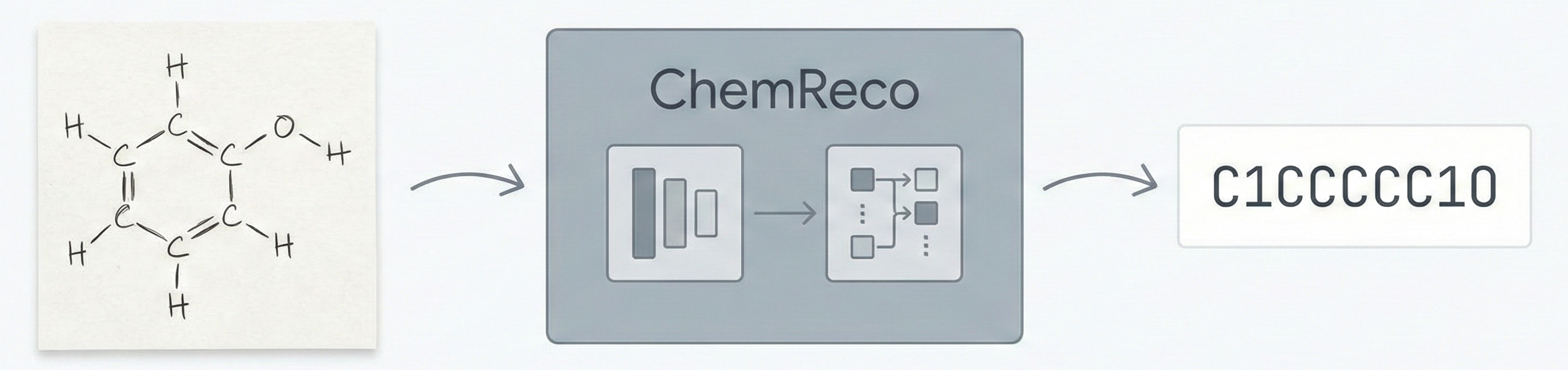
ChemReco automates the recognition of hand-drawn chemical structures using a synthetic data pipeline and an EfficientNet+Transformer architecture, achieving 96.90% accuracy on C-H-O molecules.
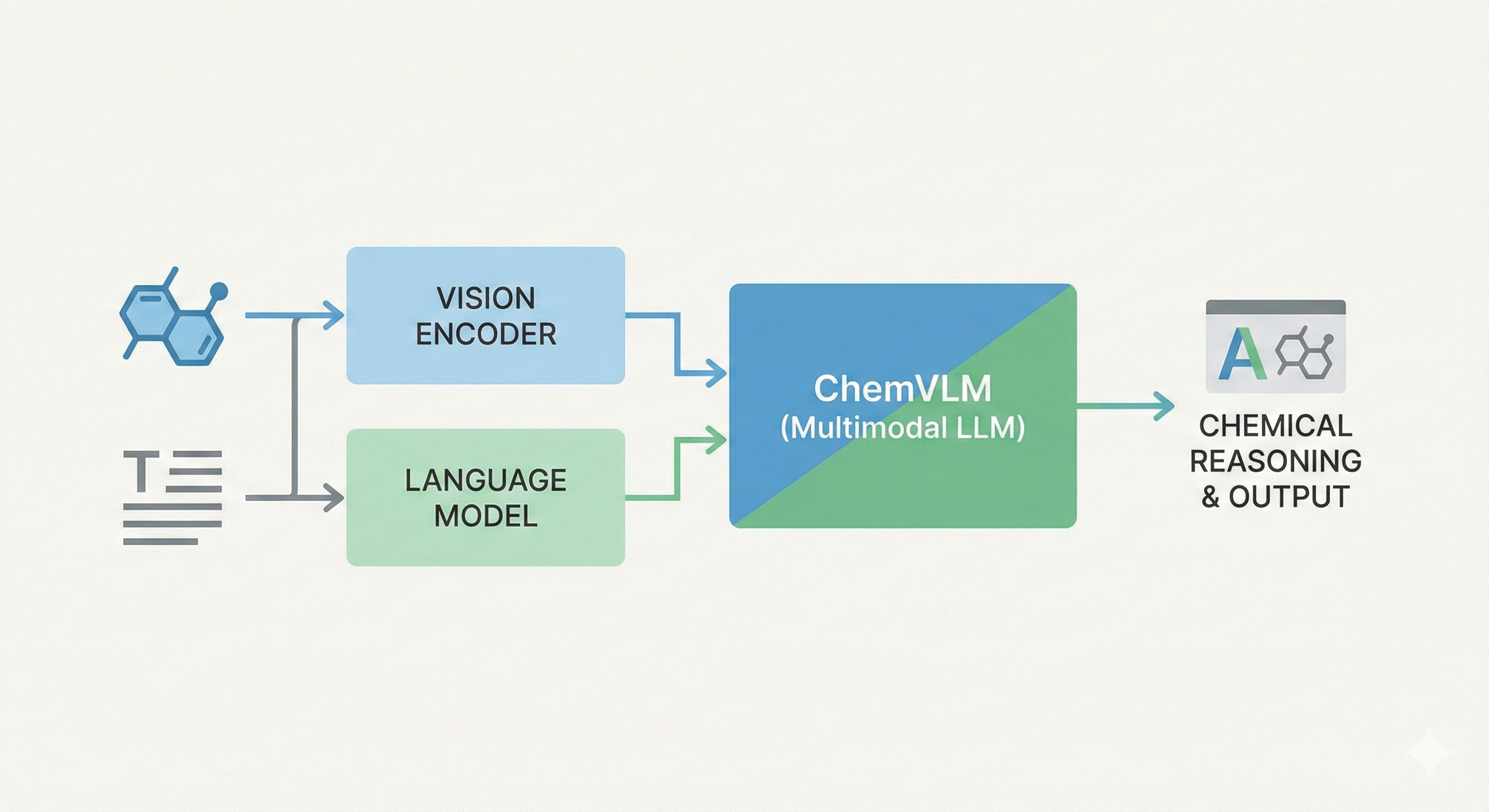
A 2025 AAAI paper introducing ChemVLM, a domain-specific multimodal LLM (26B parameters). It achieves state-of-the-art performance on chemical OCR, reasoning benchmarks, and molecular understanding tasks by combining vision and language models trained on curated chemistry data.