Lagrangian Neural Networks for Physics
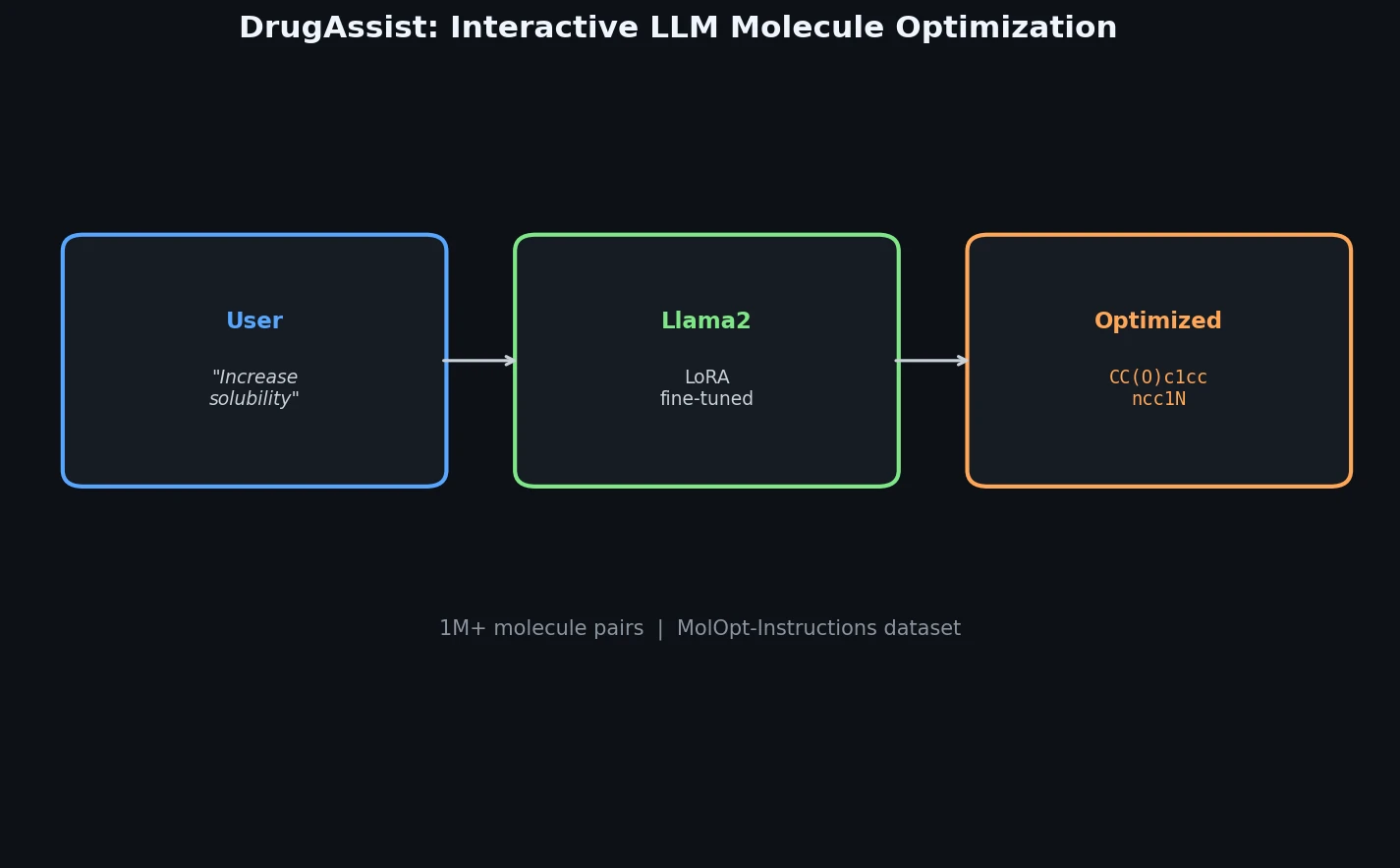
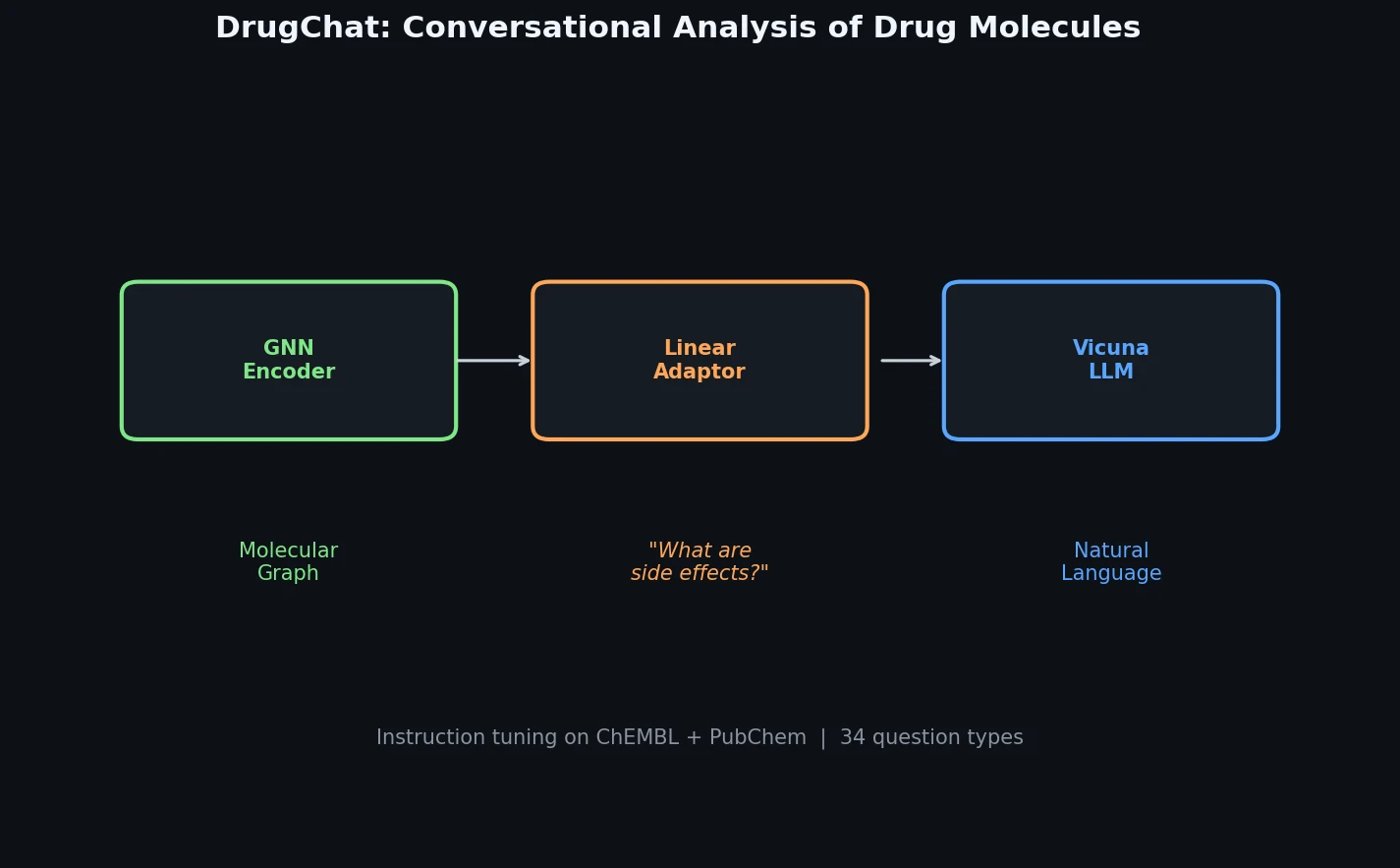
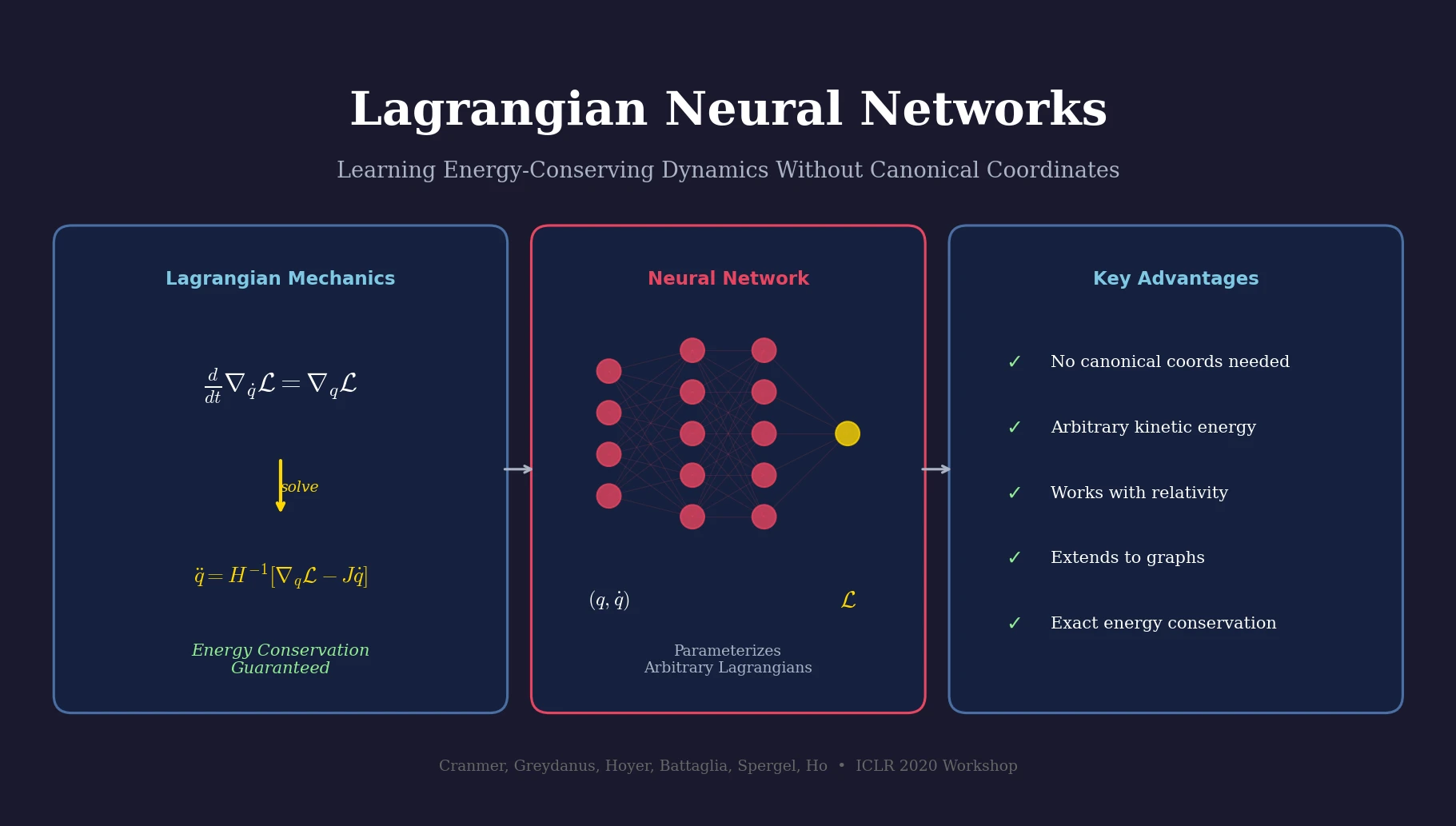
Lagrangian Neural Networks (LNNs) use neural networks to parameterize arbitrary Lagrangians, enabling energy-conserving learned dynamics without canonical coordinates. Unlike Hamiltonian approaches, LNNs handle relativistic systems and extend to graphs via Lagrangian Graph Networks.