AdaptMol: Domain Adaptation for Molecular OCSR (2026)
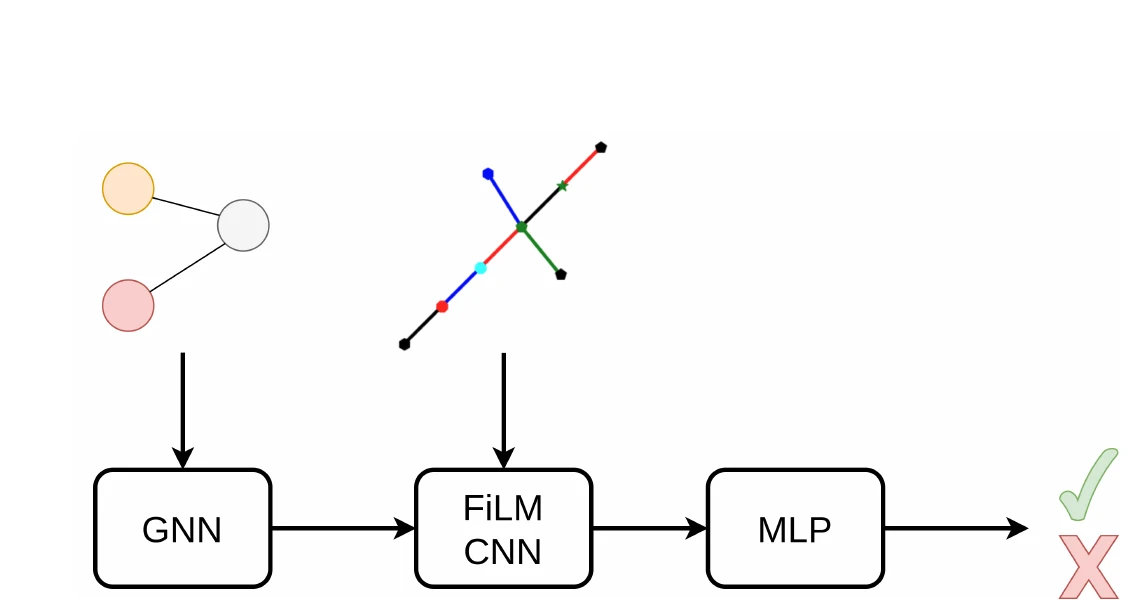
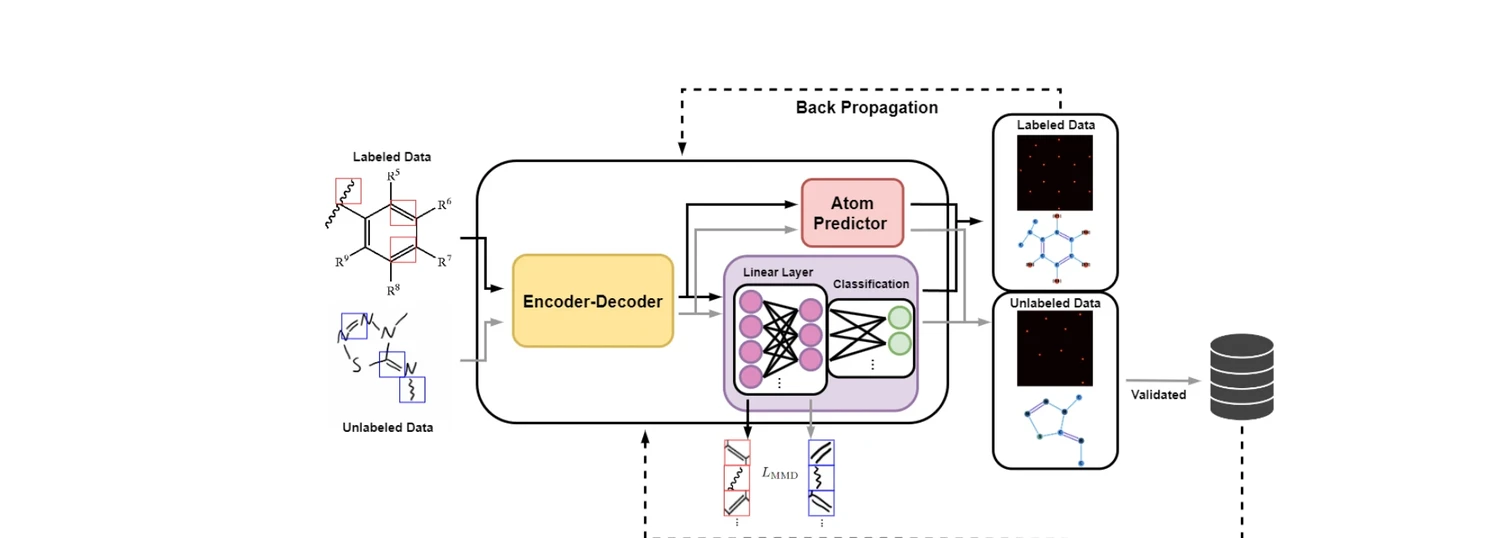
AdaptMol combines an end-to-end graph reconstruction model with unsupervised domain adaptation via class-conditional MMD on bond features and SMILES-validated self-training. Achieves 82.6% accuracy on hand-drawn molecules (10.7 points above prior best) while maintaining state-of-the-art results on four literature benchmarks, using only 4,080 real hand-drawn images for adaptation.