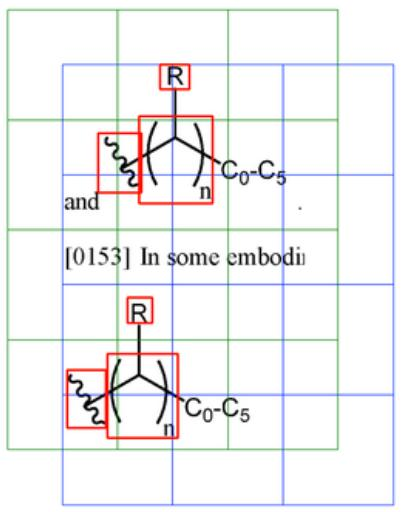
One Strike, You're Out: Detecting Markush Structures
Proposes a patch-based image processing pipeline using Inception V3 to filter Markush structures from chemical documents, outperforming traditional fixed-feature (ORB) methods on low-SNR images.

Proposes a patch-based image processing pipeline using Inception V3 to filter Markush structures from chemical documents, outperforming traditional fixed-feature (ORB) methods on low-SNR images.
This empirical study isolates the impact of chemical string representations on image-to-text translation models. It finds that while SMILES offers the highest overall accuracy, SELFIES provides a guarantee of structural validity, offering a trade-off for OCSR tasks.
Proposes an end-to-end architecture replacing standard CNN backbones with Swin Transformer to capture global image context. Introduces Multi-label Focal Loss to handle severe token imbalance in chemical datasets.
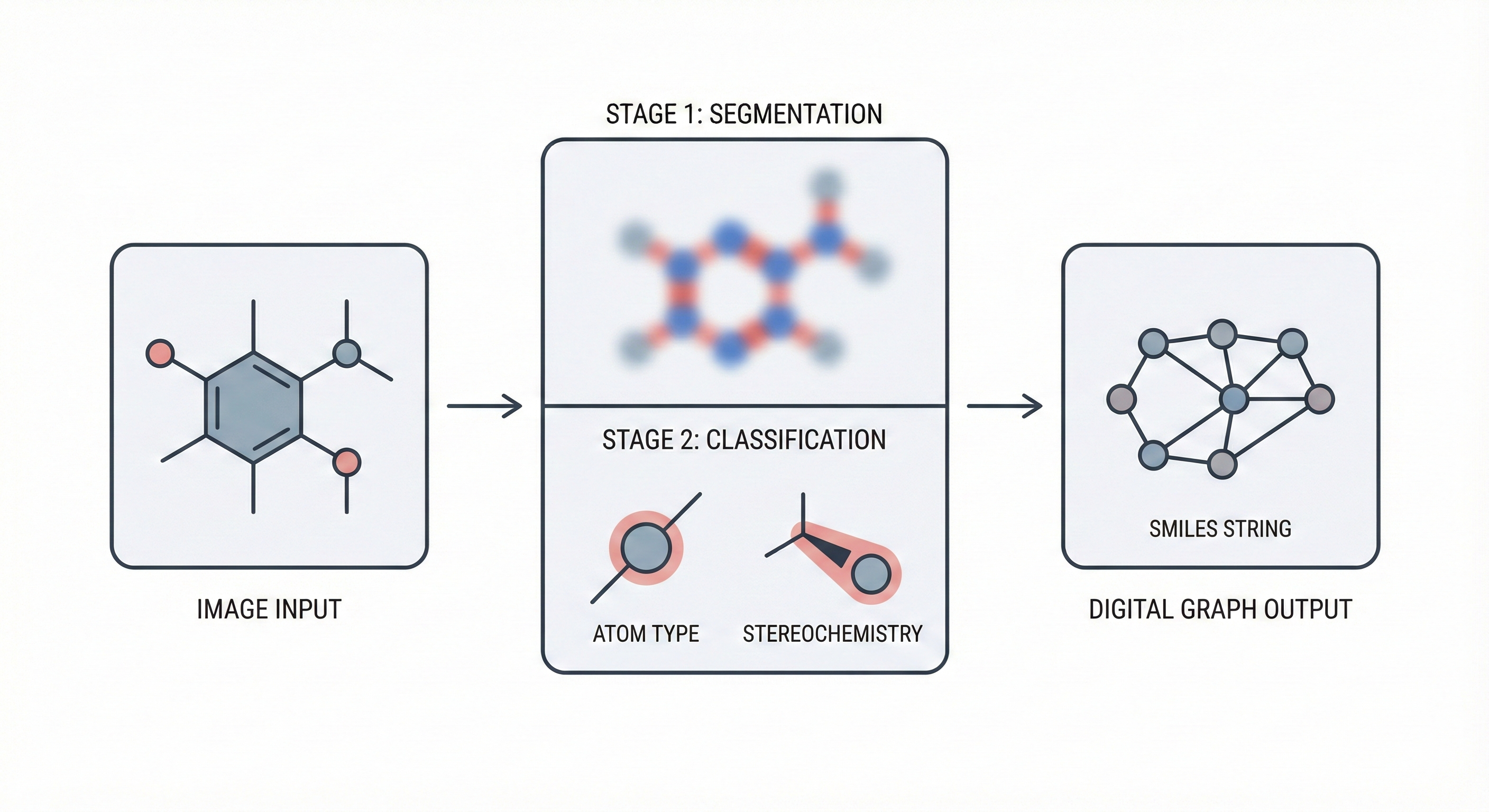
ChemGrapher replaces rule-based chemical OCR with a deep learning pipeline using semantic segmentation to identify atom and bond candidates, followed by specialized classification networks to resolve stereochemistry and bond multiplicity, reducing error rates compared to OSRA across all tested styles.
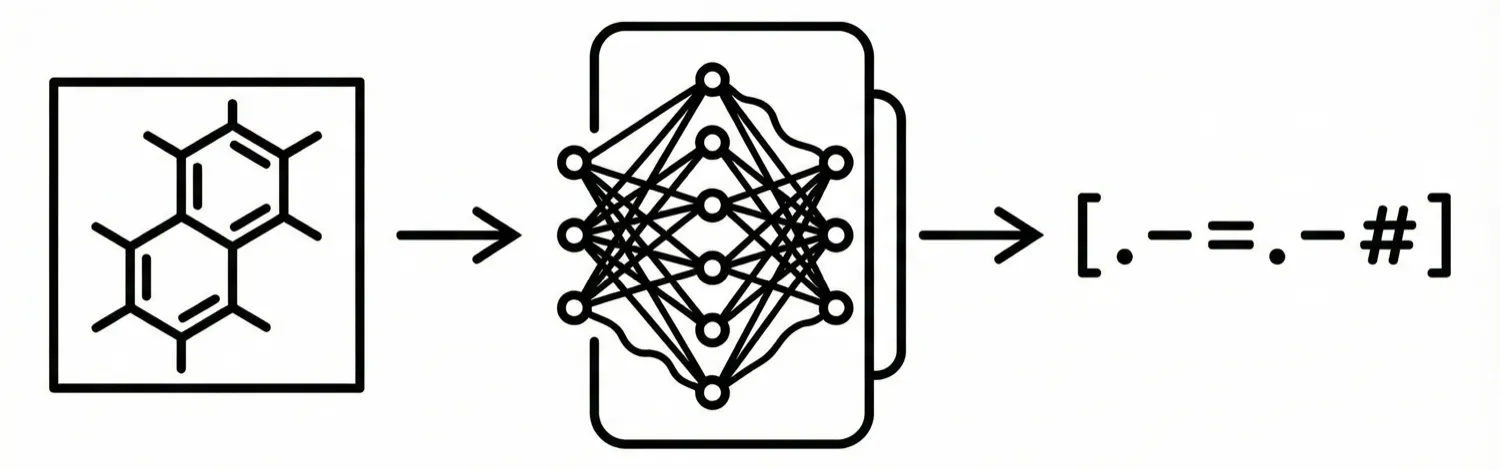
DECIMER adapts the “Show, Attend and Tell” image captioning architecture to translate chemical structure images into SMILES strings. By leveraging massive synthetic datasets generated from PubChem, it demonstrates that deep learning can perform optical chemical recognition without complex, hand-engineered rule systems.
This paper presents a two-stage deep learning pipeline to extract chemical structures from documents and convert them to SMILES strings. By training on large-scale synthetic data, the method overcomes the brittleness of rule-based systems and demonstrates high accuracy even on low-resolution and noisy input images.
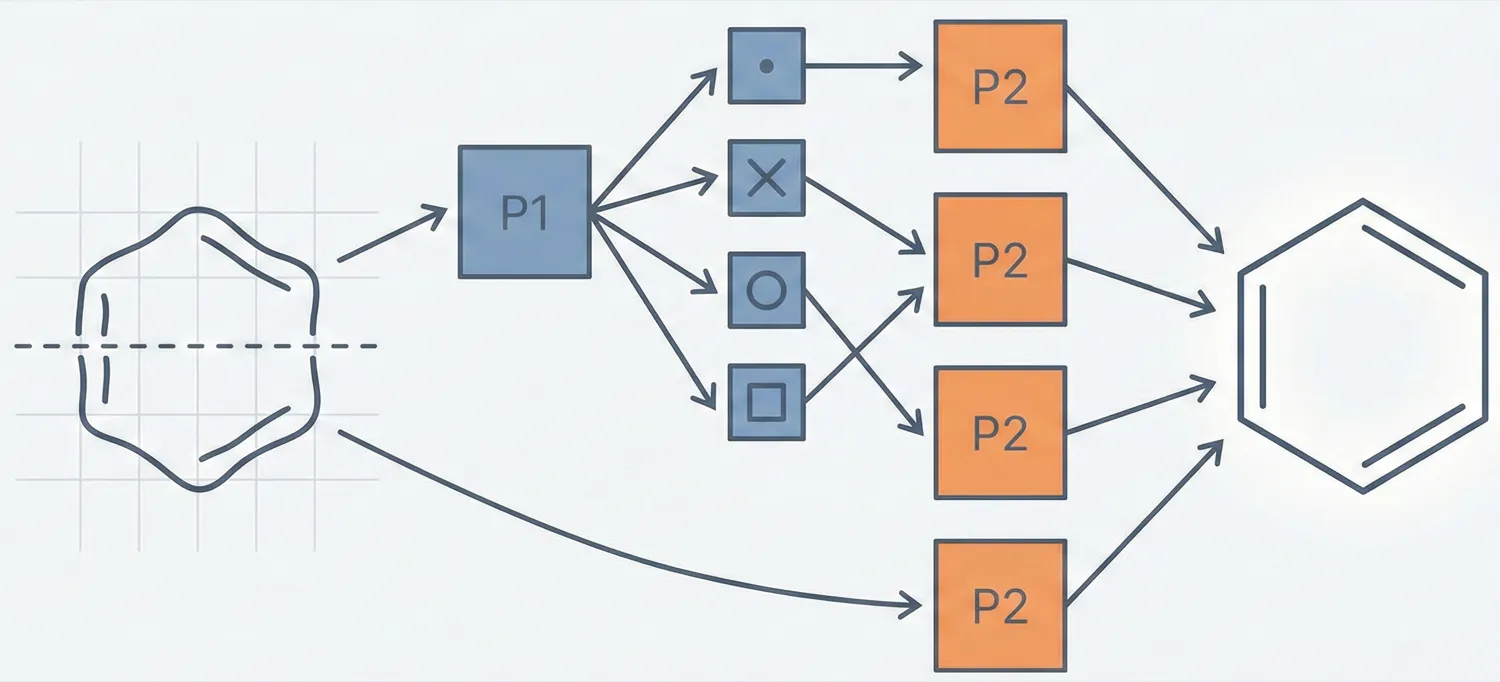
Proposes a specialized Classifier-Recognizer architecture that first categorizes rings by heteroatom (S, N, O) and then identifies the specific ring using optimized grid inputs.
A 2021 deep learning system using a two-stage approach for OCSR, encoding images into continuous CDDD embeddings before decoding to SMILES. It leverages extensive data augmentation to handle rotations, distortions, and rendering variations for fast and robust molecular structure recognition.
This paper introduces Kekulé-1, one of the first successful Optical Chemical Structure Recognition (OCSR) systems. It details a hybrid approach using neural networks for character recognition and heuristic vectorization for bond detection, achieving 98.9% accuracy on a test set of 524 structures.
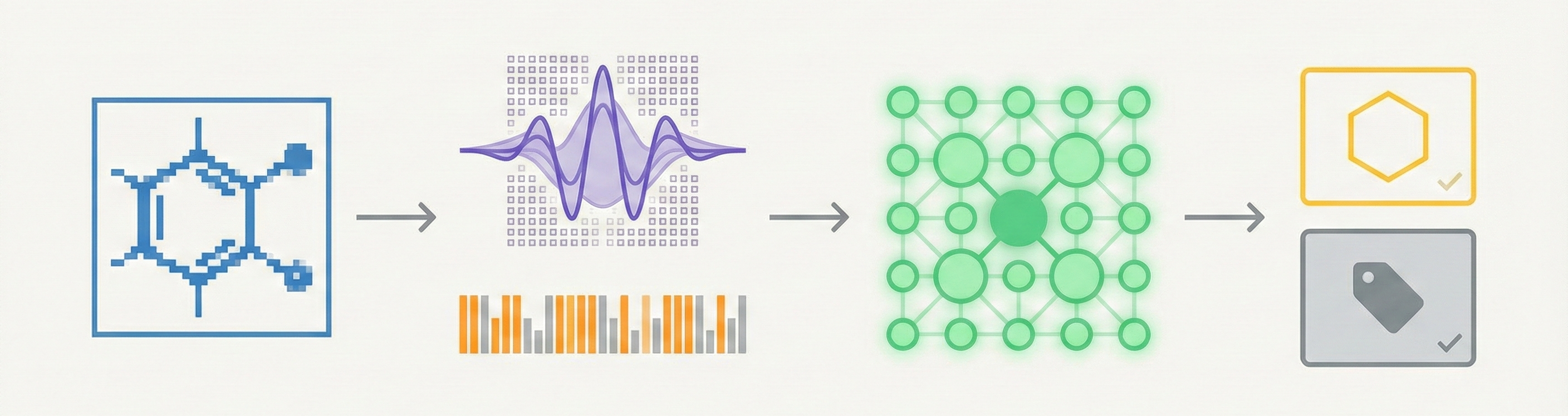
This 2003 paper introduces a machine vision approach for extracting chemical metadata from raster images. By using Gabor wavelets for feature extraction and Kohonen networks for classification, it distinguishes between chemical and non-chemical images, as well as ring and non-ring systems, without requiring high-resolution inputs.

Geoffrey Hinton’s 1984 technical report that formally derives the efficiency of distributed representations (coarse coding) and demonstrates their properties of automatic generalization, content-addressability, and robustness to damage.
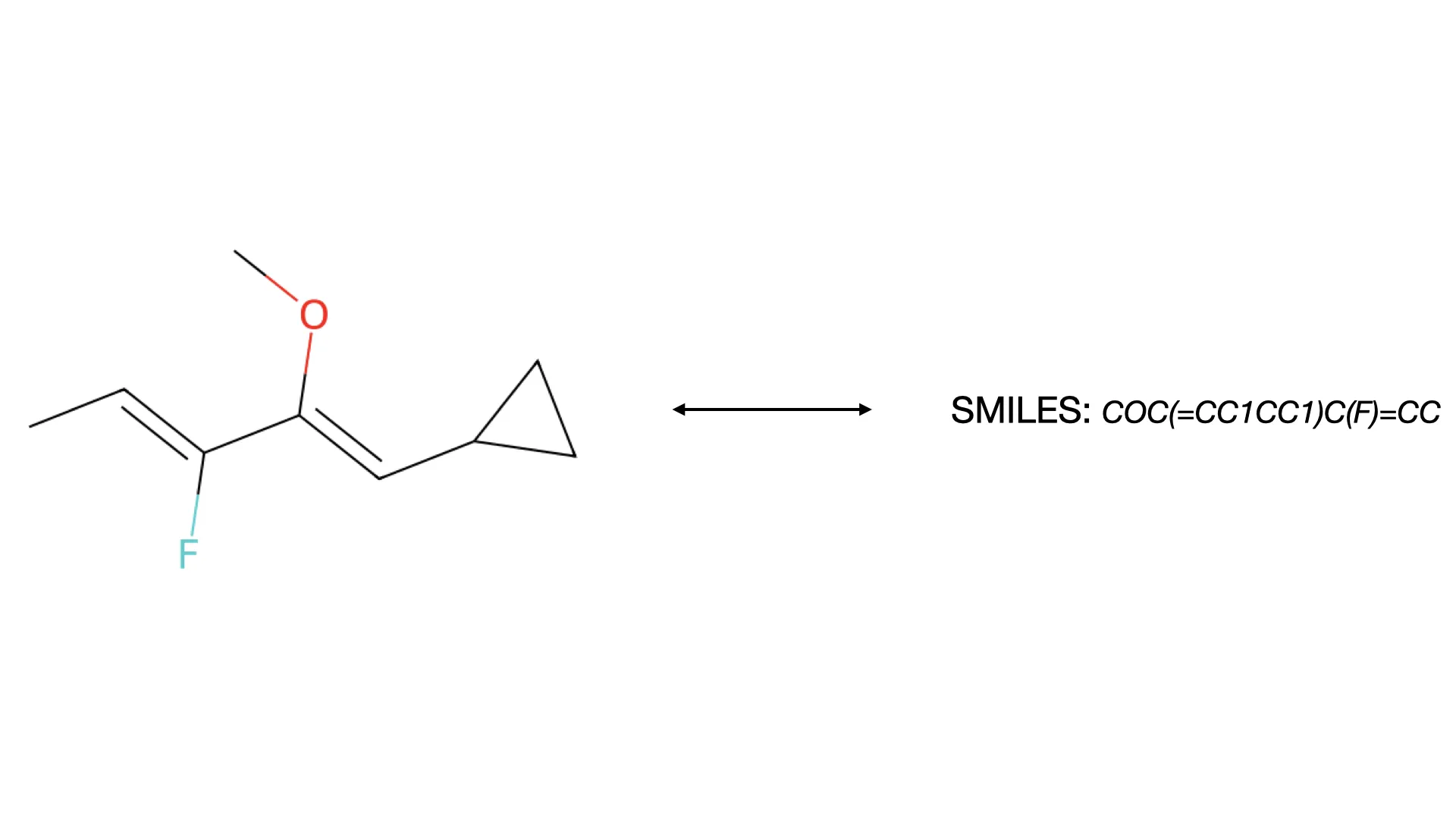
A 2021 image-to-text approach treating OCSR as an image captioning task. It uses Transformers with SELFIES representation to convert molecular structure diagrams into SMILES strings, enabling extraction of visual chemical knowledge from scientific literature.