Translating InChI to IUPAC Names with Transformers
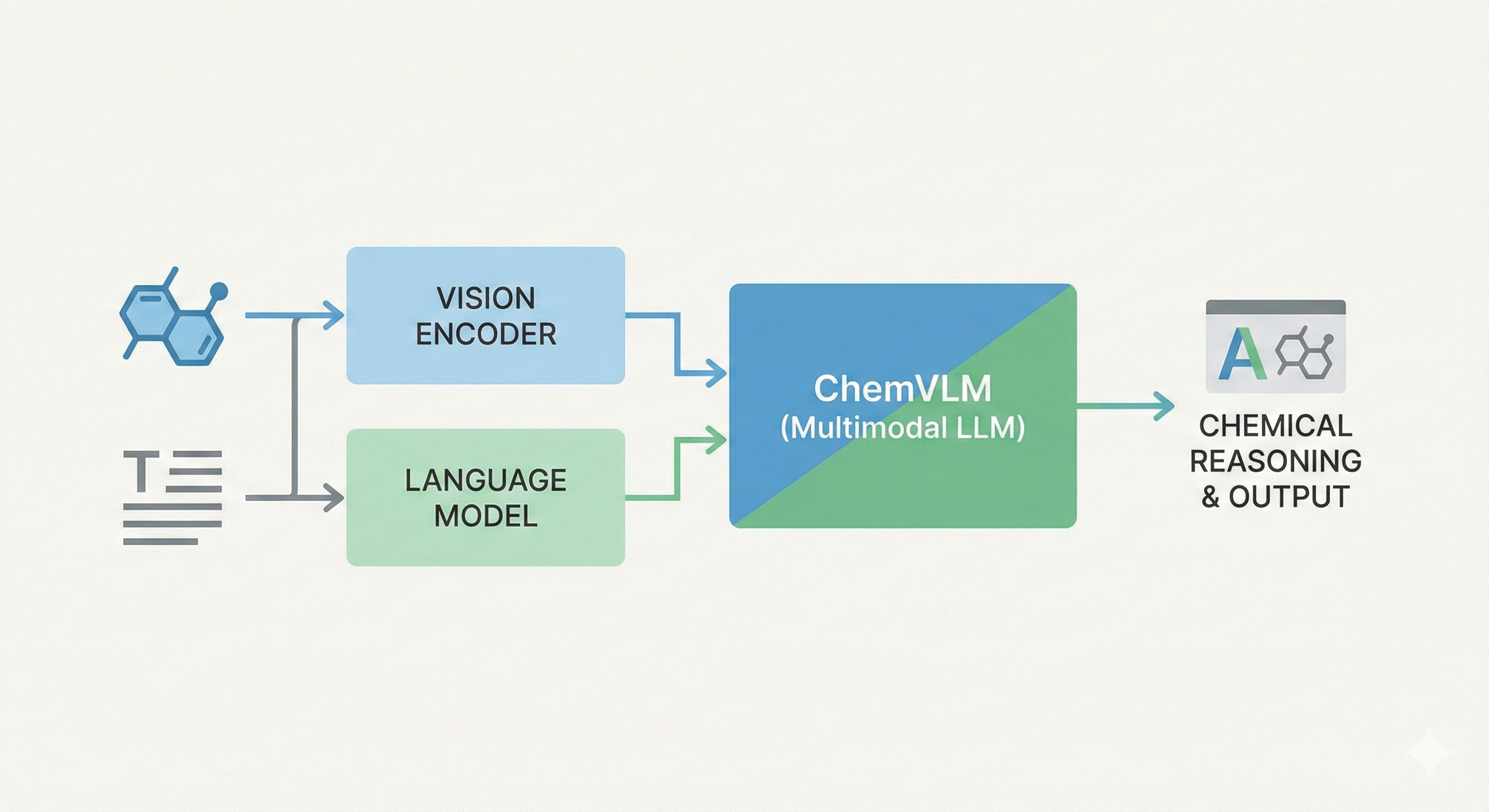
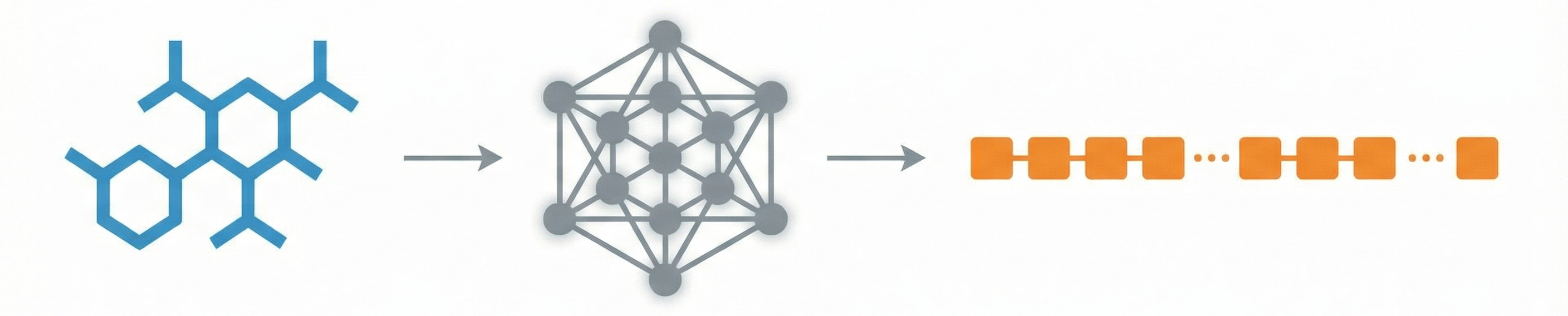
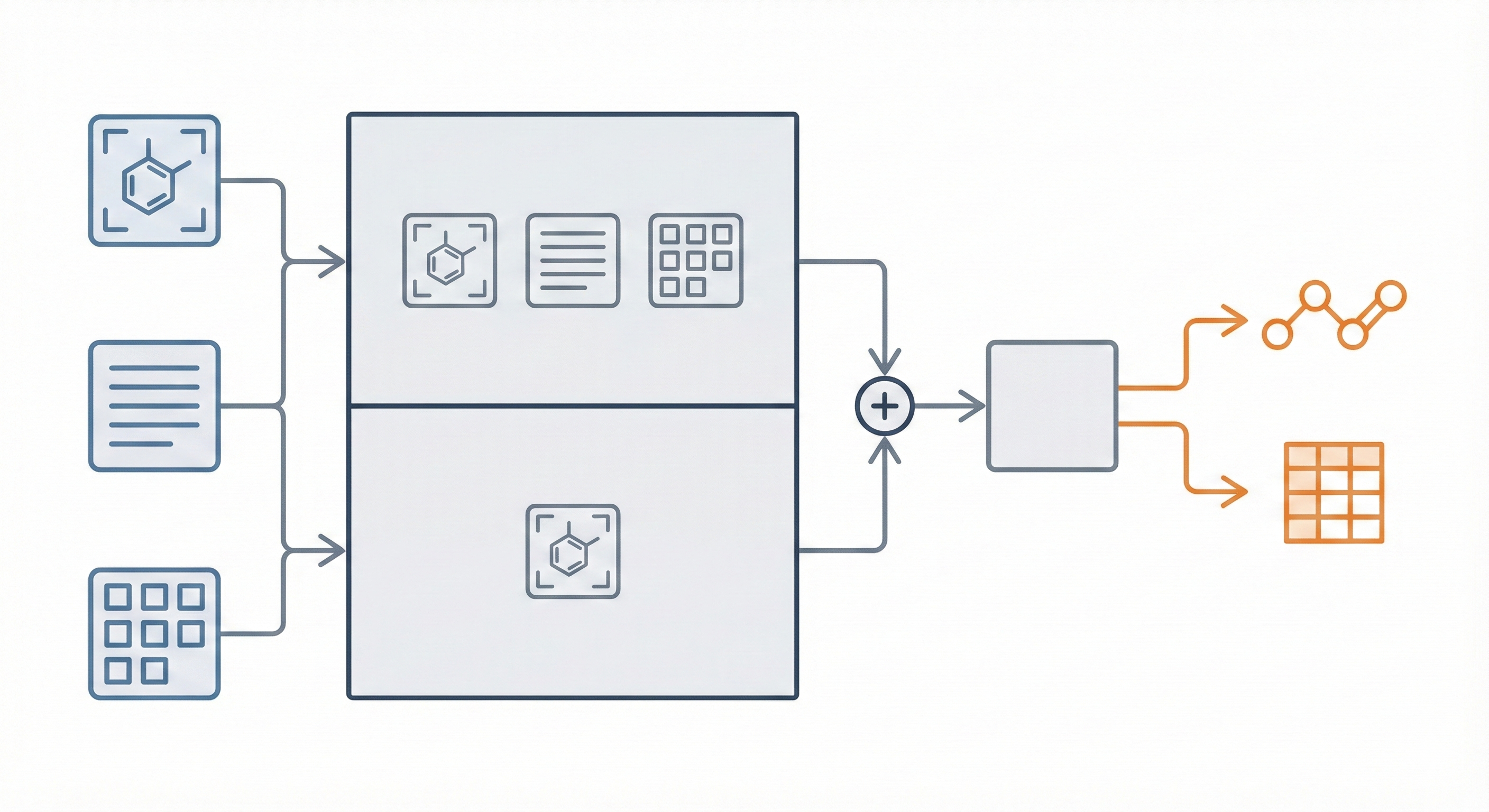
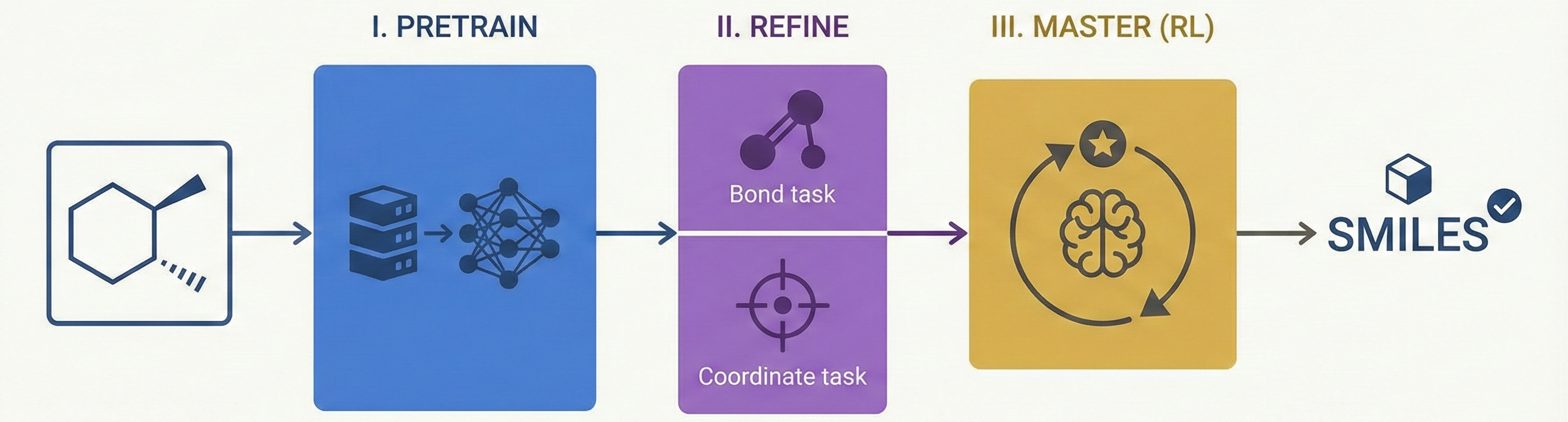
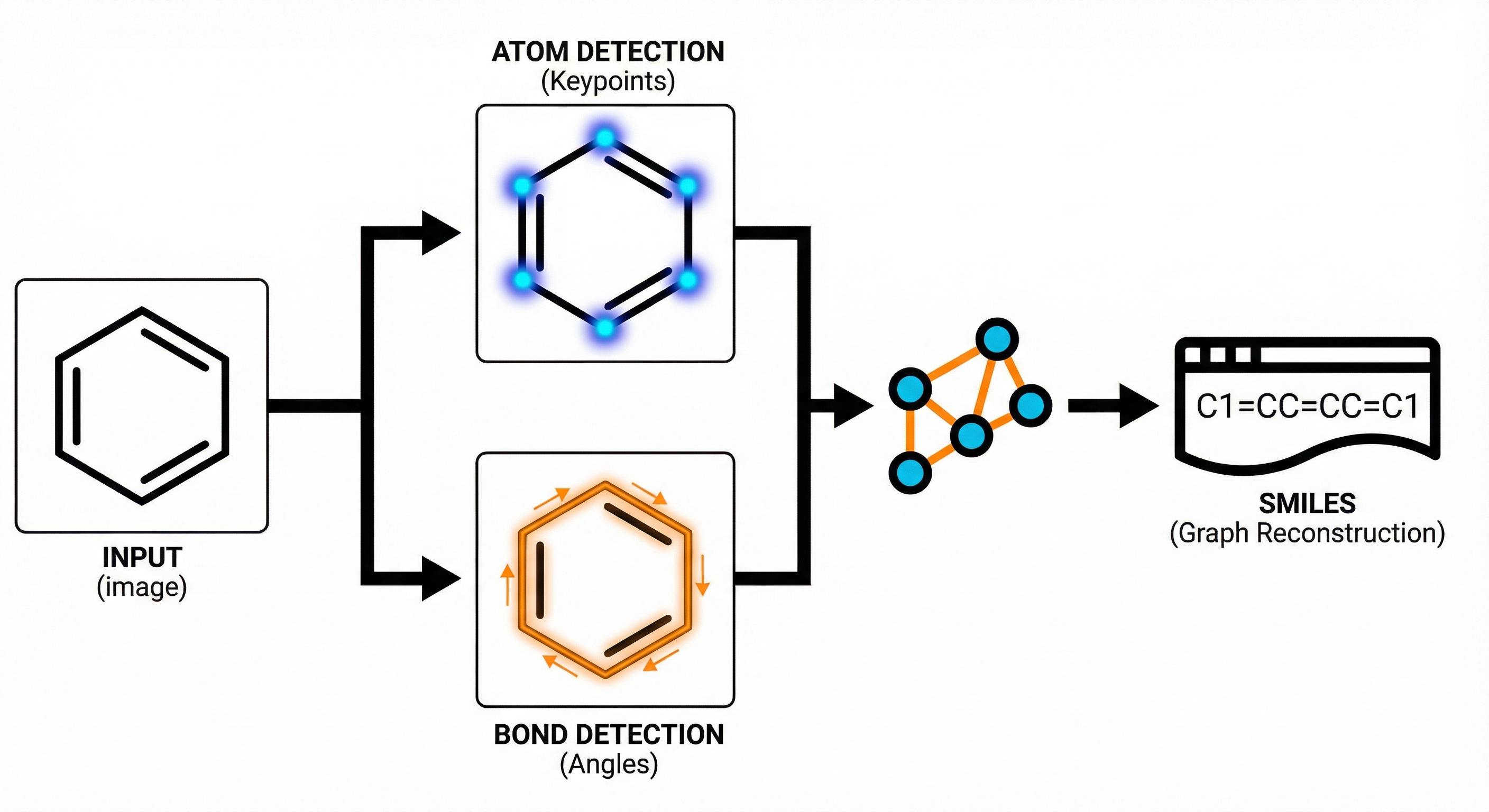
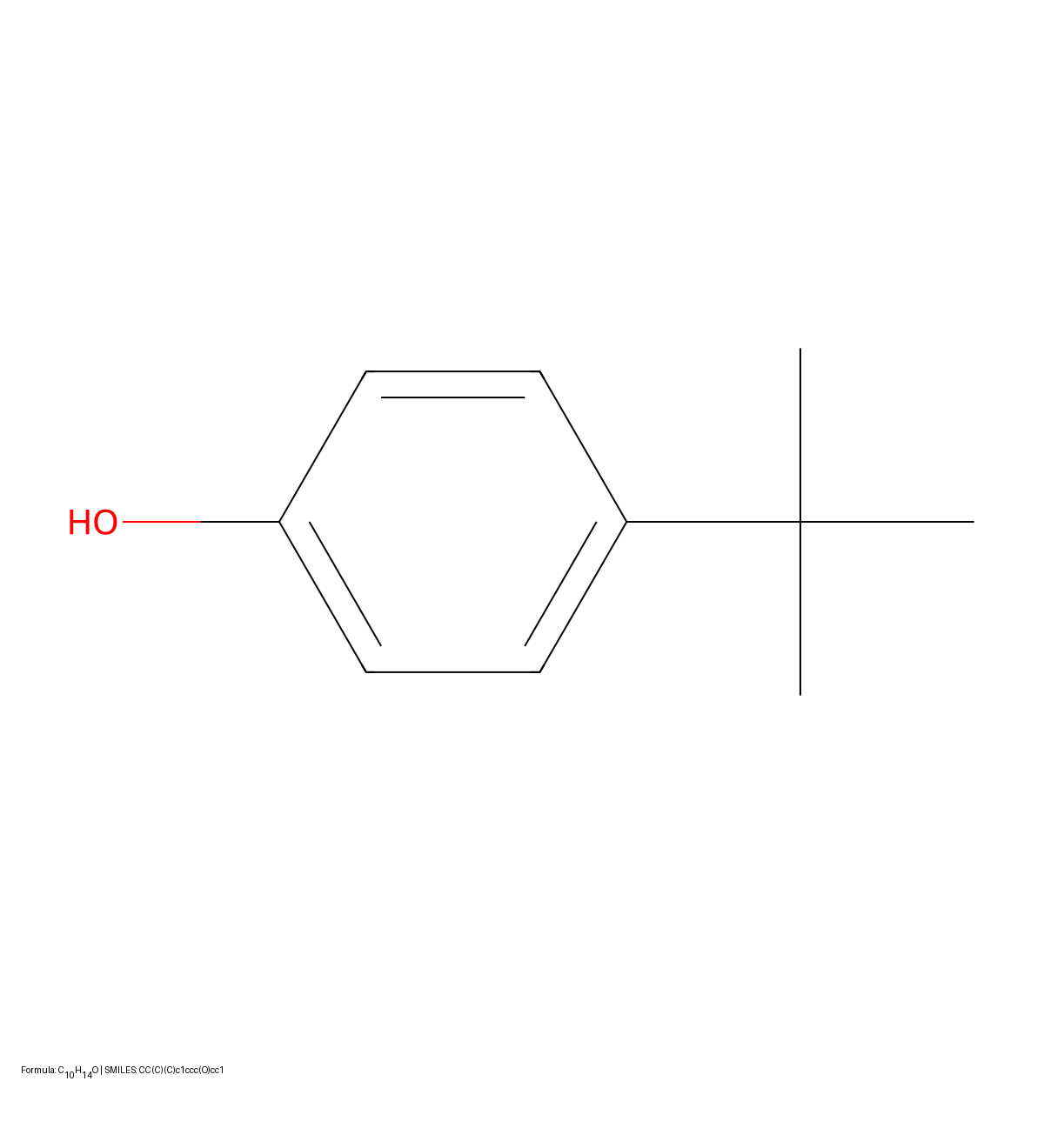
This study presents a sequence-to-sequence Transformer model that translates InChI identifiers into IUPAC names character-by-character. Trained on 10 million PubChem pairs, it achieves 91% accuracy on organic compounds, performing comparably to commercial software.