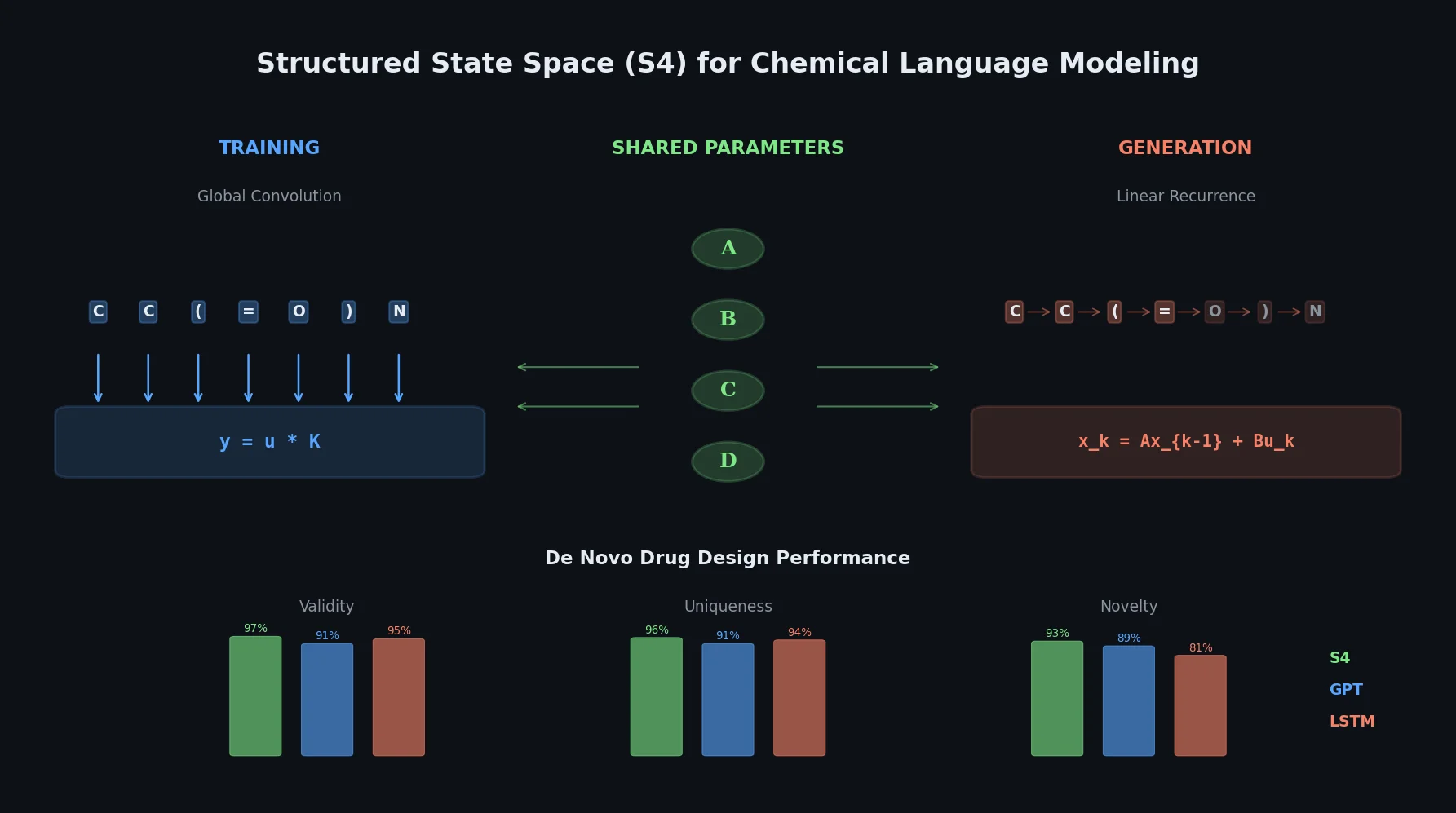
S4 Structured State Space Models for De Novo Drug Design
This paper introduces structured state space sequence (S4) models to chemical language modeling, showing they combine the strengths of LSTMs (efficient recurrent generation) and GPTs (holistic sequence learning) for de novo molecular design.