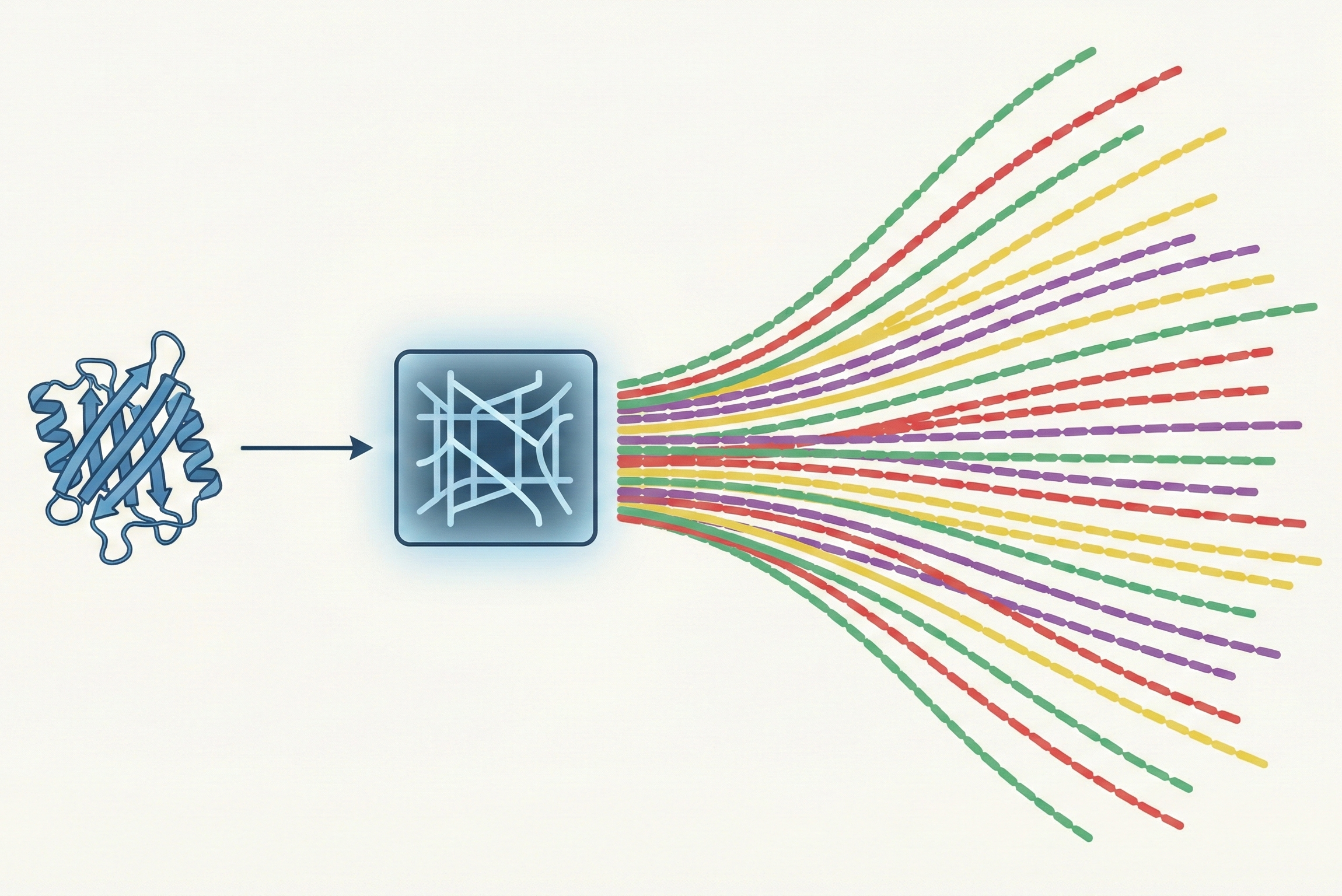
InvMSAFold: Generative Inverse Folding with Potts Models
InvMSAFold replaces autoregressive decoding with a Potts model parameter generator, enabling diverse protein sequence sampling orders of magnitude faster than ESM-IF1.

InvMSAFold replaces autoregressive decoding with a Potts model parameter generator, enabling diverse protein sequence sampling orders of magnitude faster than ESM-IF1.
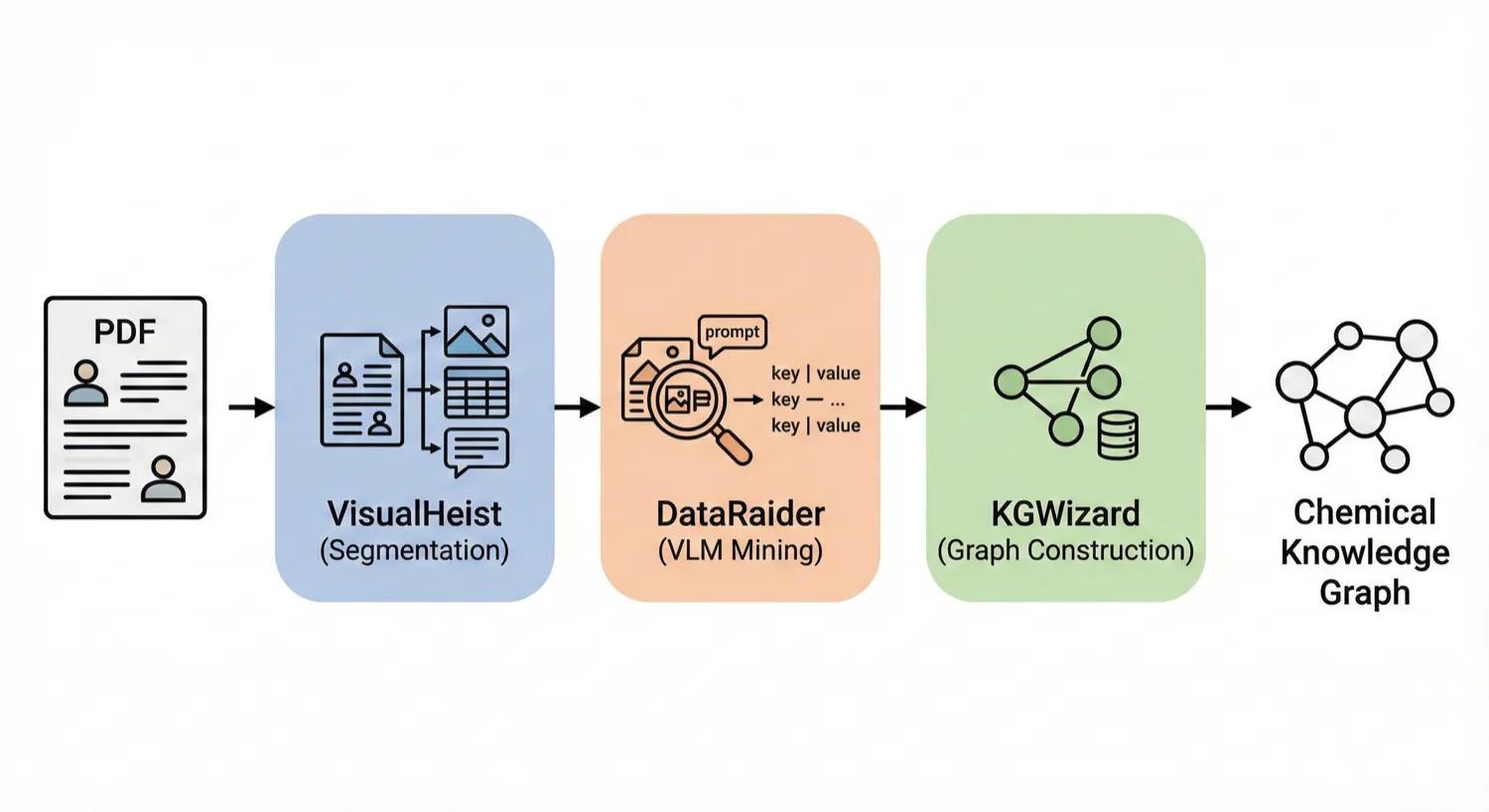
MERMaid leverages fine-tuned vision models and VLM reasoning to mine chemical reaction data directly from PDF figures and tables. By handling context inference and coreference resolution, it builds high-fidelity knowledge graphs with 87% end-to-end accuracy.
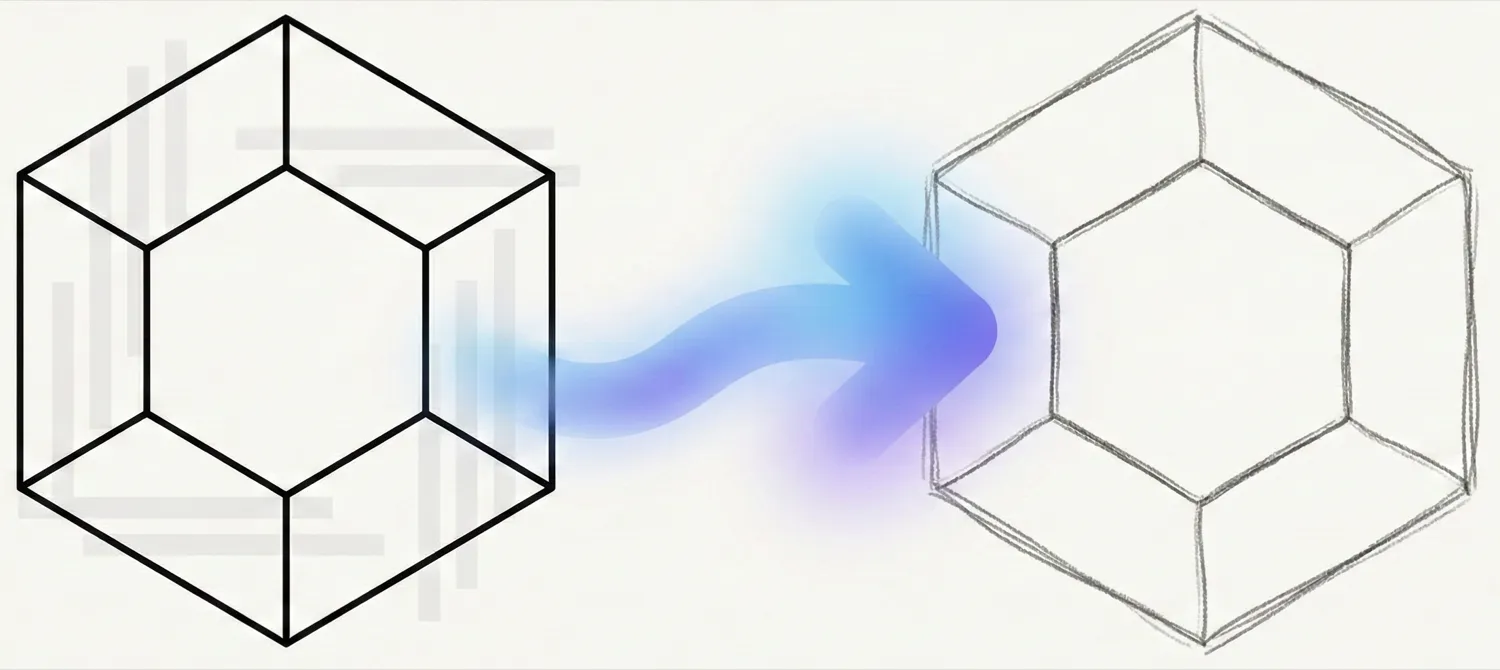
OCSAug uses Denoising Diffusion Probabilistic Models (DDPM) and the RePaint algorithm with custom masking to generate synthetic hand-drawn chemical structure images, improving OCSR performance by 1.918-3.820x on the DECIMER benchmark.
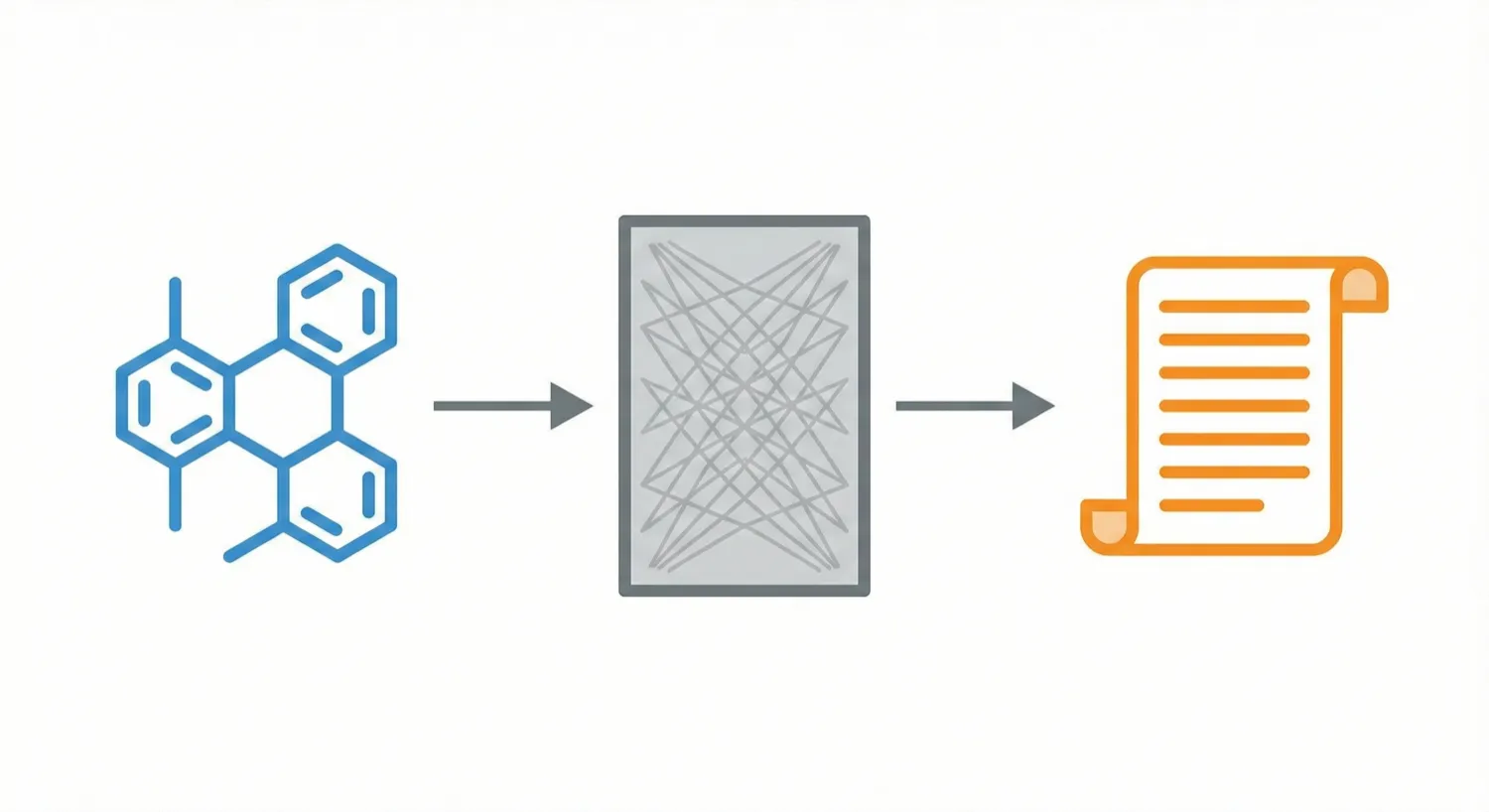
STOUT V2.0 uses Transformers trained on ~1 billion SMILES-IUPAC pairs to accurately translate chemical structures into systematic names (and vice-versa), outperforming its RNN predecessor.
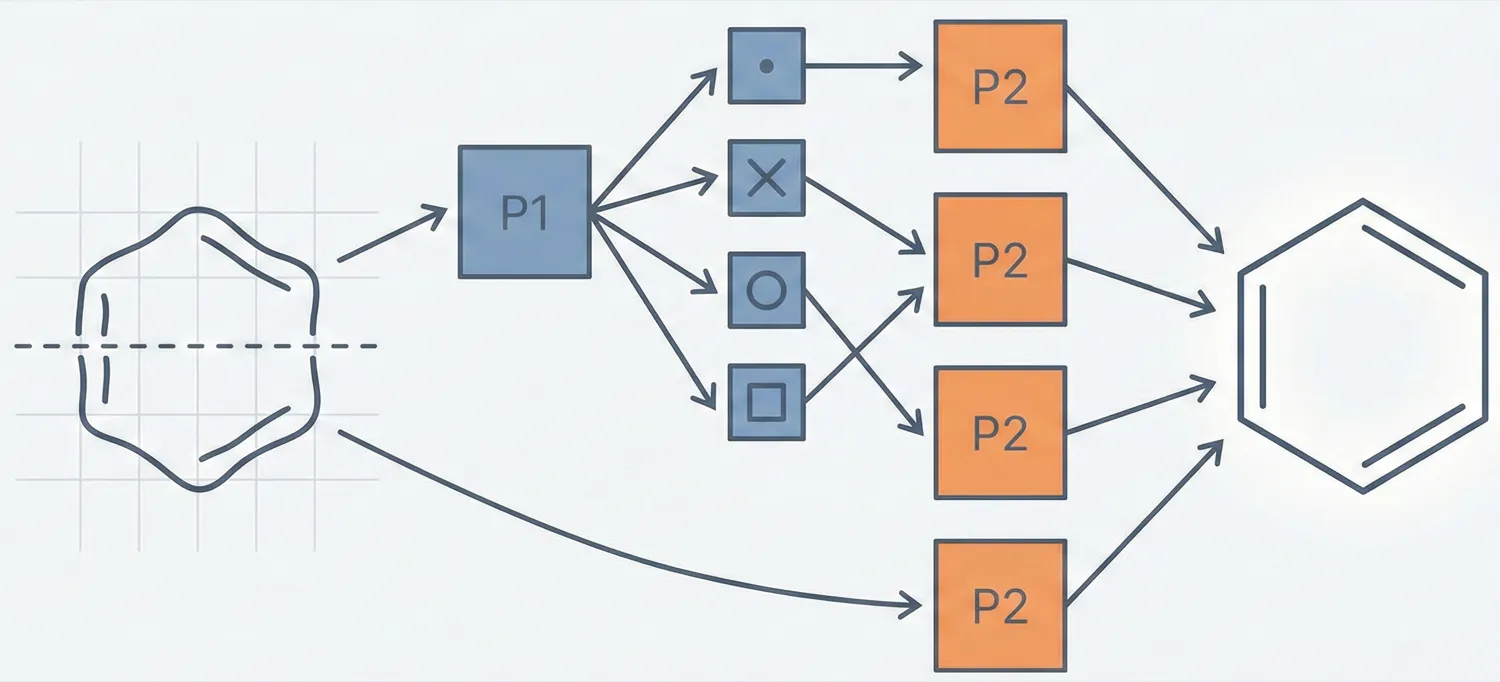
Proposes a specialized Classifier-Recognizer architecture that first categorizes rings by heteroatom (S, N, O) and then identifies the specific ring using optimized grid inputs.
A 2013 paper introducing a hybrid recognition system for handwritten chemical symbols on touch devices. Combines Support Vector Machines (SVM) for classification with elastic matching for geometric verification, achieving 89.7% top-1 accuracy on pen-based input for chemical structure drawing applications.
HMM-based method for recognizing online handwritten chemical symbols using 11-dimensional local features including derivatives, curvature, and linearity. Achieves 89.5% top-1 accuracy and 98.7% top-3 accuracy on a custom dataset of 64 chemical symbols.
Yang et al. propose a two-level recognition system for handwritten chemical formulas, combining global structural analysis to identify substances with local character recognition using ANNs, achieving ~96% accuracy on a dataset of 1197 expressions.
A three-level grammatical framework (formula, molecule, text) for parsing online handwritten chemical formulas, generating semantic graphs that capture both connectivity and layout using context-free grammars and HMMs.
Proposes a novel two-level algorithm for on-line handwritten chemical expression recognition, combining substance-level matching with character-level segmentation to achieve 96% accuracy.
This paper proposes a double-stage architecture using SVM for rough classification and HMM for fine recognition. It features a novel Point Sequence Reordering (PSR) algorithm that significantly improves accuracy on organic ring structures.
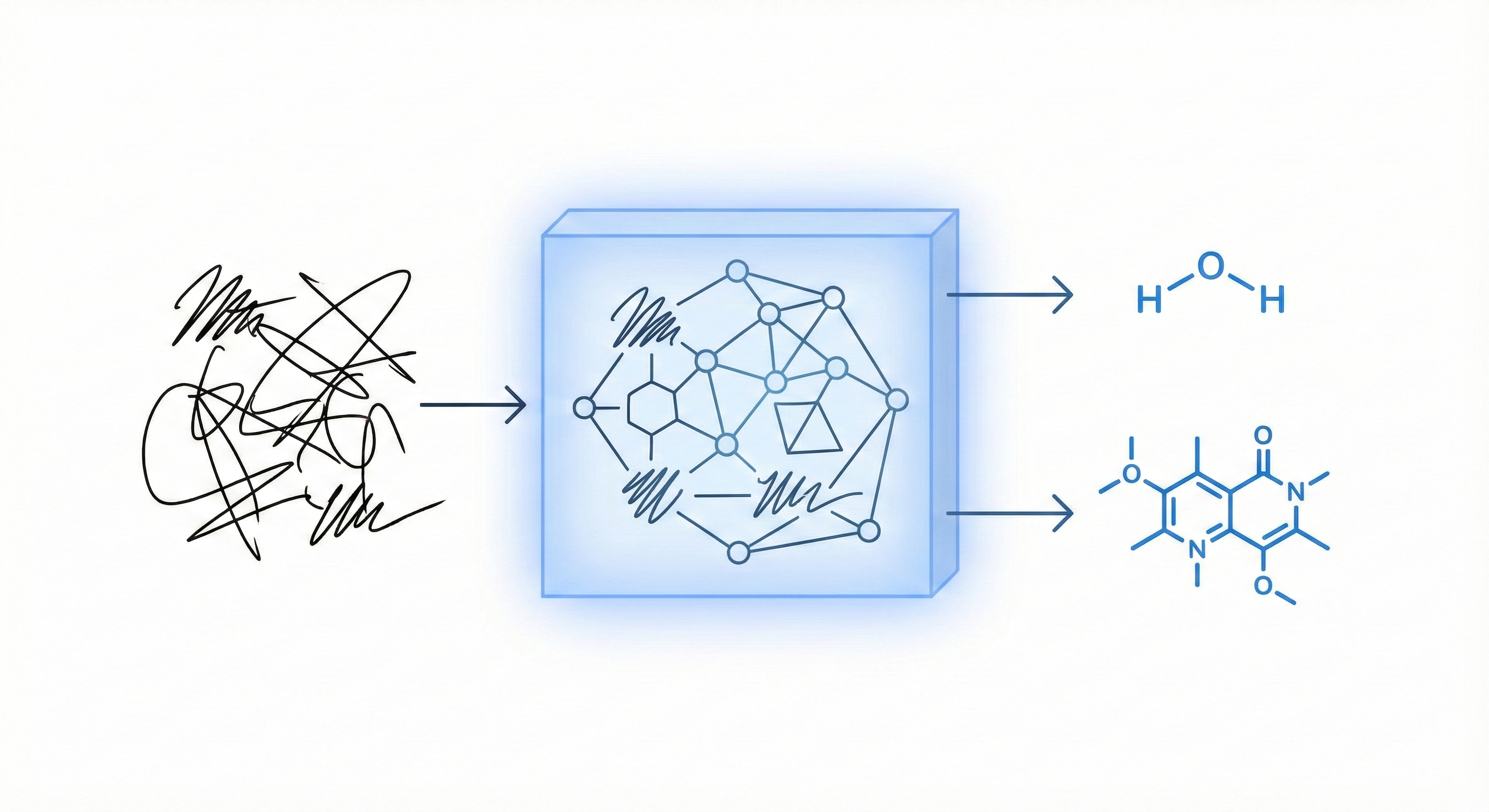
Proposes a unified statistical framework for recognizing both inorganic and organic handwritten chemical expressions. Introduces the Chemical Expression Structure Graph (CESG) and uses a weighted direction graph search for structural analysis, achieving 83.1% top-5 accuracy on a large proprietary dataset.