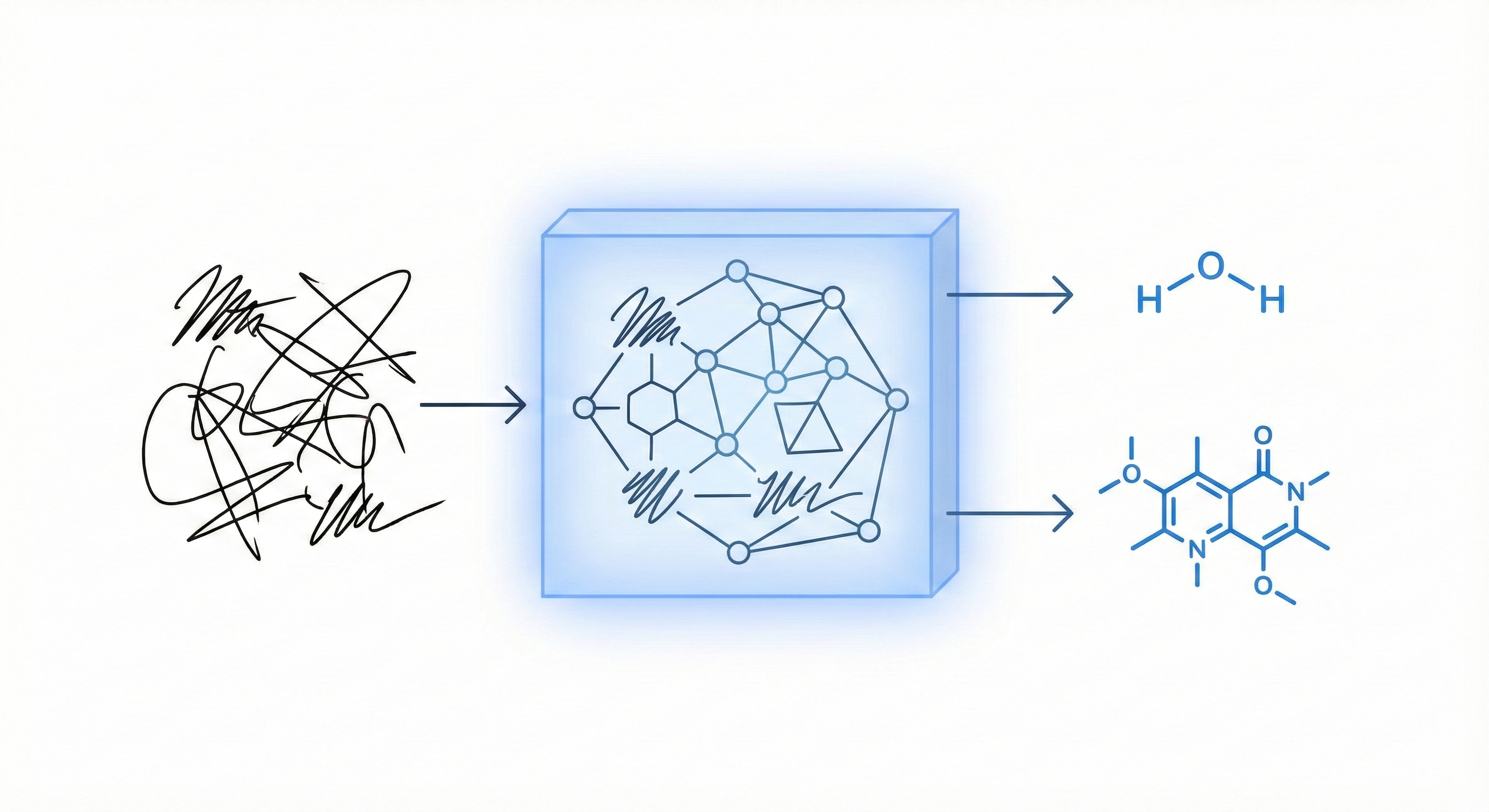
Unified Framework for Handwritten Chemical Expressions
Proposes a unified statistical framework for recognizing both inorganic and organic handwritten chemical expressions. Introduces the Chemical Expression Structure Graph (CESG) and uses a weighted direction graph search for structural analysis, achieving 83.1% top-5 accuracy on a large proprietary dataset.