Survey of Transformer Architectures in Molecular Science
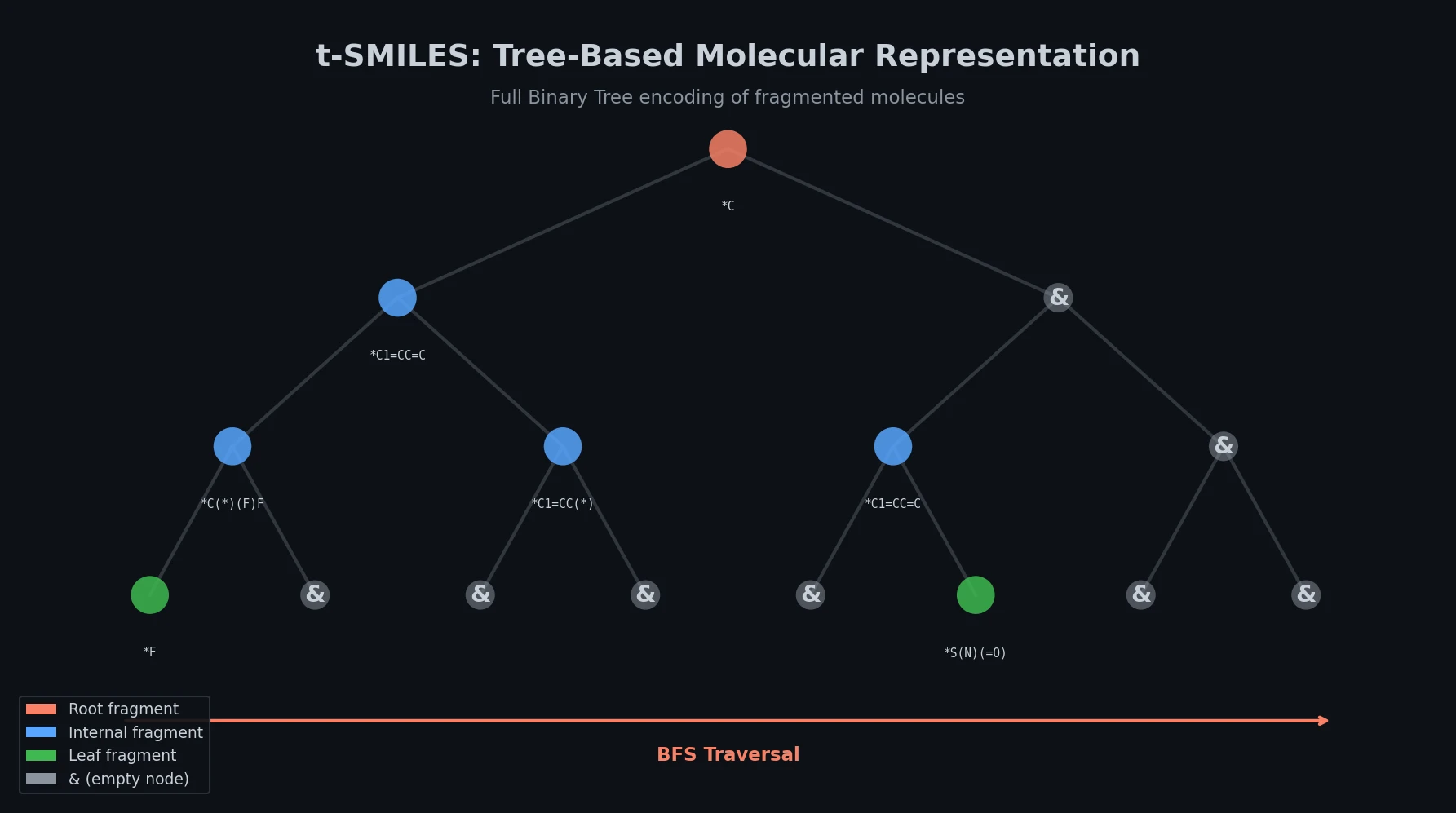
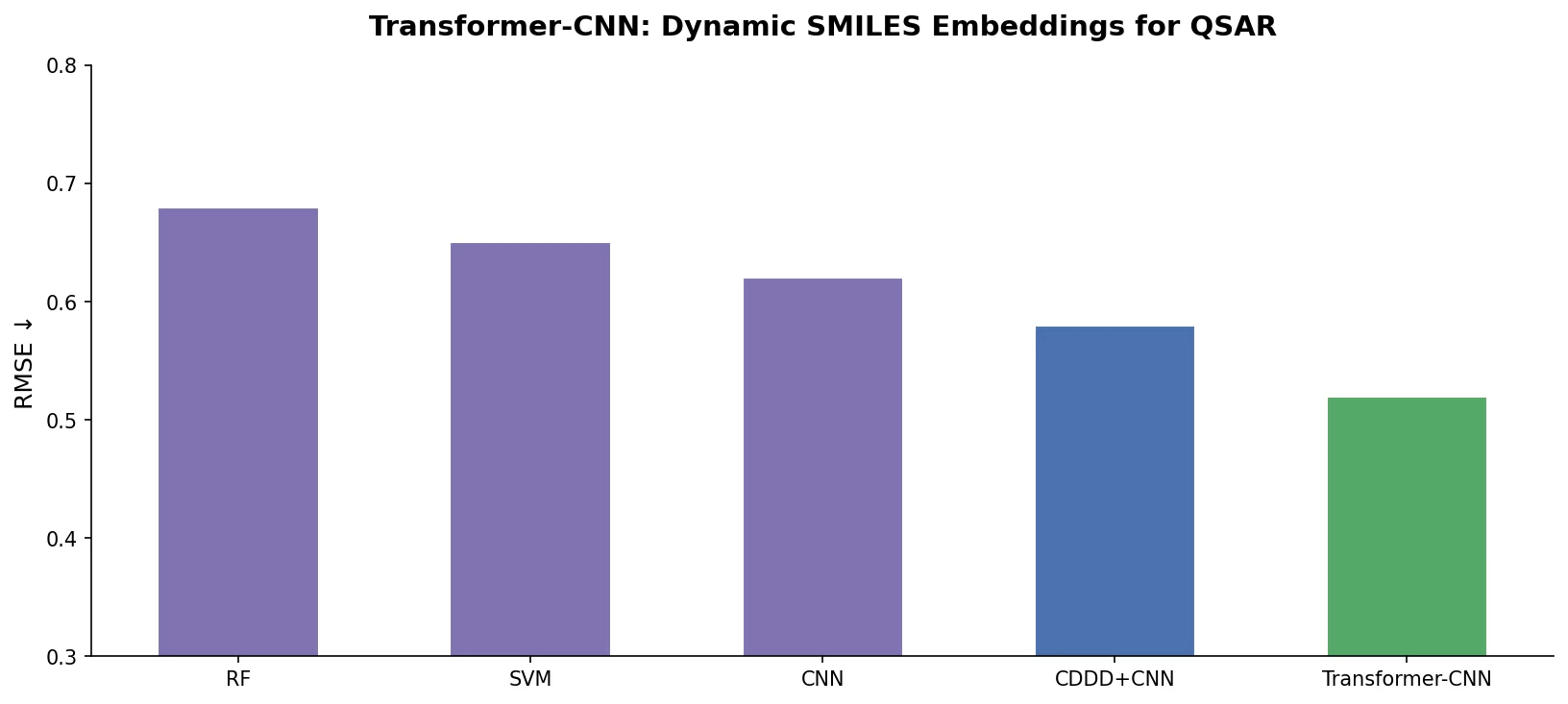
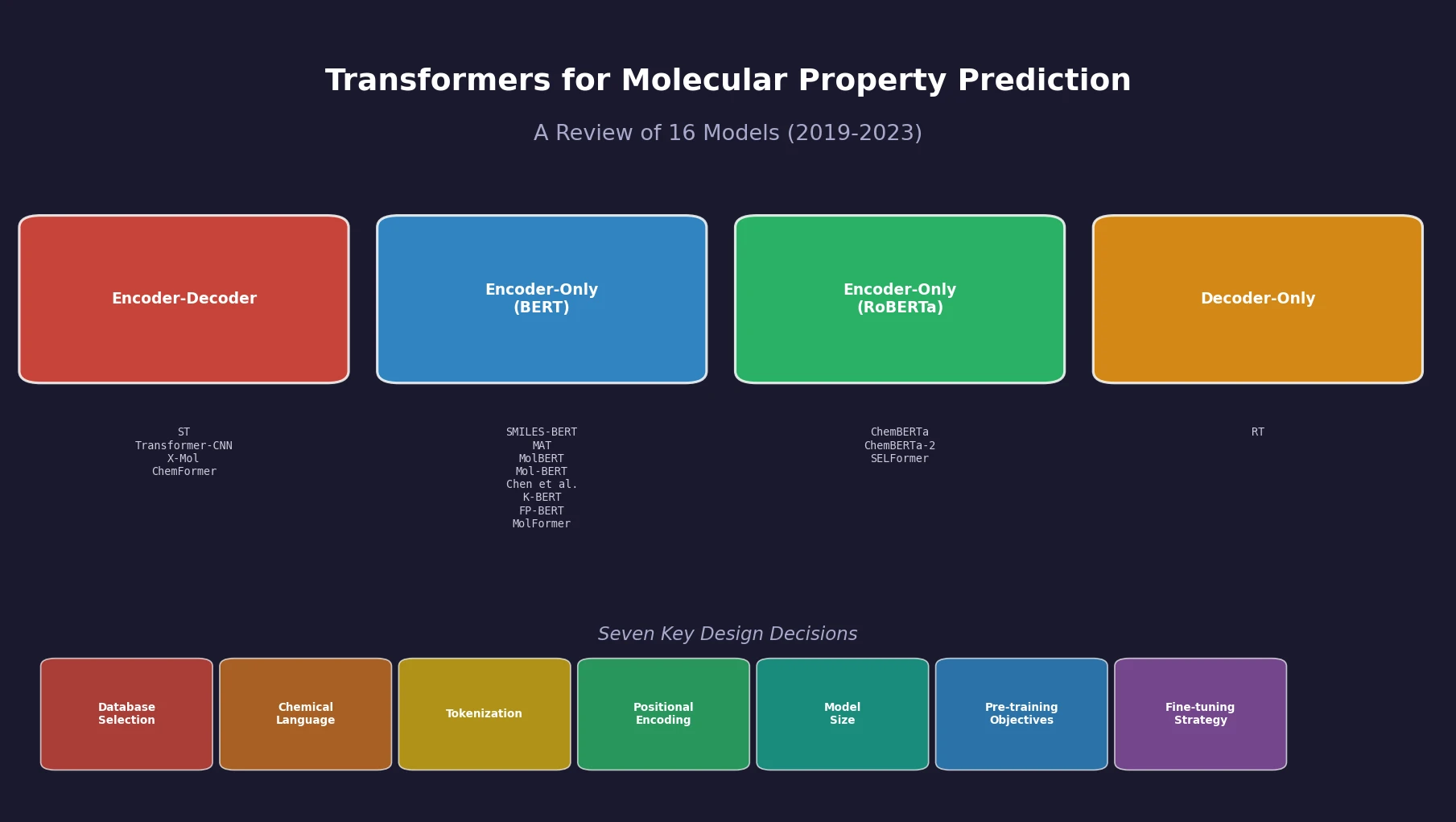
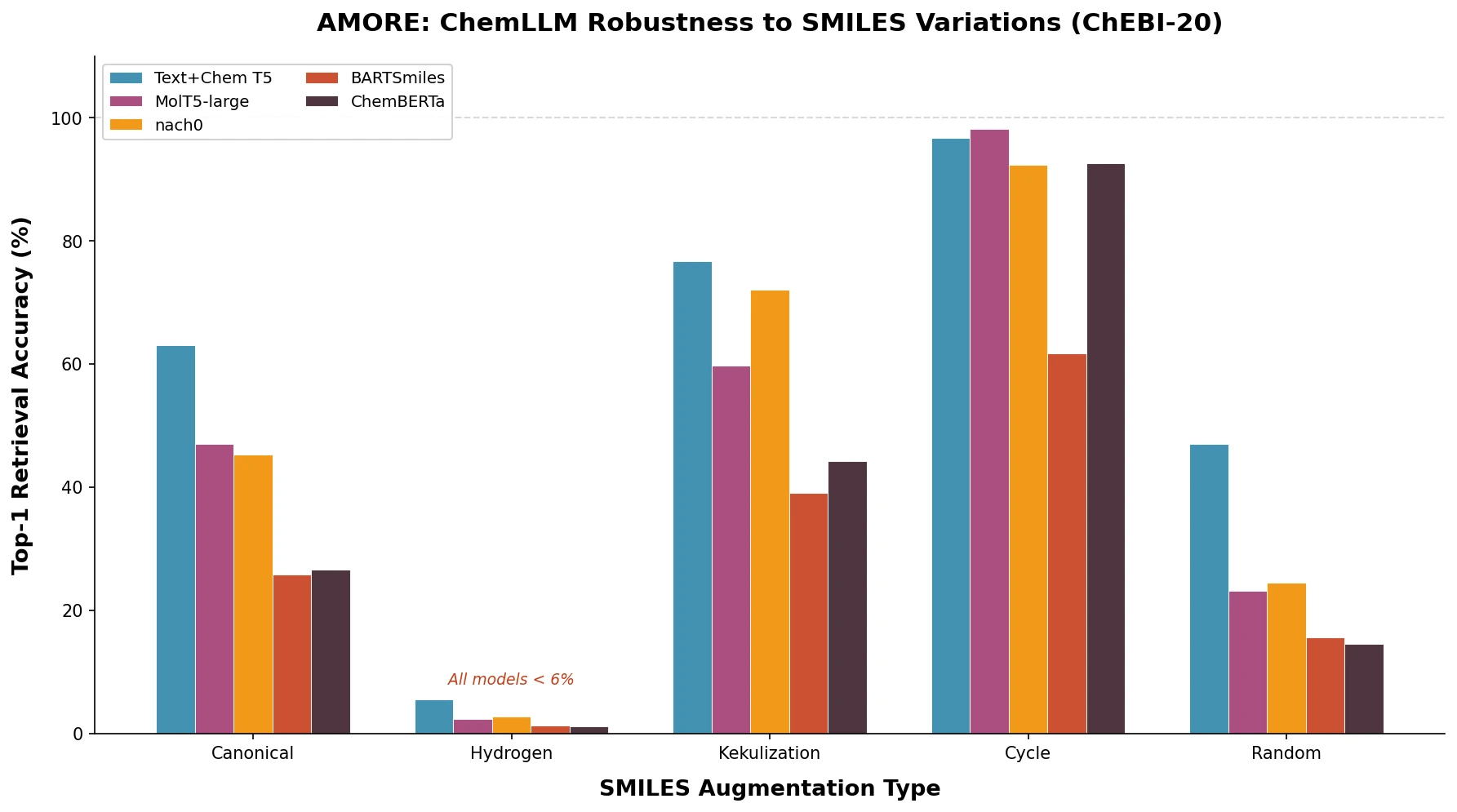
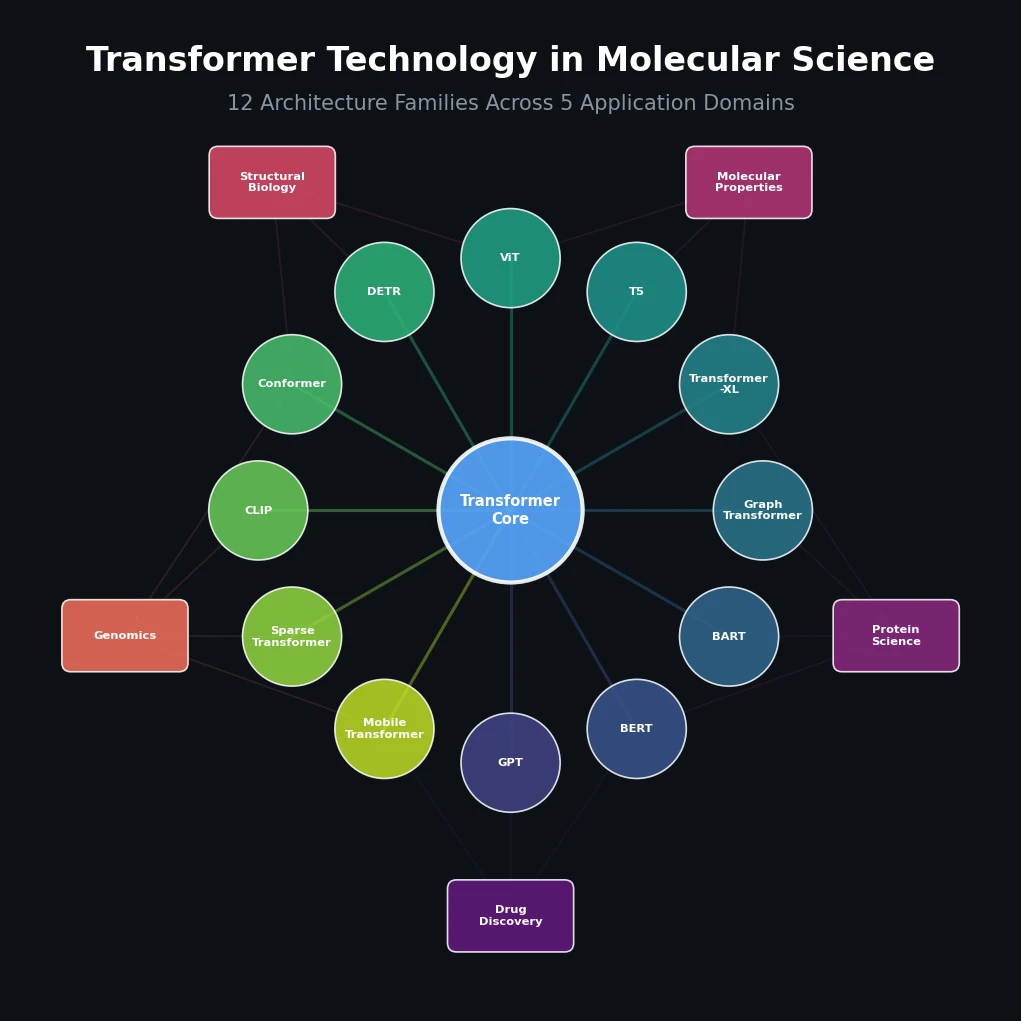
Jiang et al. survey 12 families of transformer architectures in molecular science, covering GPT, BERT, BART, graph transformers, Transformer-XL, T5, ViT, DETR, Conformer, CLIP, sparse transformers, and mobile/efficient variants, with detailed algorithmic descriptions and molecular applications.