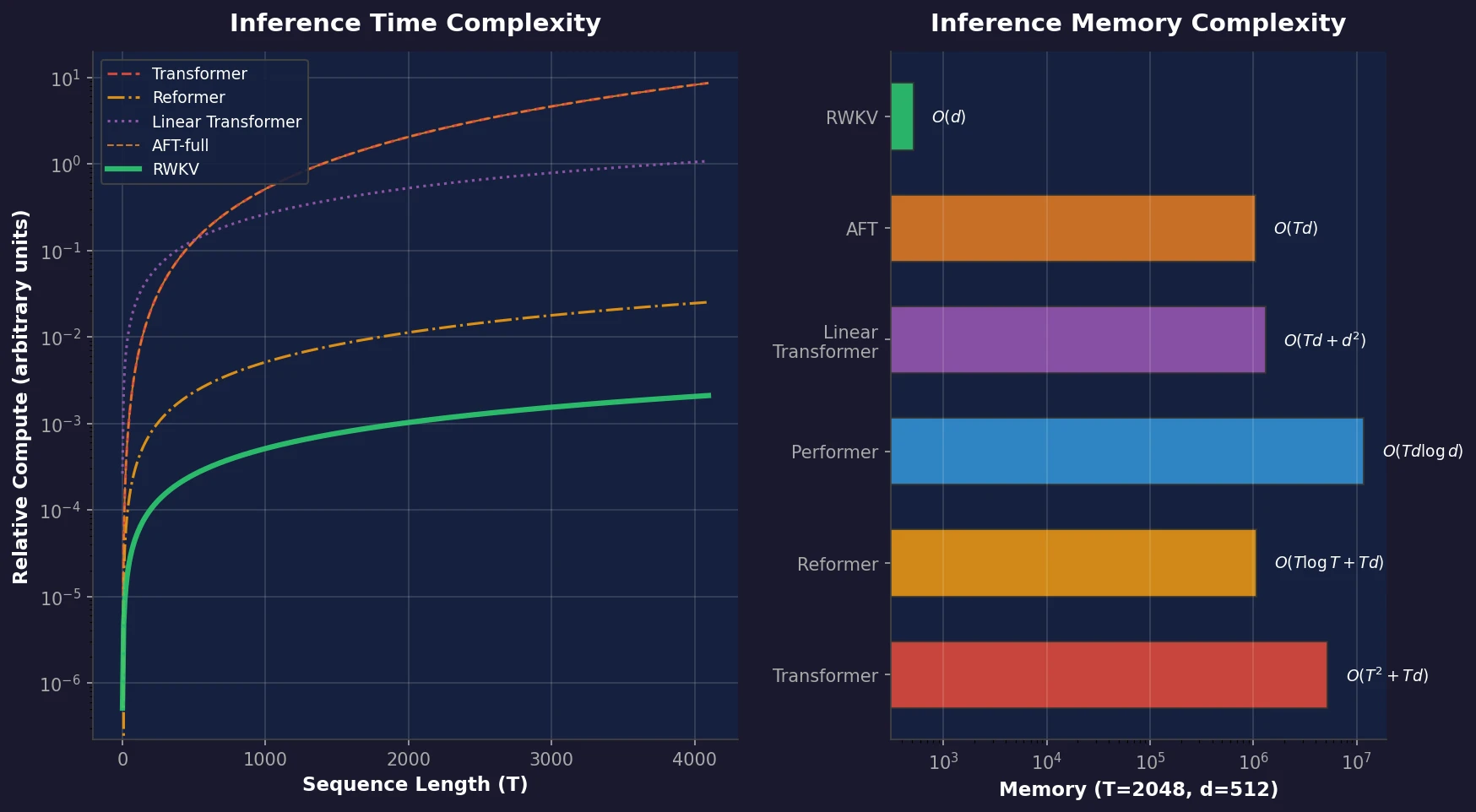
RWKV: Linear-Cost RNN with Transformer Training
RWKV is a novel sequence model that achieves transformer-level performance while maintaining linear time and constant memory complexity during inference, scaled up to 14 billion parameters.

RWKV is a novel sequence model that achieves transformer-level performance while maintaining linear time and constant memory complexity during inference, scaled up to 14 billion parameters.
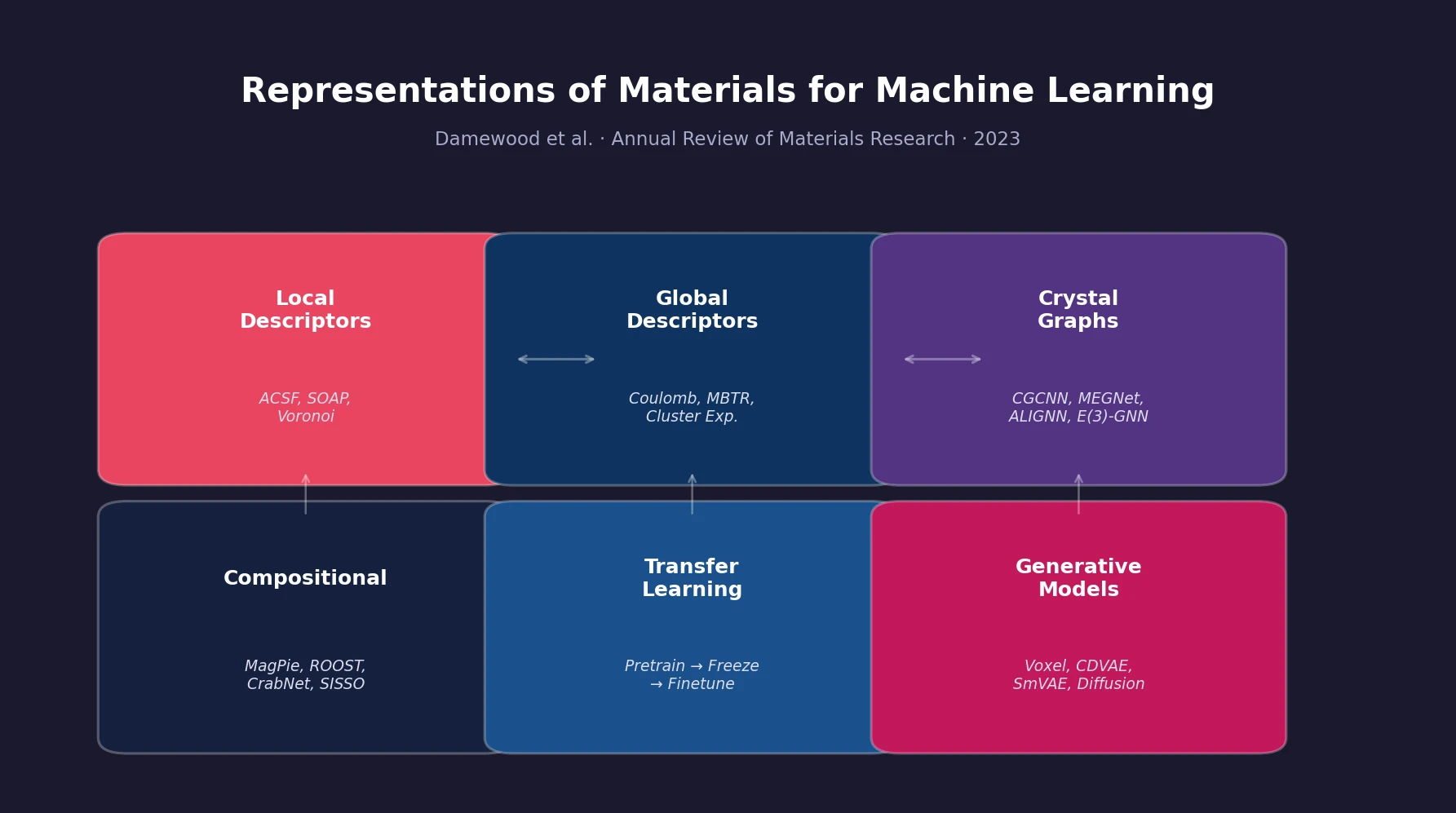
A comprehensive review of how solid-state materials can be numerically represented for machine learning, spanning structural features, graph neural networks, compositional descriptors, transfer learning, and generative models for inverse design.
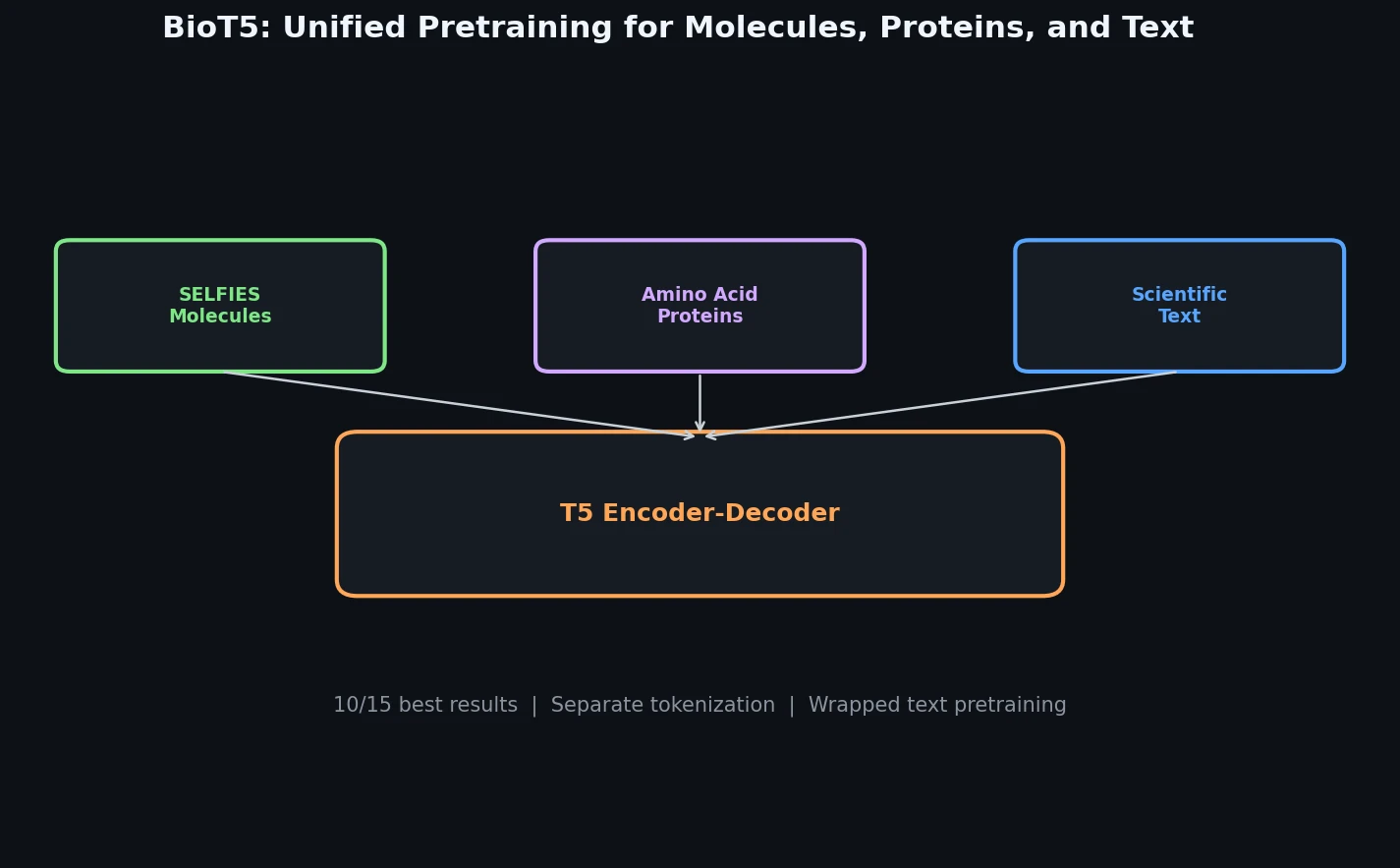
BioT5 uses SELFIES representations and separate tokenization to pre-train a unified T5 model across molecules, proteins, and text, achieving state-of-the-art results on 10 of 15 downstream tasks.

ChatDrug is a parameter-free framework that combines ChatGPT with retrieval-augmented domain feedback and iterative conversation to edit drugs across small molecules, peptides, and proteins.
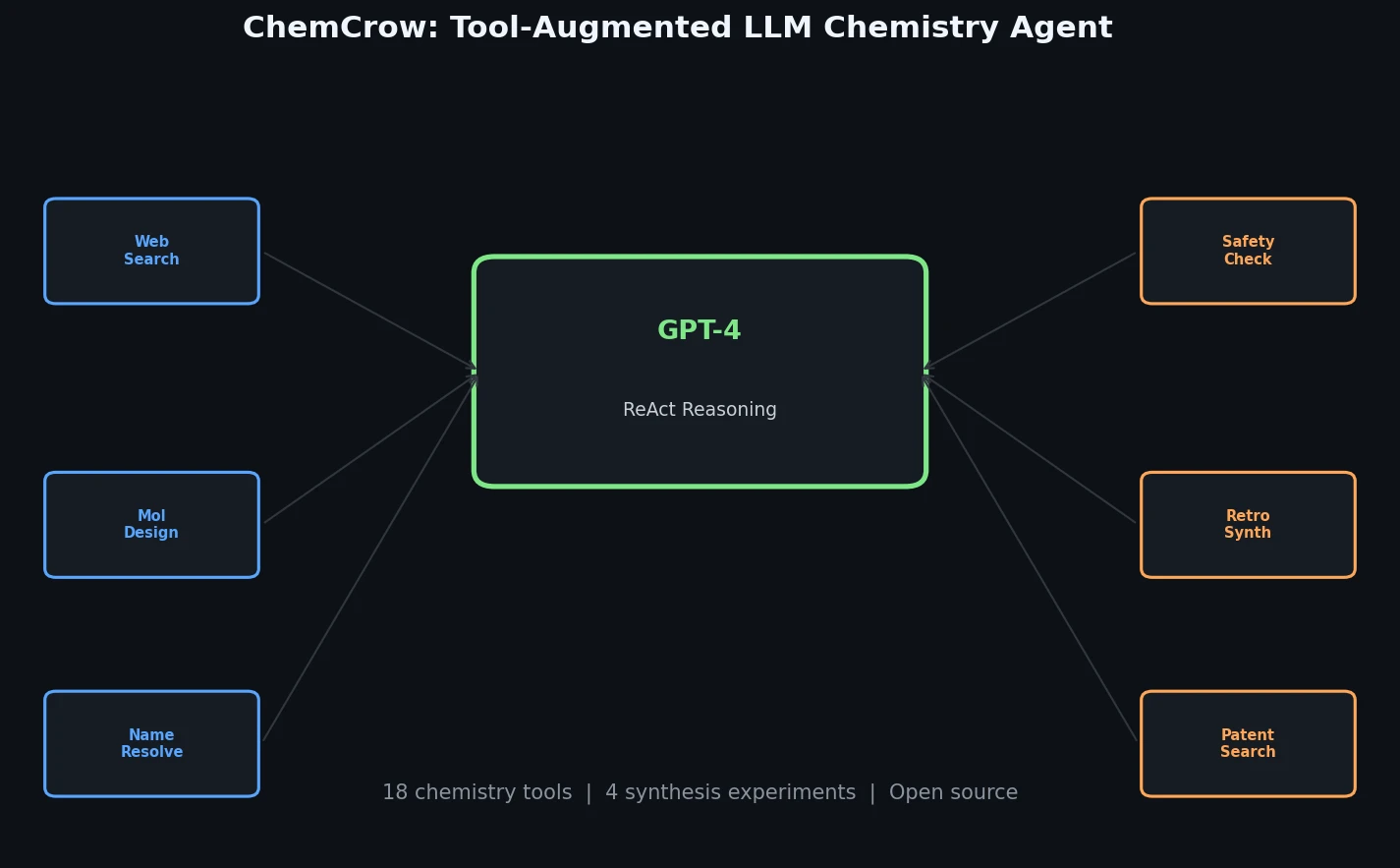
ChemCrow augments GPT-4 with 18 chemistry tools to autonomously plan and execute syntheses, discover novel chromophores, and solve diverse chemical reasoning tasks.
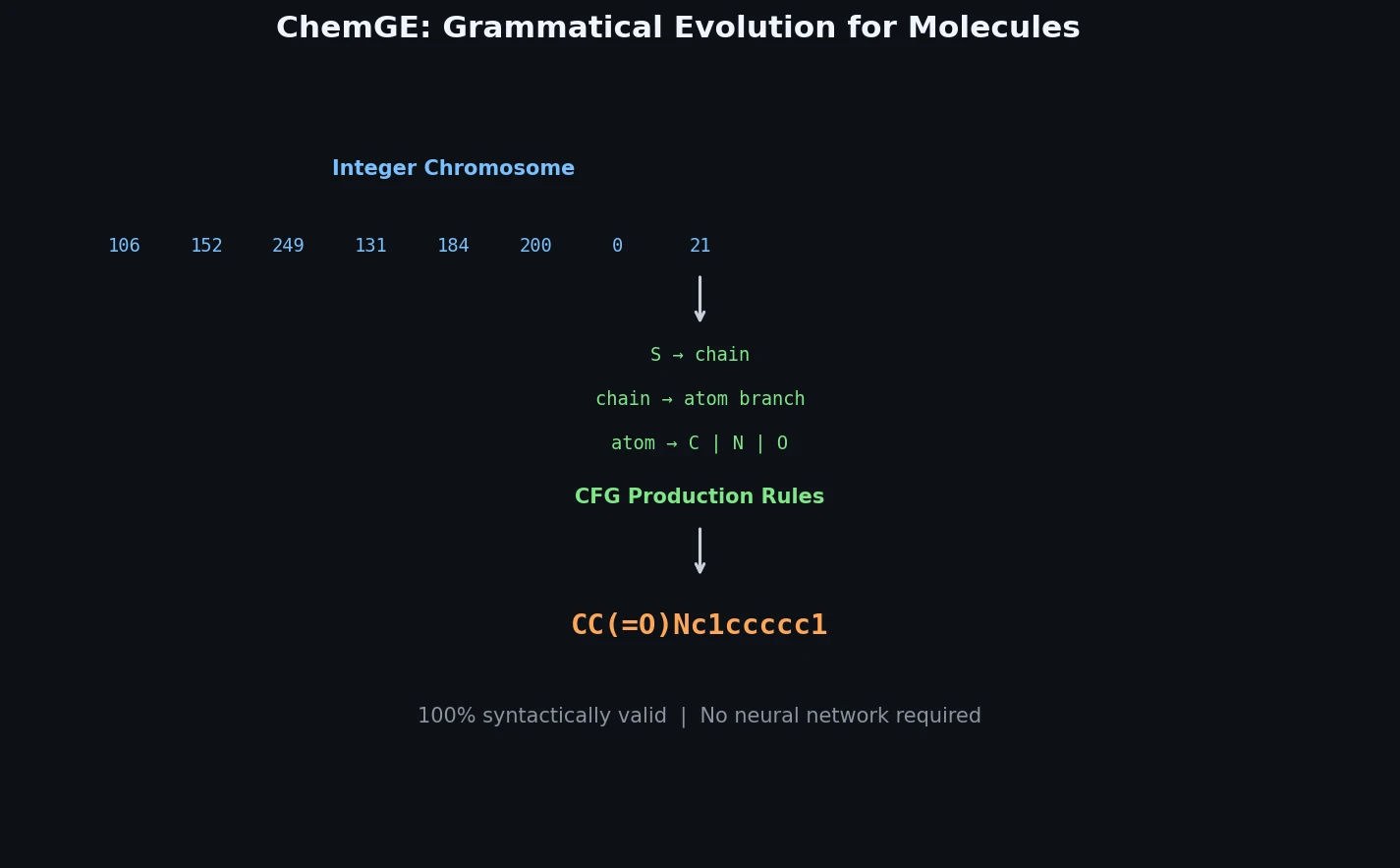
ChemGE uses grammatical evolution over SMILES context-free grammars to generate diverse drug-like molecules in parallel, outperforming deep learning baselines in throughput and molecular diversity.
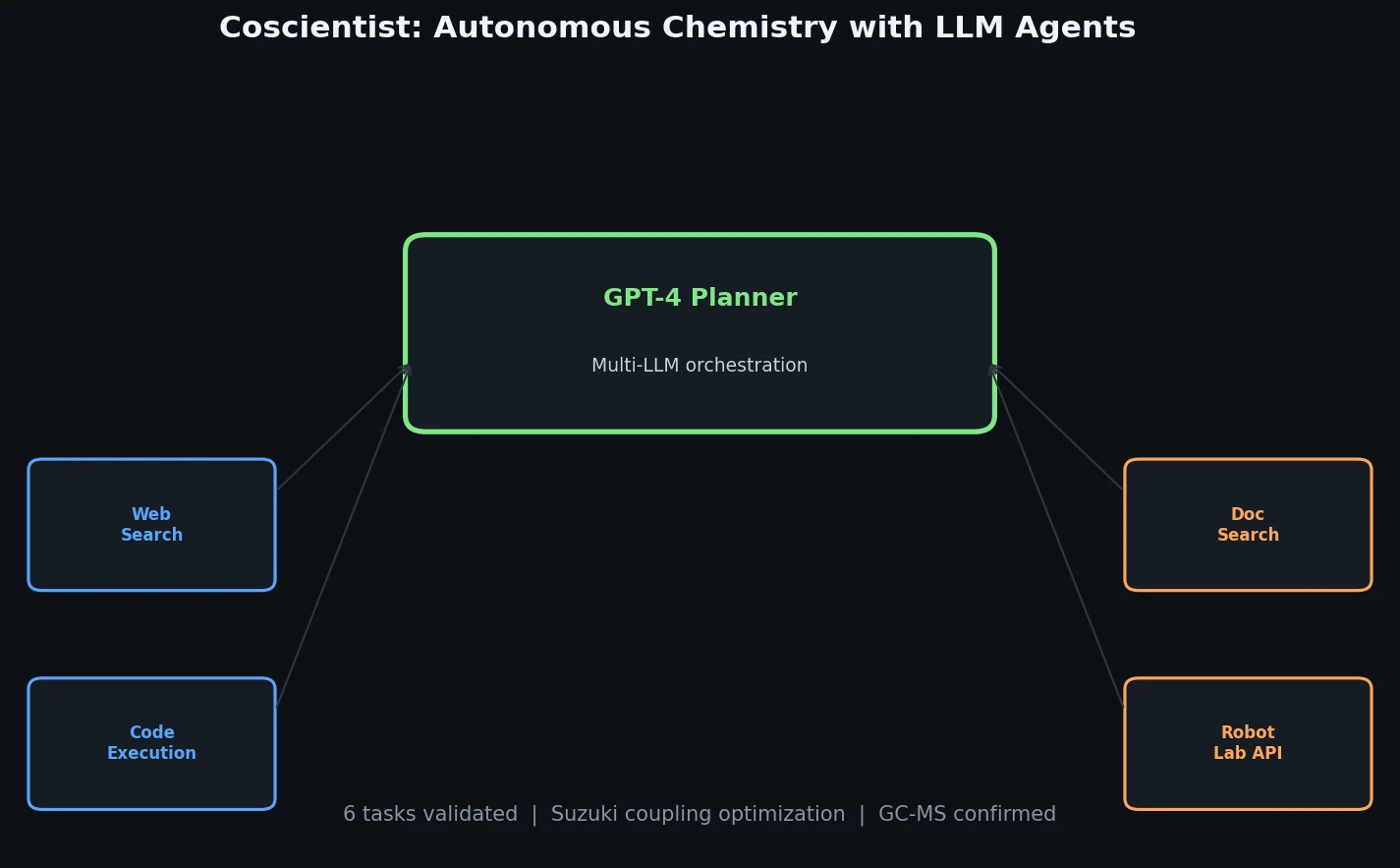
Introduces Coscientist, a GPT-4-driven AI system that autonomously designs and executes chemical experiments using web search, code execution, and robotic lab automation.
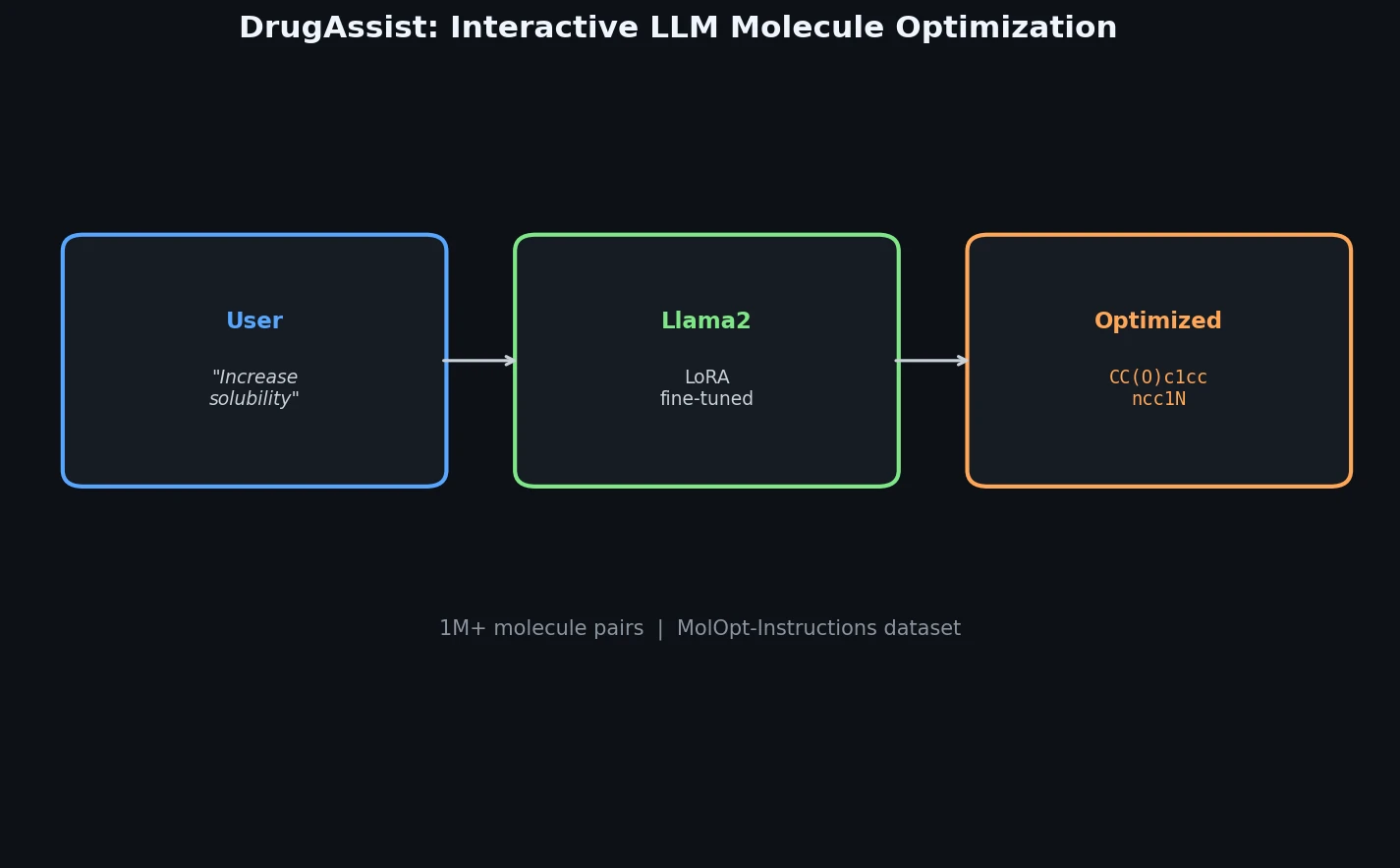
DrugAssist fine-tunes Llama2-7B-Chat on over one million molecule pairs for interactive, dialogue-based molecule optimization across six molecular properties.
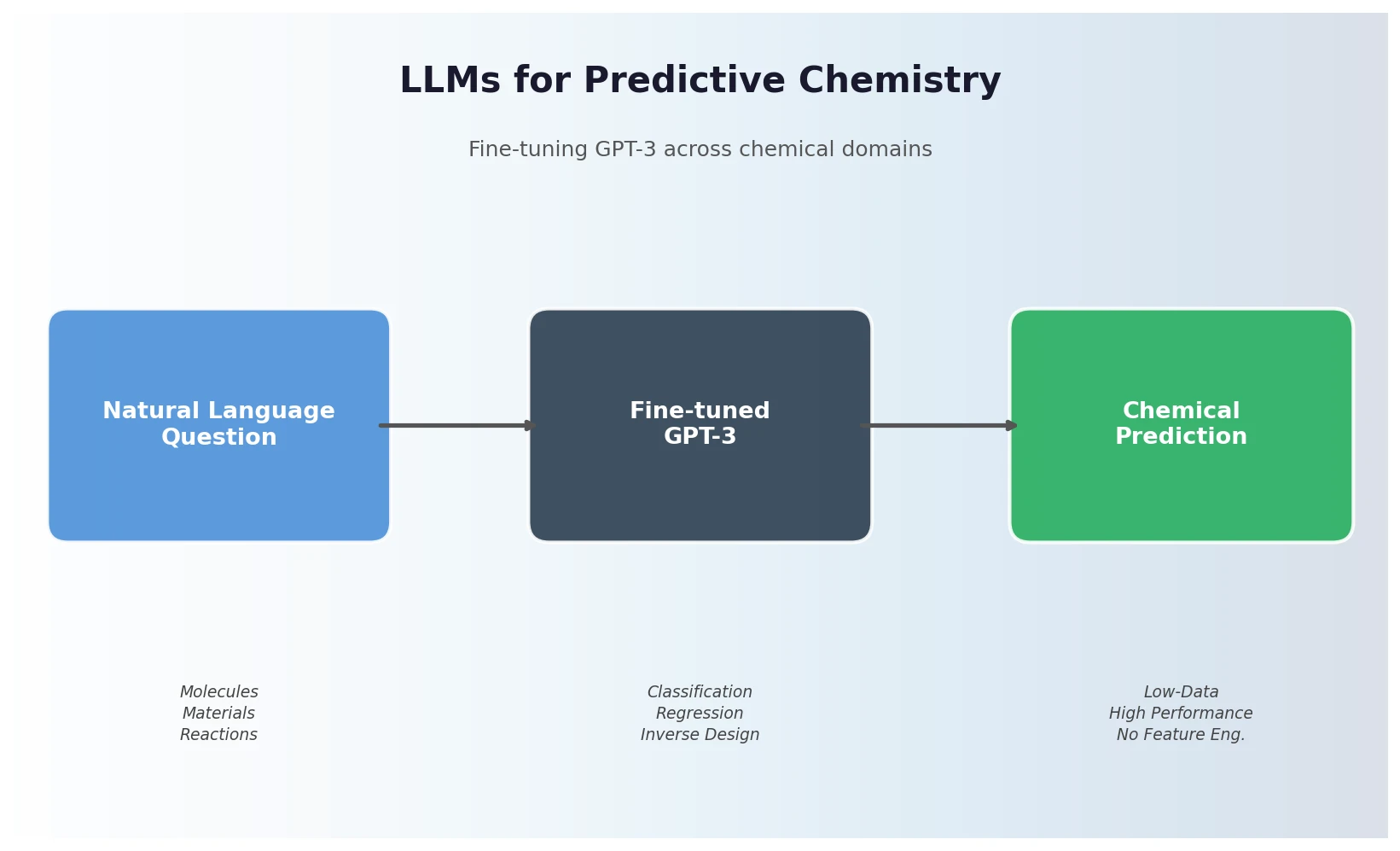
Jablonka et al. show that fine-tuning GPT-3 on natural language chemistry questions achieves competitive or superior performance to dedicated ML models across 15 benchmarks, with particular strength in low-data settings and inverse molecular design.
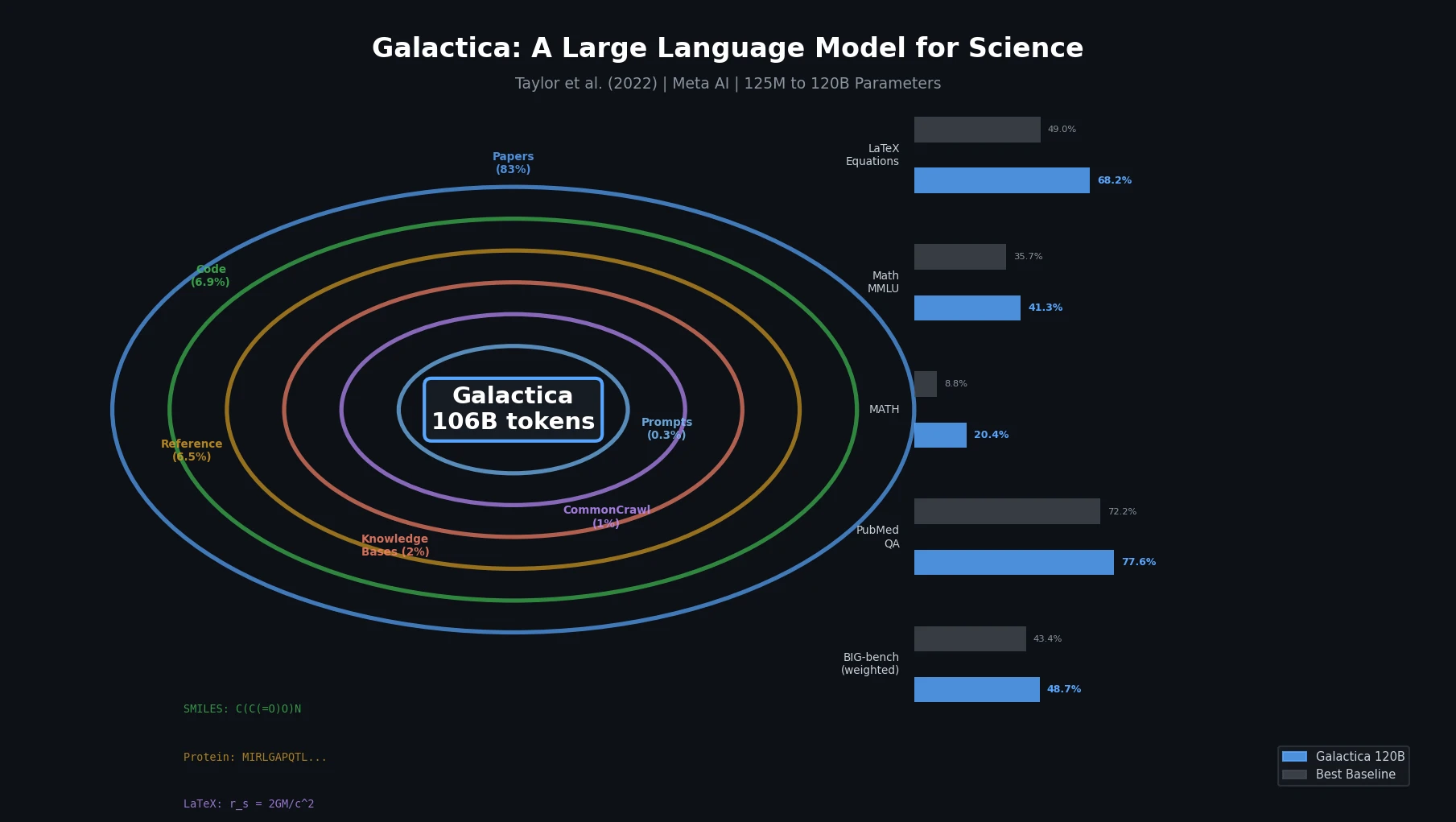
Galactica trains a decoder-only Transformer on a curated 106B-token scientific corpus spanning papers, proteins, and molecules, achieving strong results on scientific QA, mathematical reasoning, and citation prediction.
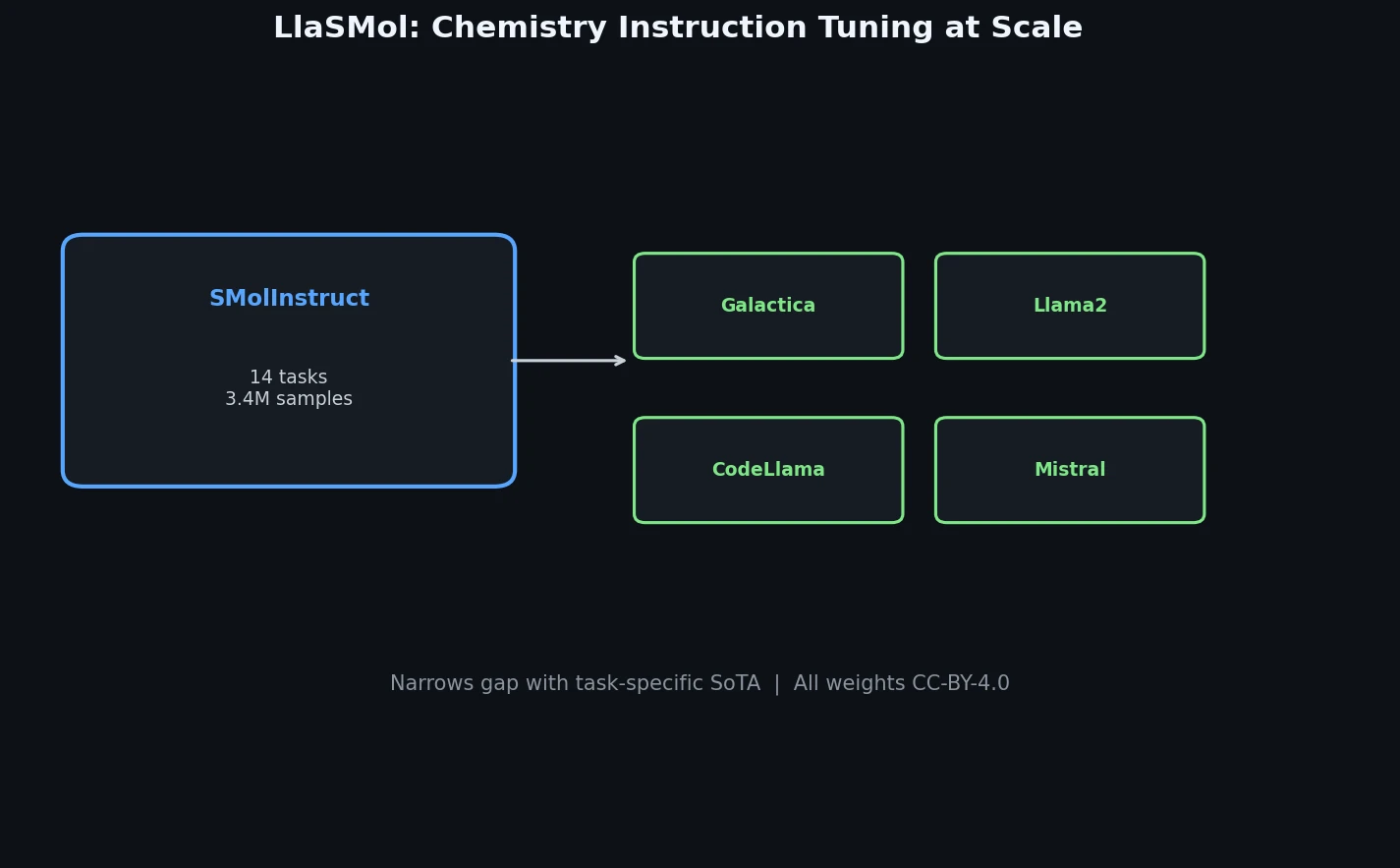
LlaSMol fine-tunes Mistral, Llama 2, and other open-source LLMs on SMolInstruct, a 3.3M-sample instruction tuning dataset covering 14 chemistry tasks. The Mistral-based model outperforms GPT-4 and Claude 3 Opus across all tasks.
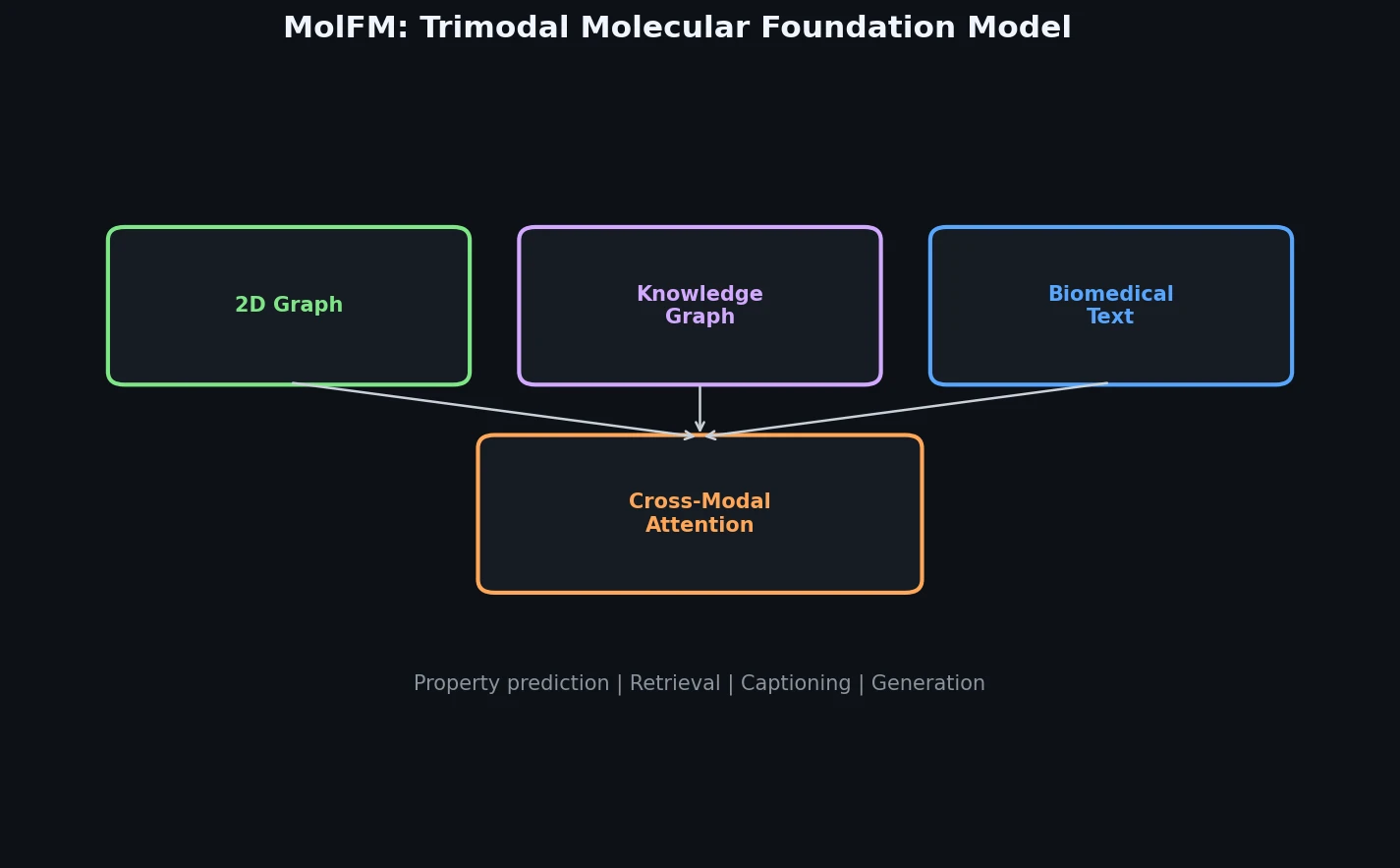
MolFM pre-trains a multimodal encoder that fuses 2D molecular graphs, biomedical text, and knowledge graph entities through fine-grained cross-modal attention, achieving strong gains on cross-modal retrieval, molecule captioning, text-based generation, and property prediction.