GP-MoLFormer: Molecular Generation via Transformers
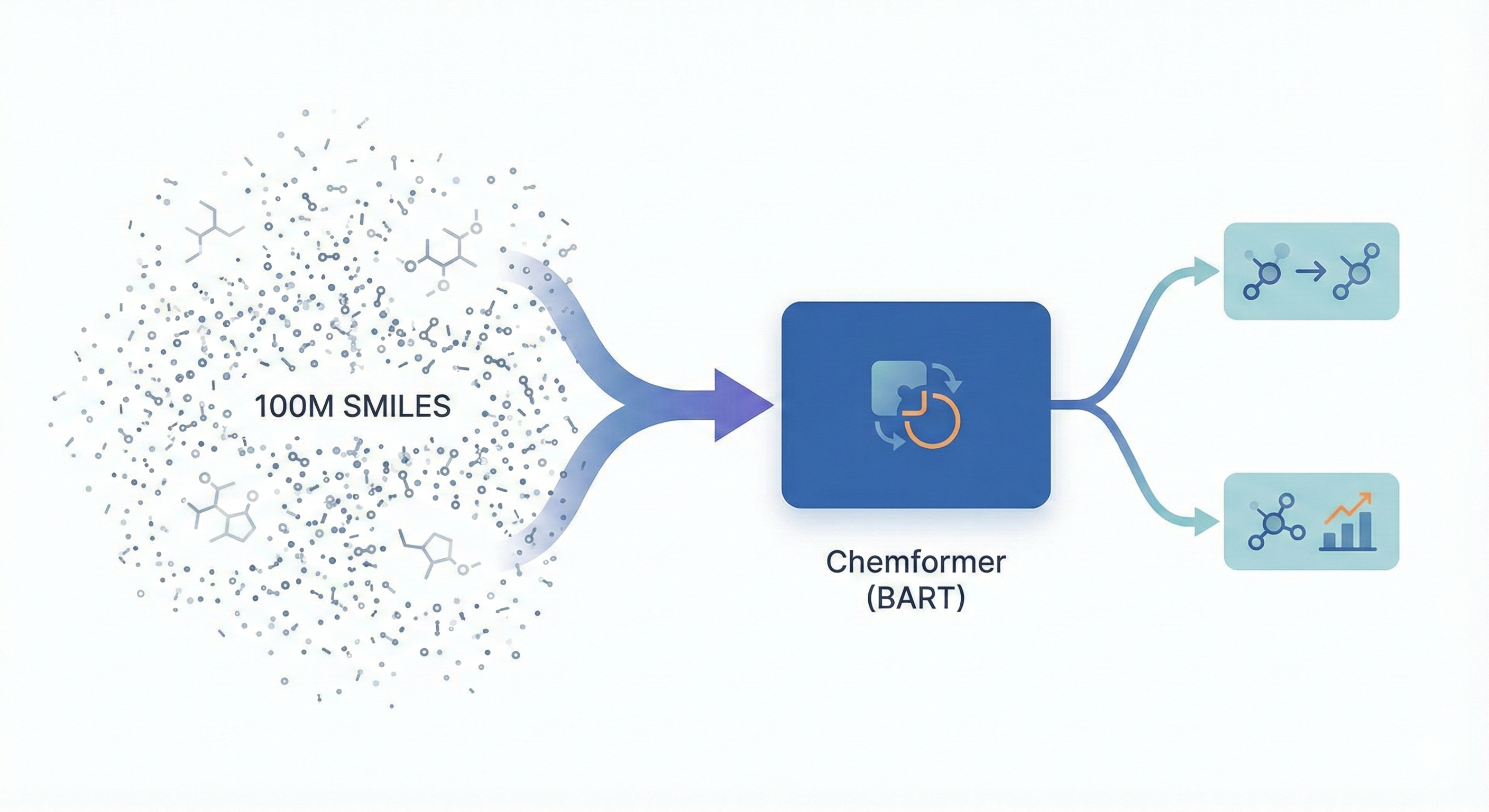
This methodological paper proposes a linear-attention transformer decoder trained on 1.1 billion molecules. It introduces pair-tuning for efficient property optimization and establishes empirical scaling laws relating inference compute to generation novelty.