Recent Advances in the SELFIES Library: 2023 Update
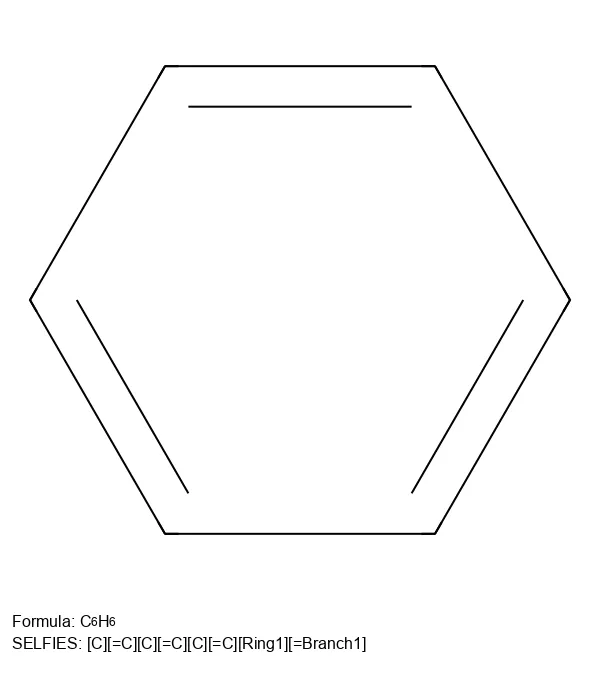
A 2023 software update paper documenting improvements to the SELFIES Python library (v2.1.1), including a streamlined context-free grammar, expanded support for aromatic systems and stereochemistry, customizable semantic constraints, ML utility functions, and performance benchmarks on 300K+ molecules.