SMINA Docking Benchmark for De Novo Drug Design Models
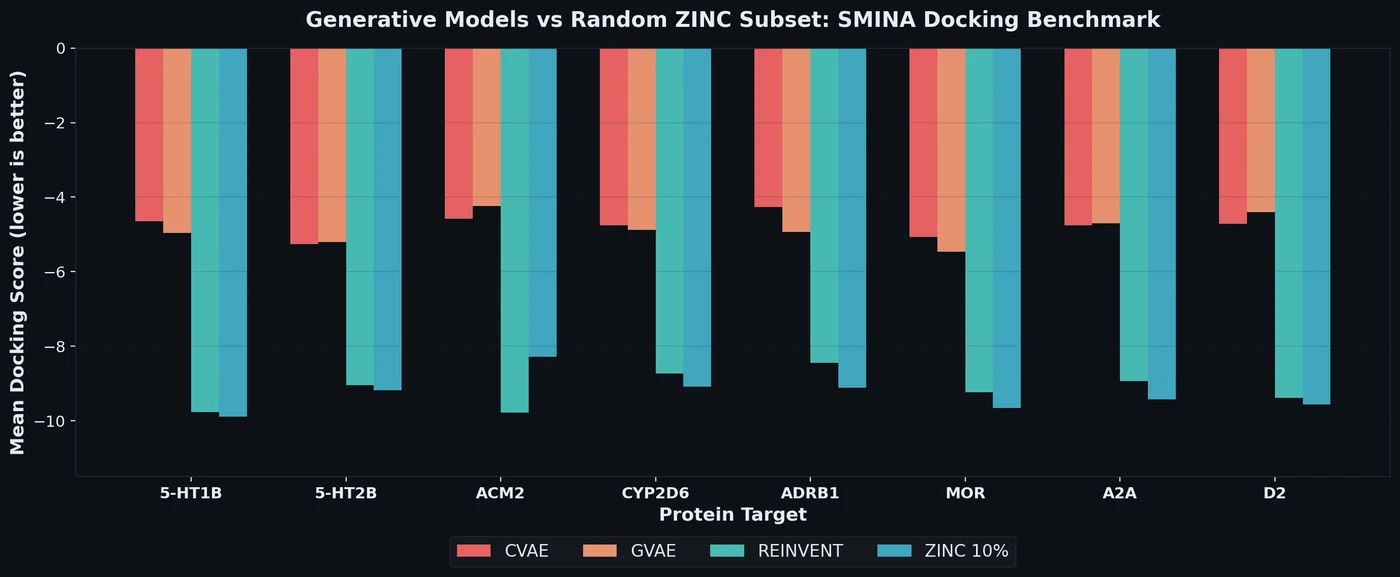
Proposes a benchmark for de novo drug design using SMINA docking scores across eight drug targets, revealing that popular generative models fail to outperform random ZINC subsets.

Proposes a benchmark for de novo drug design using SMINA docking scores across eight drug targets, revealing that popular generative models fail to outperform random ZINC subsets.
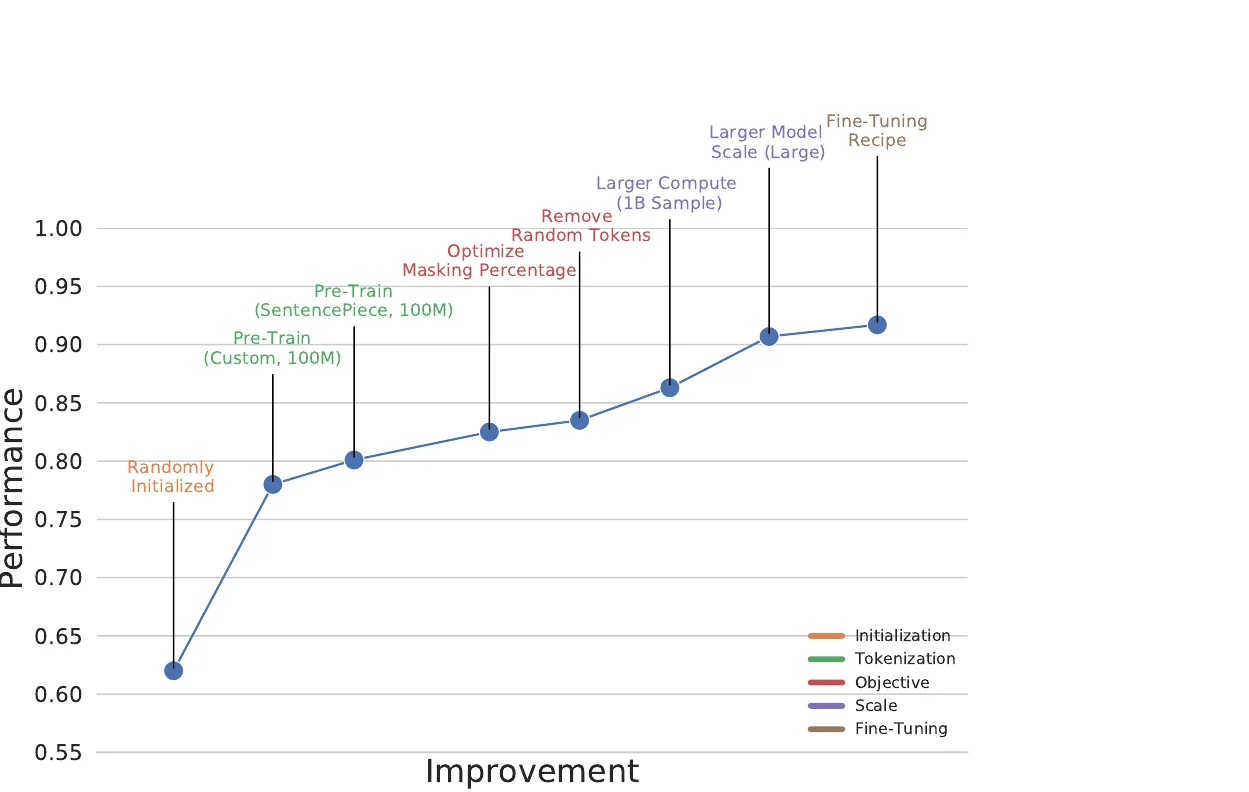
BARTSmiles pre-trains a BART-large model on 1.7 billion SMILES strings from ZINC20 and achieves the best reported results on 11 classification, regression, and generation benchmarks.
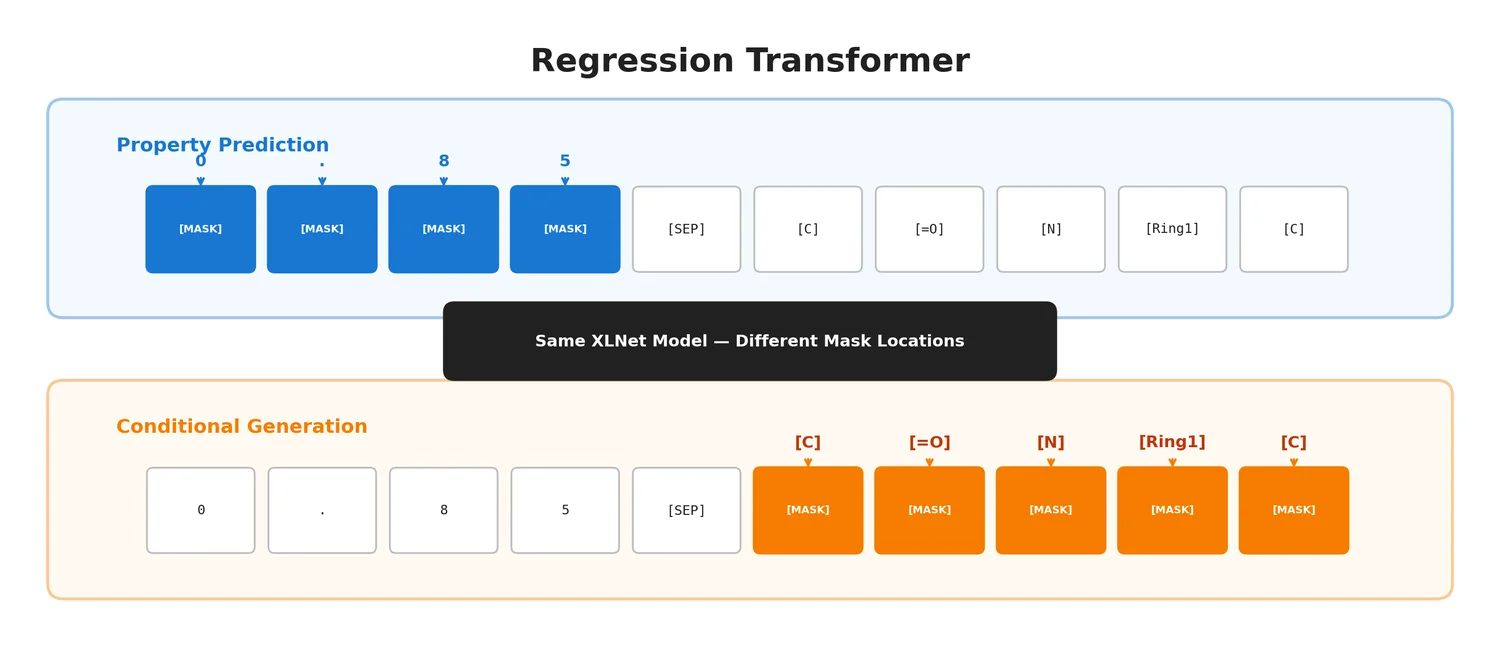
The Regression Transformer (RT) reformulates regression as conditional sequence modelling, enabling a single XLNet-based model to both predict continuous molecular properties and generate novel molecules conditioned on desired property values.
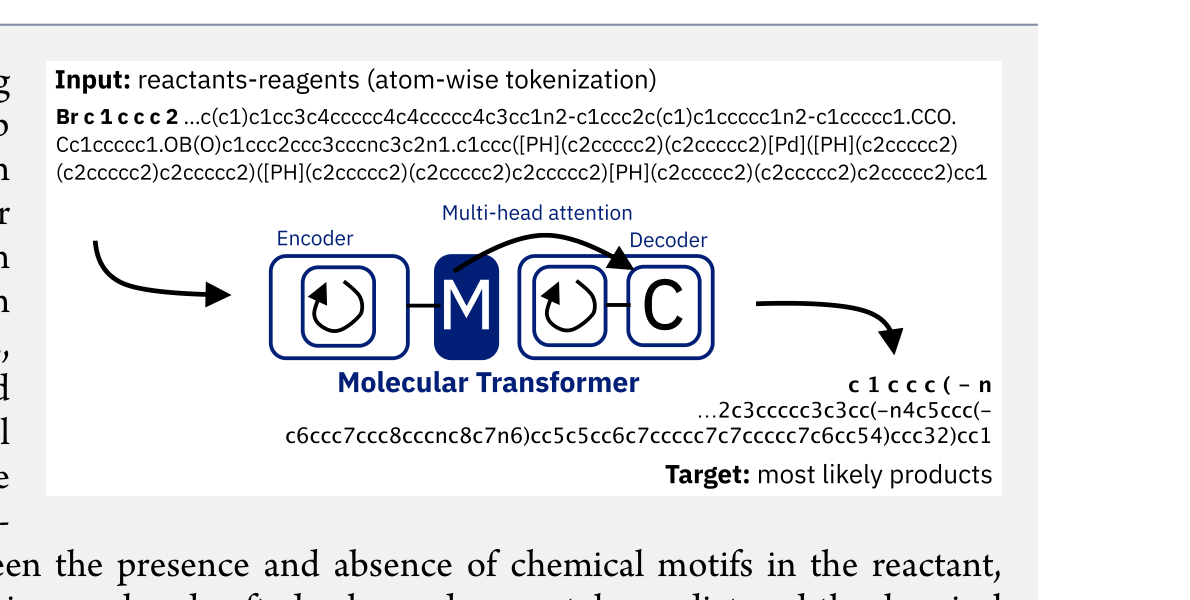
The Molecular Transformer applies the Transformer architecture to forward reaction prediction, treating it as SMILES-to-SMILES machine translation. It achieves 90.4% top-1 accuracy on USPTO_MIT, outperforms quantum-chemistry baselines on regioselectivity, and provides calibrated uncertainty scores (0.89 AUC-ROC) for ranking synthesis pathways.
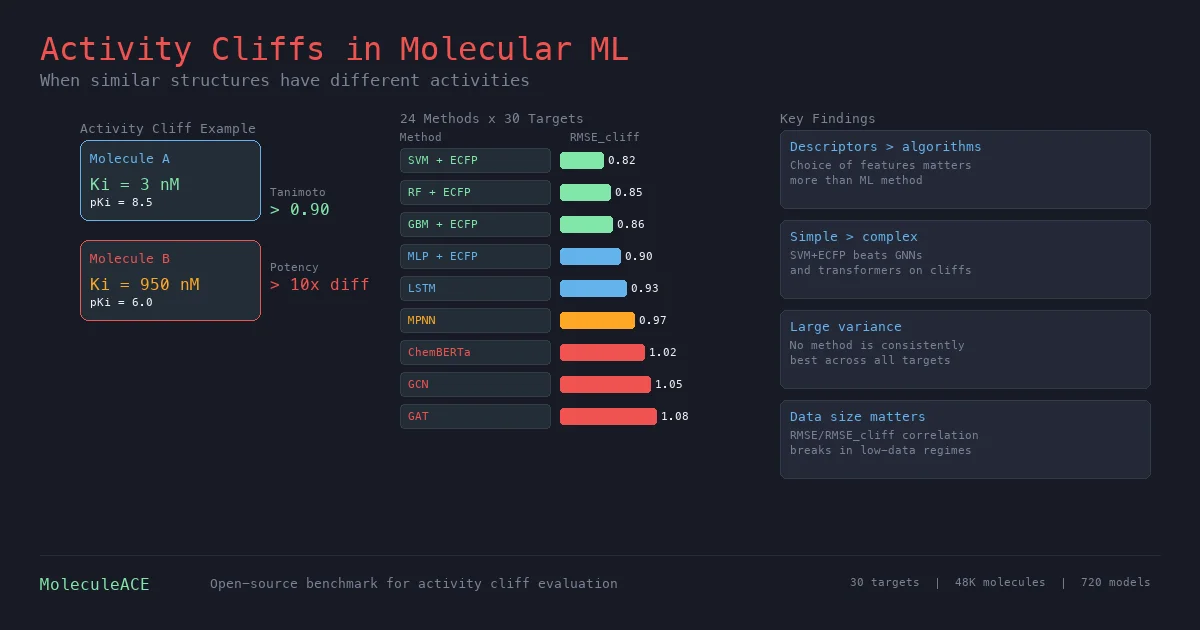
This paper benchmarks 24 machine and deep learning methods on activity cliff compounds (structurally similar molecules with large potency differences) across 30 macromolecular targets. Traditional ML with molecular fingerprints consistently outperforms graph neural networks and SMILES-based transformers on these challenging cases, especially in low-data regimes.
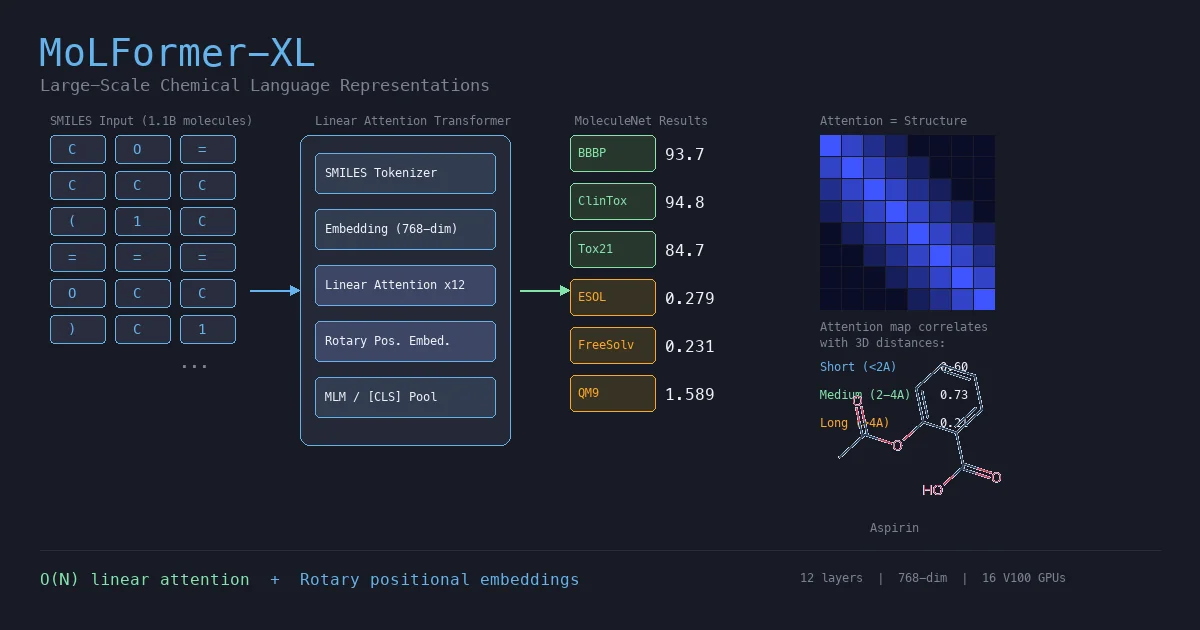
MoLFormer is a transformer encoder with linear attention and rotary positional embeddings, pretrained via masked language modeling on 1.1 billion molecules from PubChem and ZINC. MoLFormer-XL outperforms GNN baselines on most MoleculeNet classification and regression tasks, and attention analysis reveals that the model learns interatomic spatial relationships directly from SMILES strings.
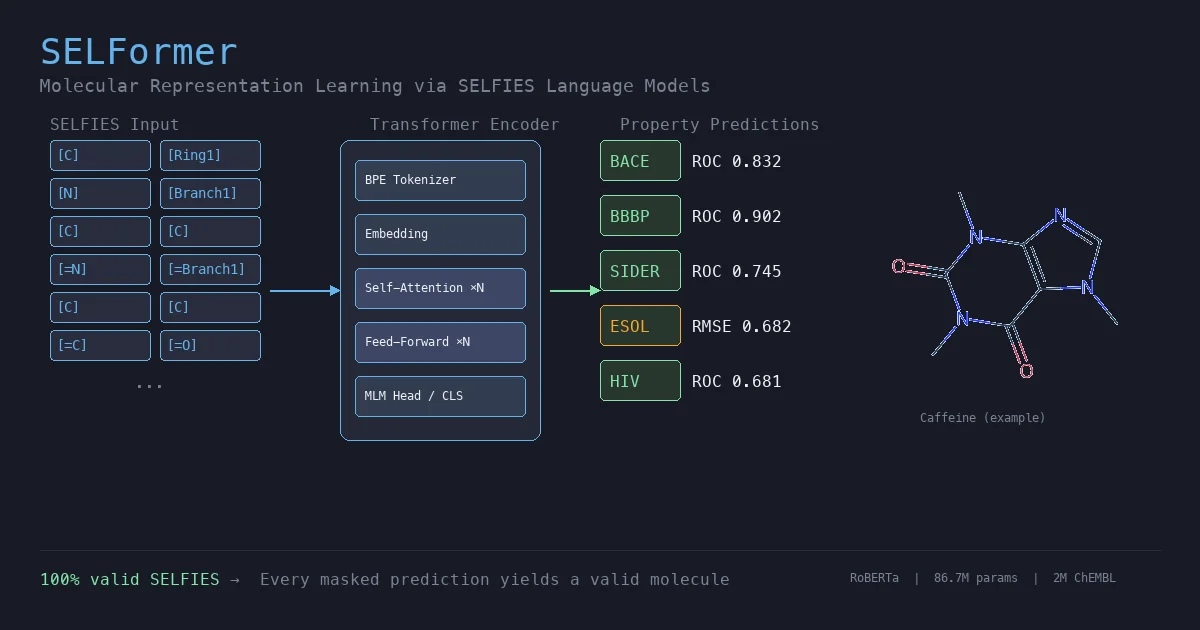
SELFormer is a transformer-based chemical language model that uses SELFIES instead of SMILES as input. Pretrained on 2M ChEMBL compounds via masked language modeling, it achieves strong classification performance on MoleculeNet tasks, outperforming ChemBERTa-2 by ~12% on average across BACE, BBBP, and HIV.
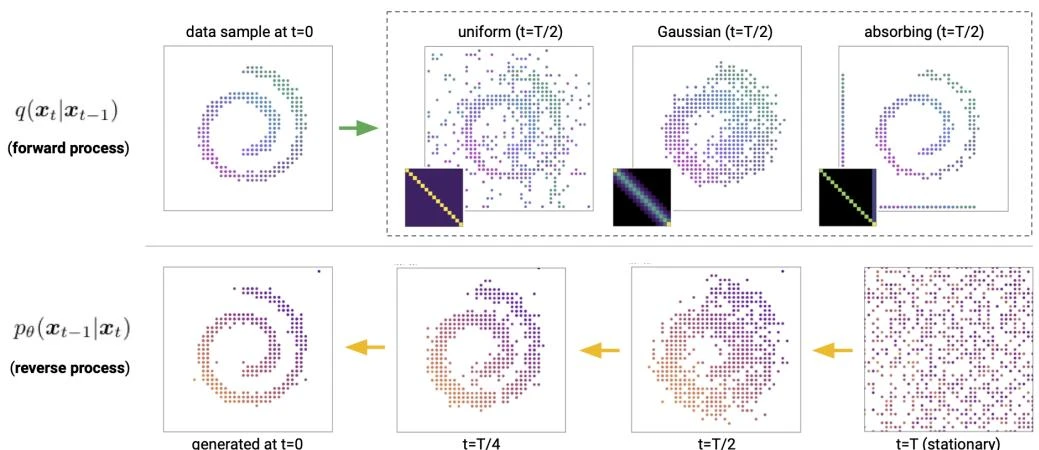
This paper introduces Discrete Denoising Diffusion Probabilistic Models (D3PMs), which generalize diffusion to discrete state-spaces using structured Markov transition matrices. D3PMs include uniform, absorbing-state, and discretized Gaussian corruption processes, drawing a connection between diffusion and masked language models.
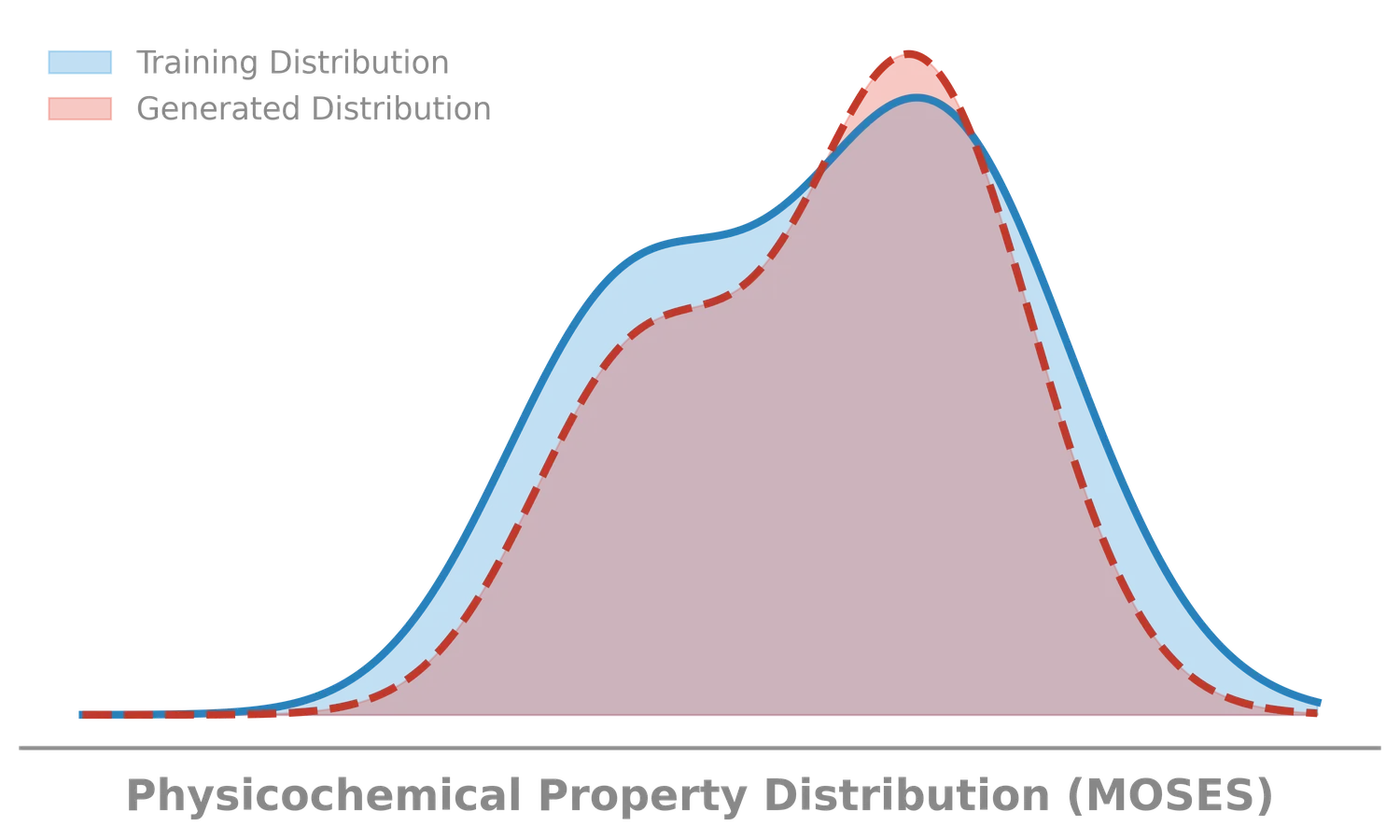
MOSES introduces a comprehensive benchmarking platform for molecular generative models, offering standardized datasets, evaluation metrics, and baselines. By providing a unified measuring stick, it aims to resolve reproducibility challenges in chemical distribution learning.
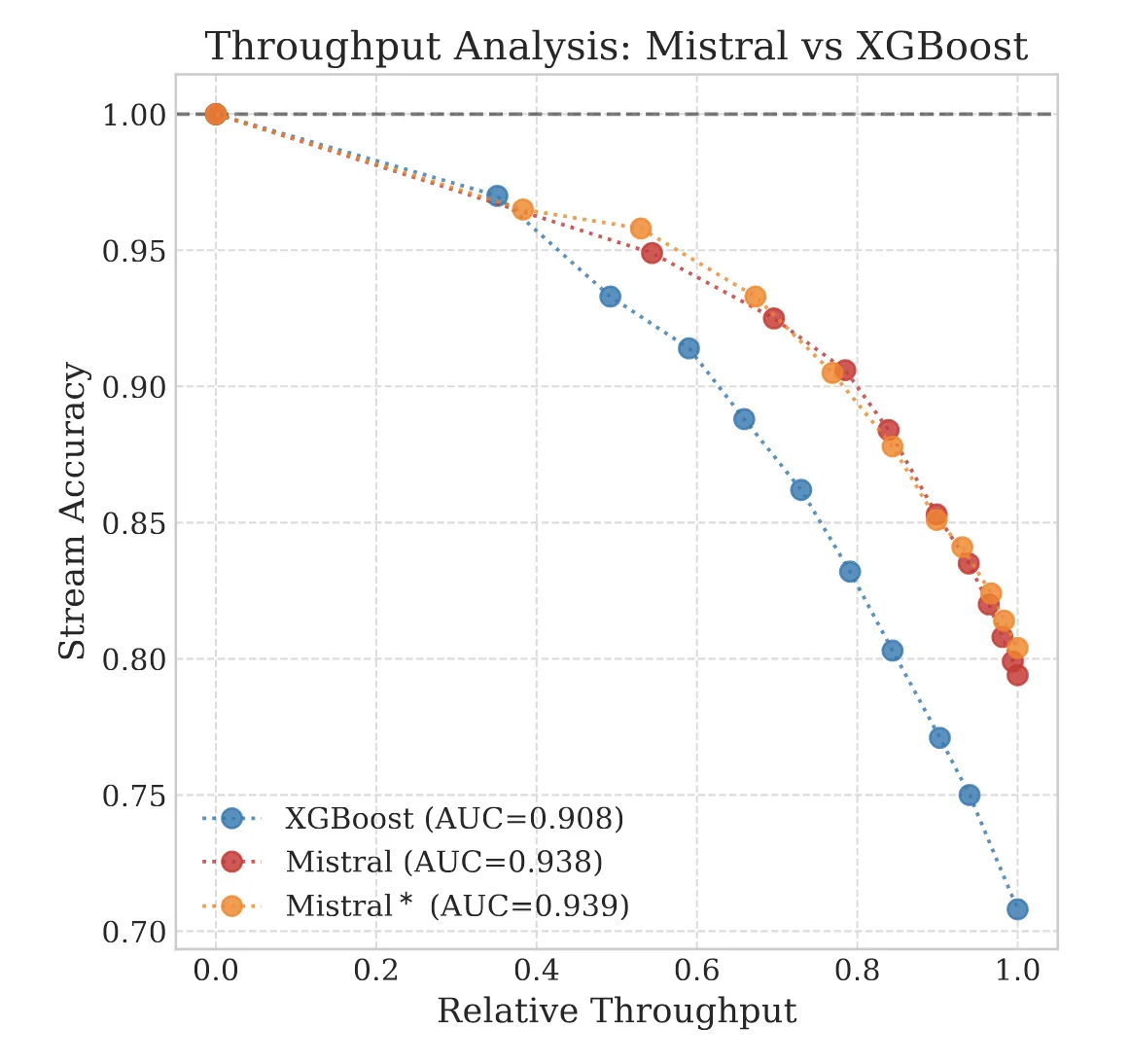
We explore the ‘Silent Failure’ mode of LLMs in production: the limits of 99% accuracy for reliability, how confidence decays in long documents, and why standard calibration techniques struggle to fix it.
We trace the history of Page Stream Segmentation (PSS) through three eras (Heuristic, Encoder, and Decoder) and explain how privacy-preserving, localized LLMs enable true semantic processing.

This work investigates the scaling hypothesis for molecular transformers, training RoBERTa models on 77M SMILES from PubChem. It compares Masked Language Modeling (MLM) against Multi-Task Regression (MTR) pretraining, finding that MTR yields better downstream performance but is computationally heavier.