Re-evaluating Sample Efficiency in Molecule Generation
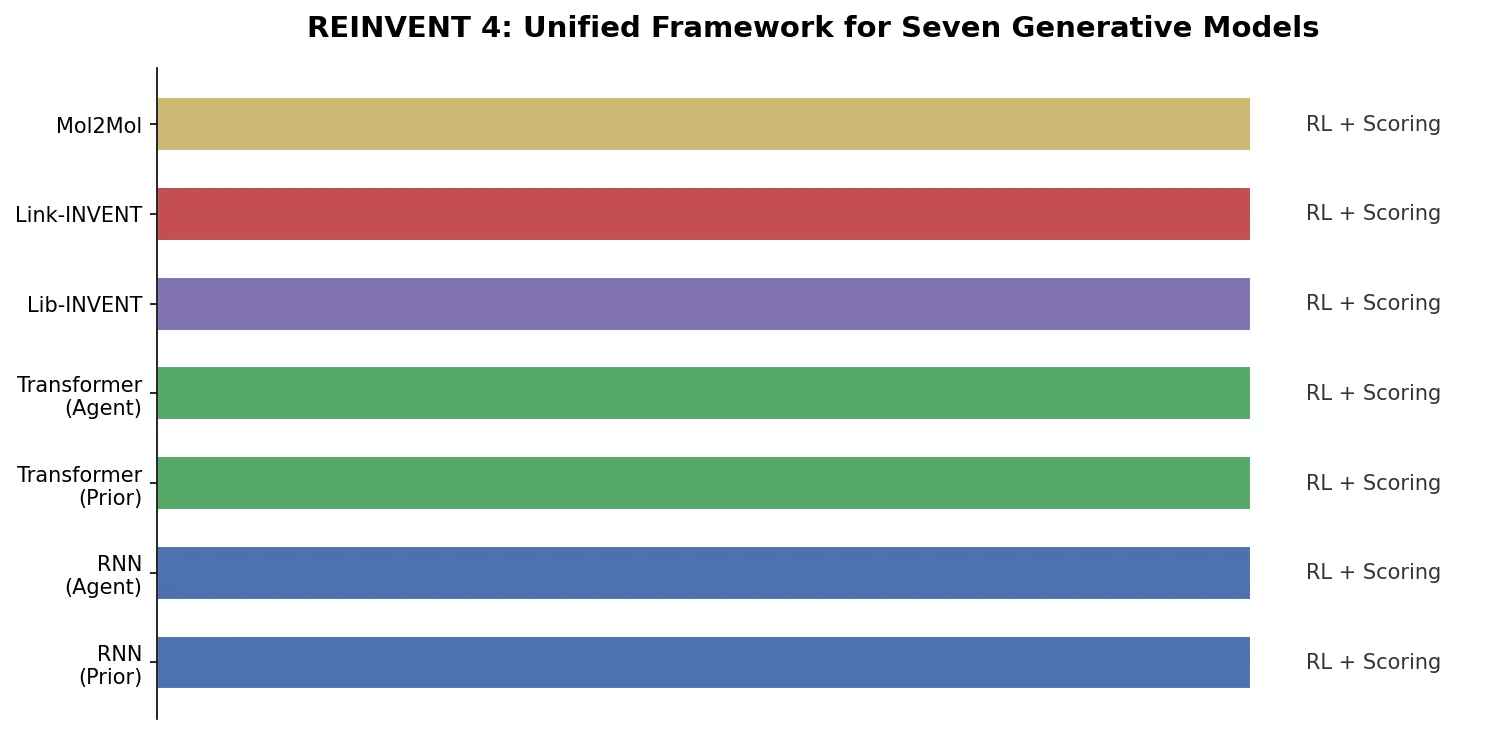
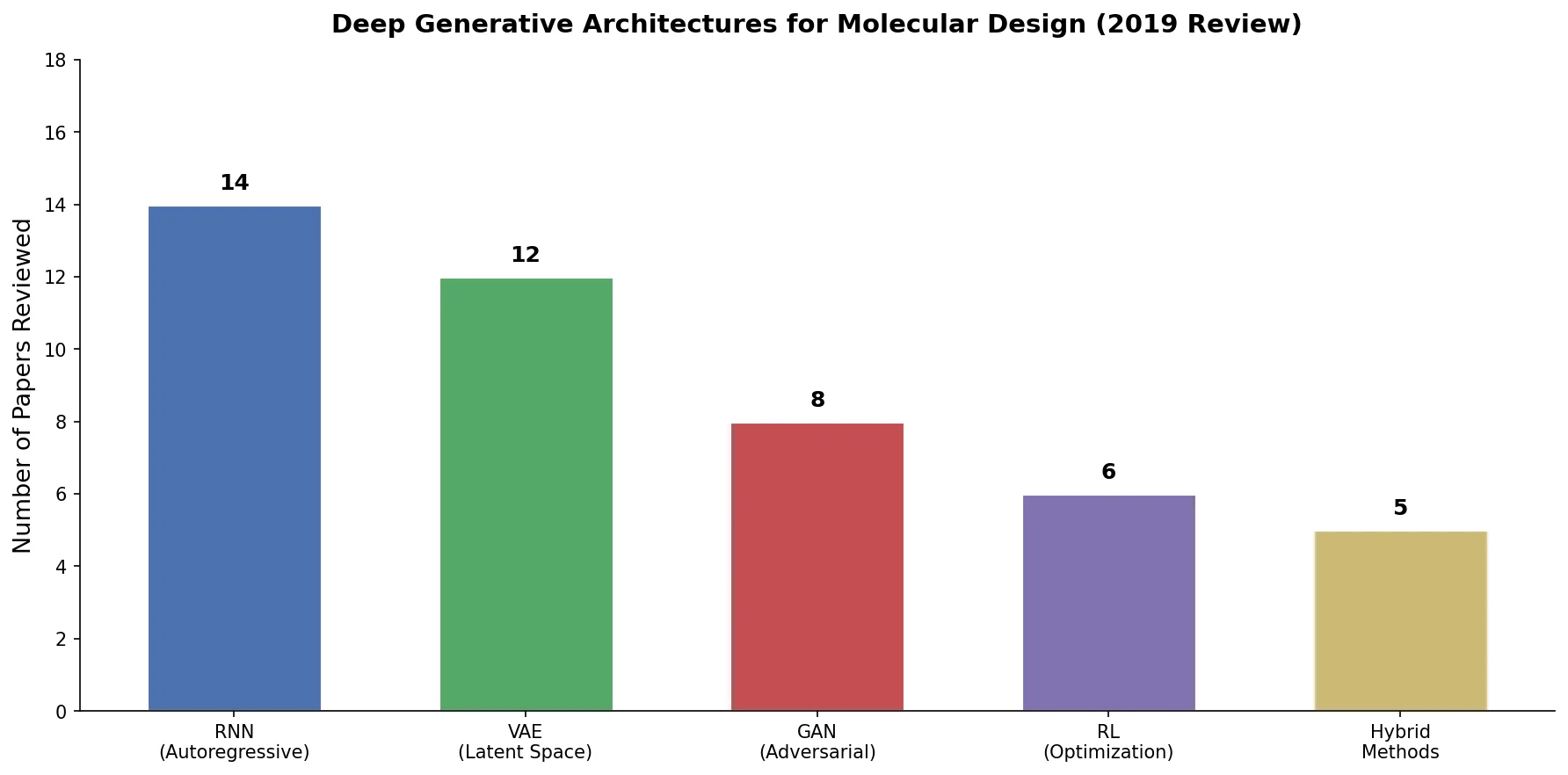
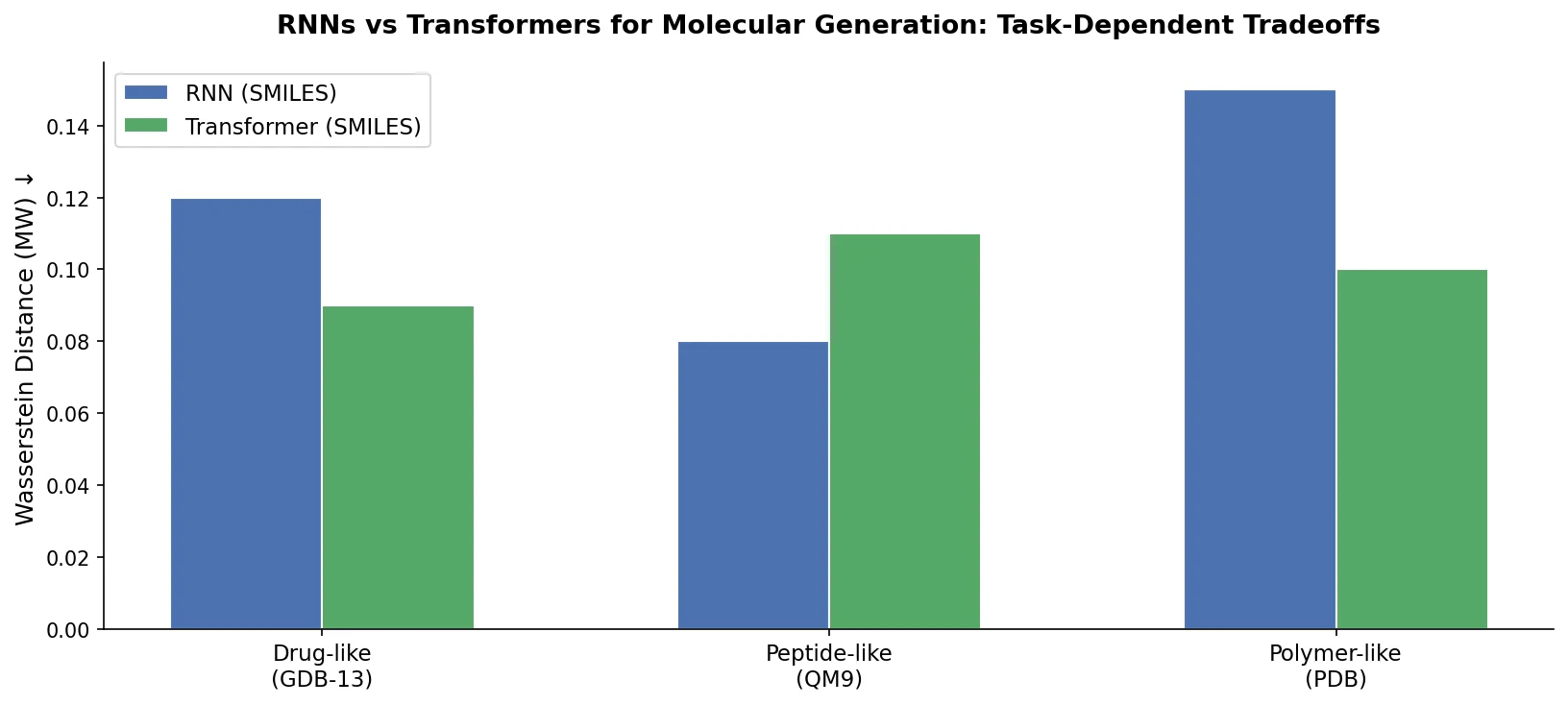
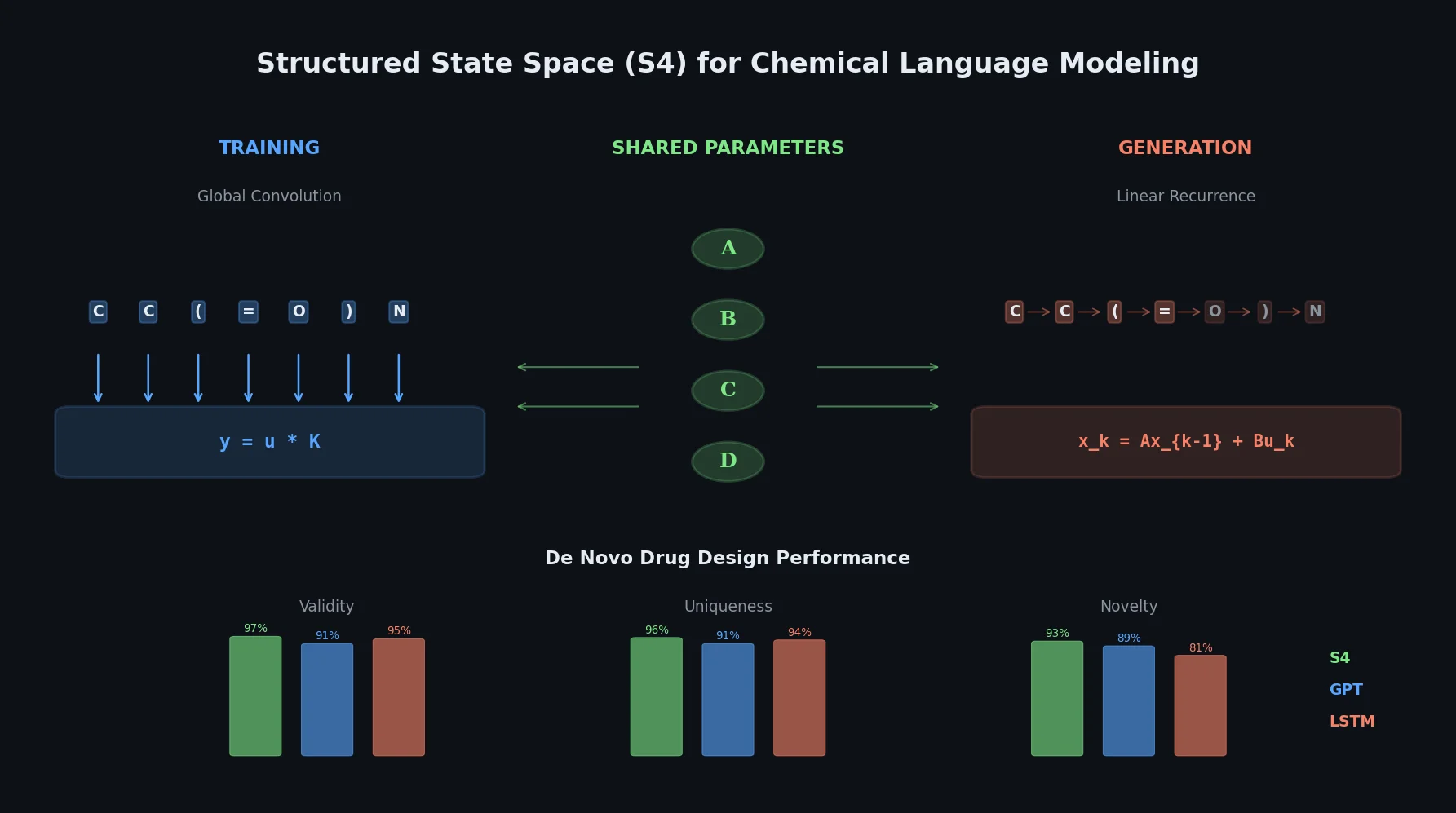
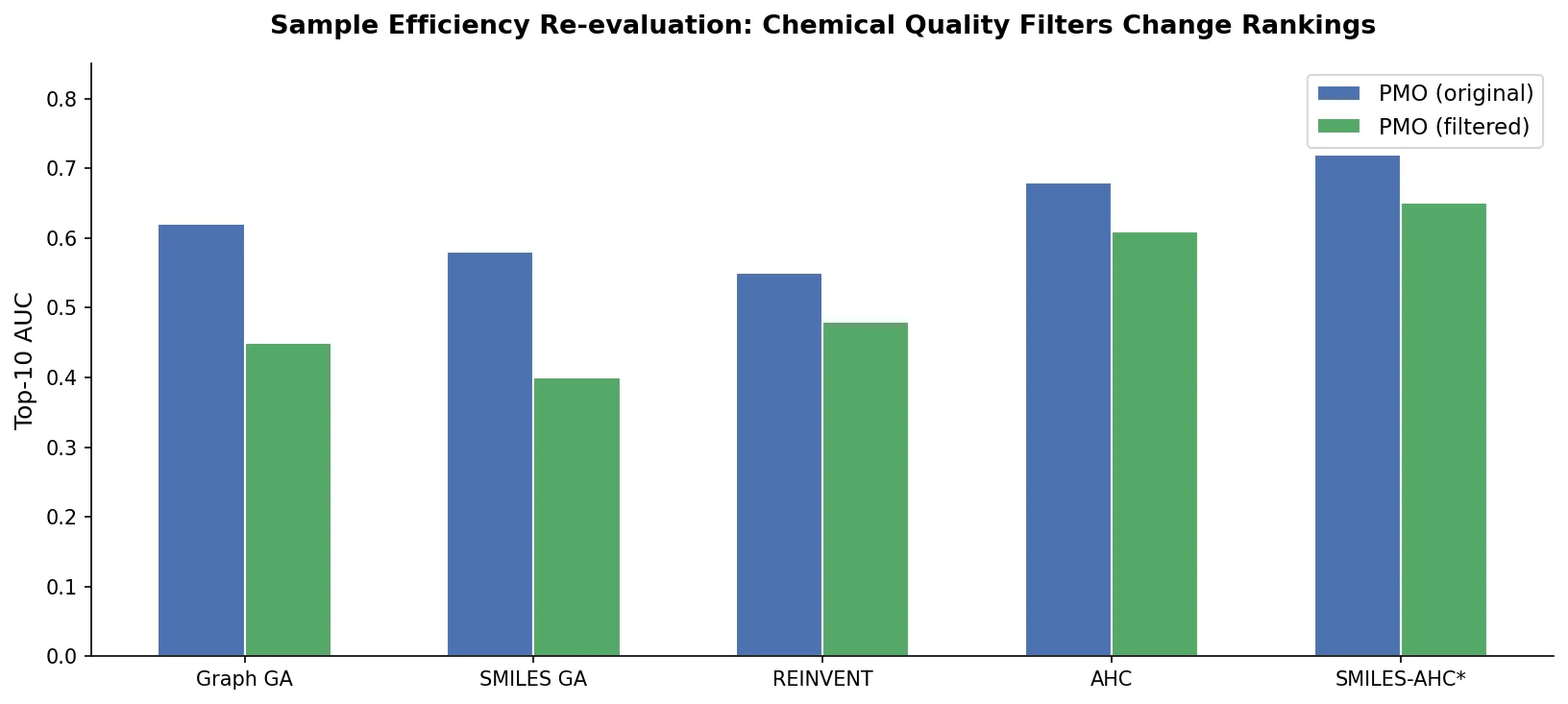
A critical reassessment of the PMO benchmark for de novo molecule generation, showing that adding molecular weight, LogP, and diversity filters substantially re-ranks generative models, with Augmented Hill-Climb emerging as the top method.