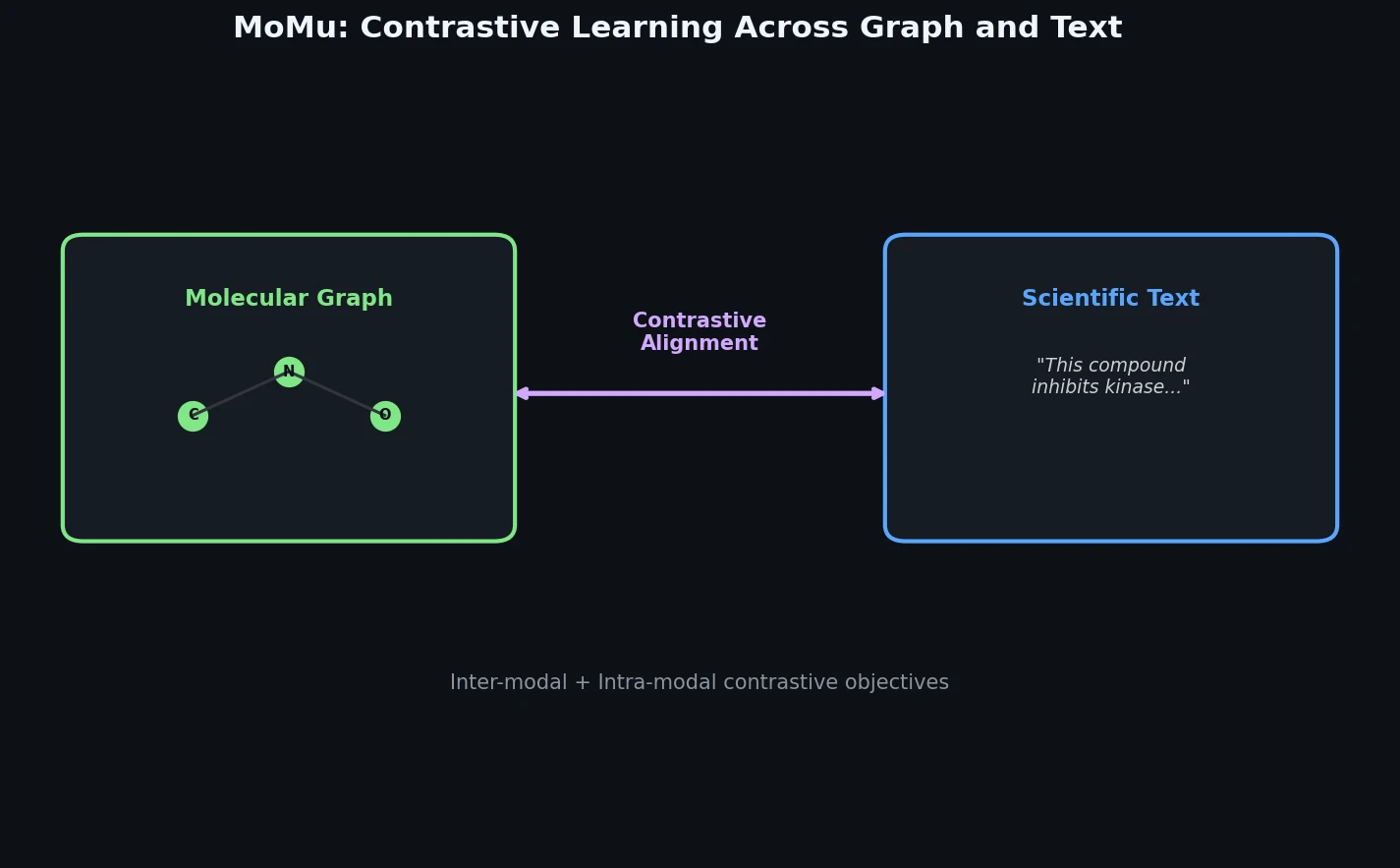
MoMu: Bridging Molecular Graphs and Natural Language
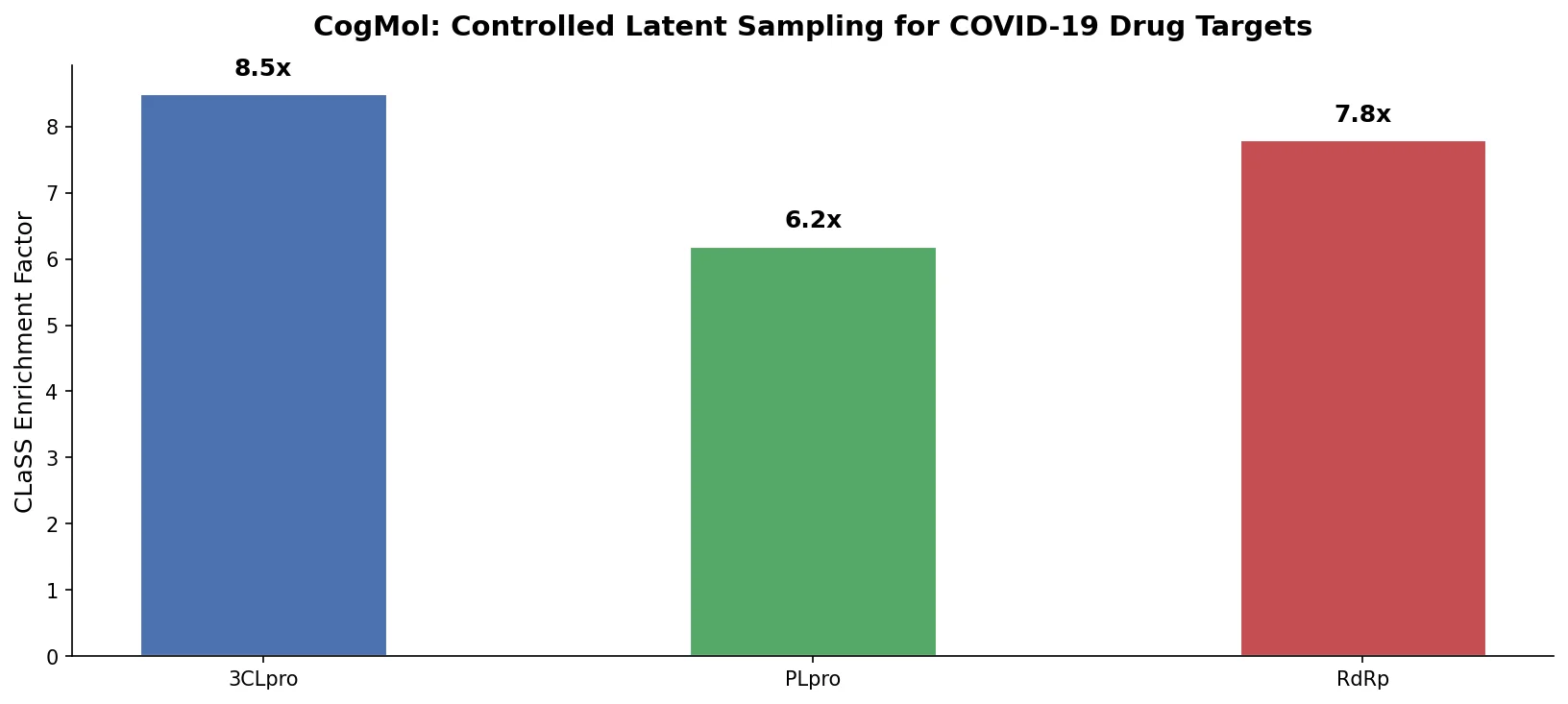
MoMu pre-trains dual graph and text encoders on 15K molecule graph-text pairs using contrastive learning, enabling cross-modal retrieval, molecule captioning, zero-shot text-to-graph generation, and improved molecular property prediction.