
GutenOCR: A Grounded Vision-Language Front-End for Documents
GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.

GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.
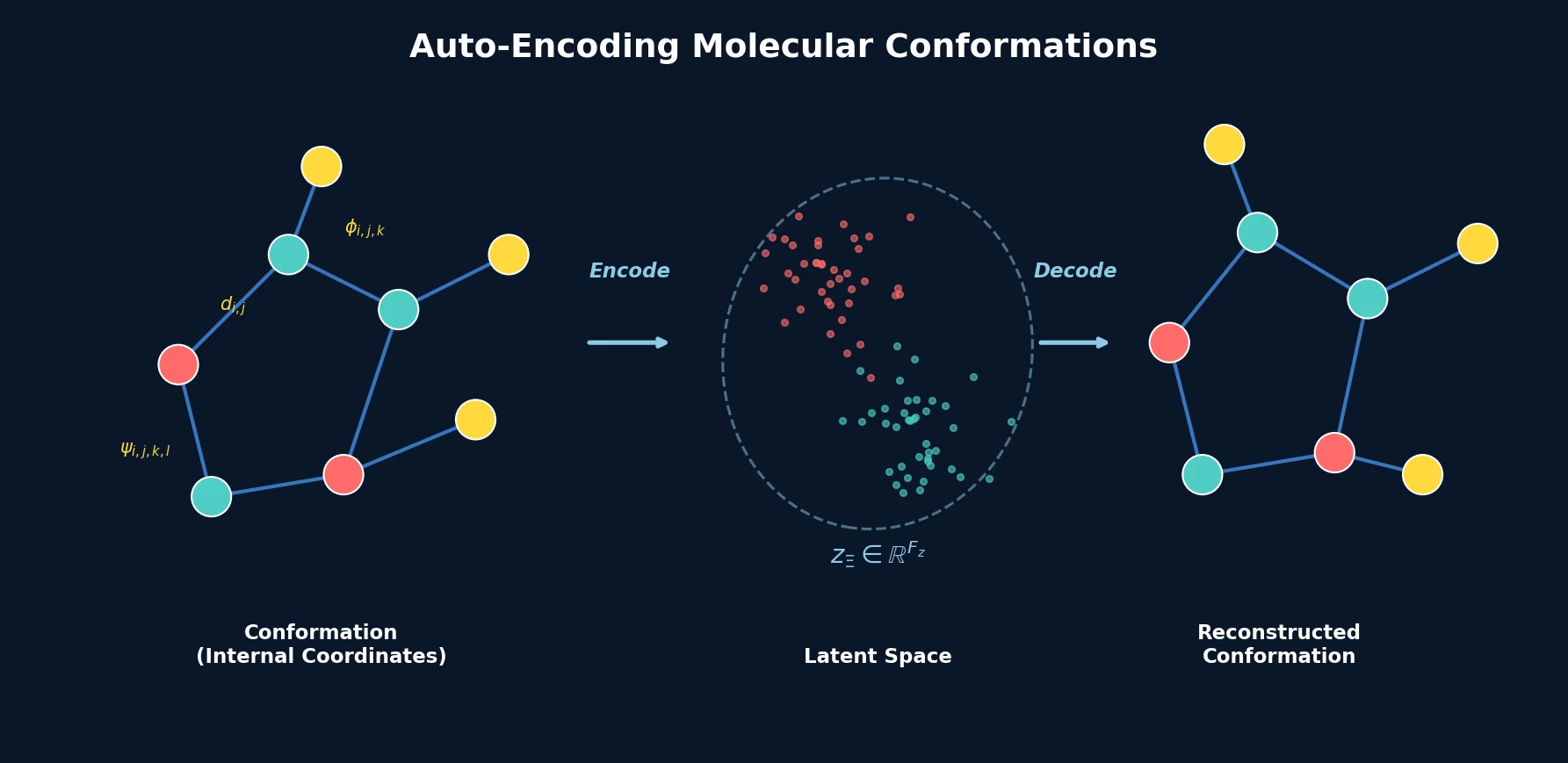
A conformation autoencoder converts molecular 3D arrangements into fixed-size latent representations using internal coordinates and graph neural networks, enabling conformer generation and spatial property optimization.
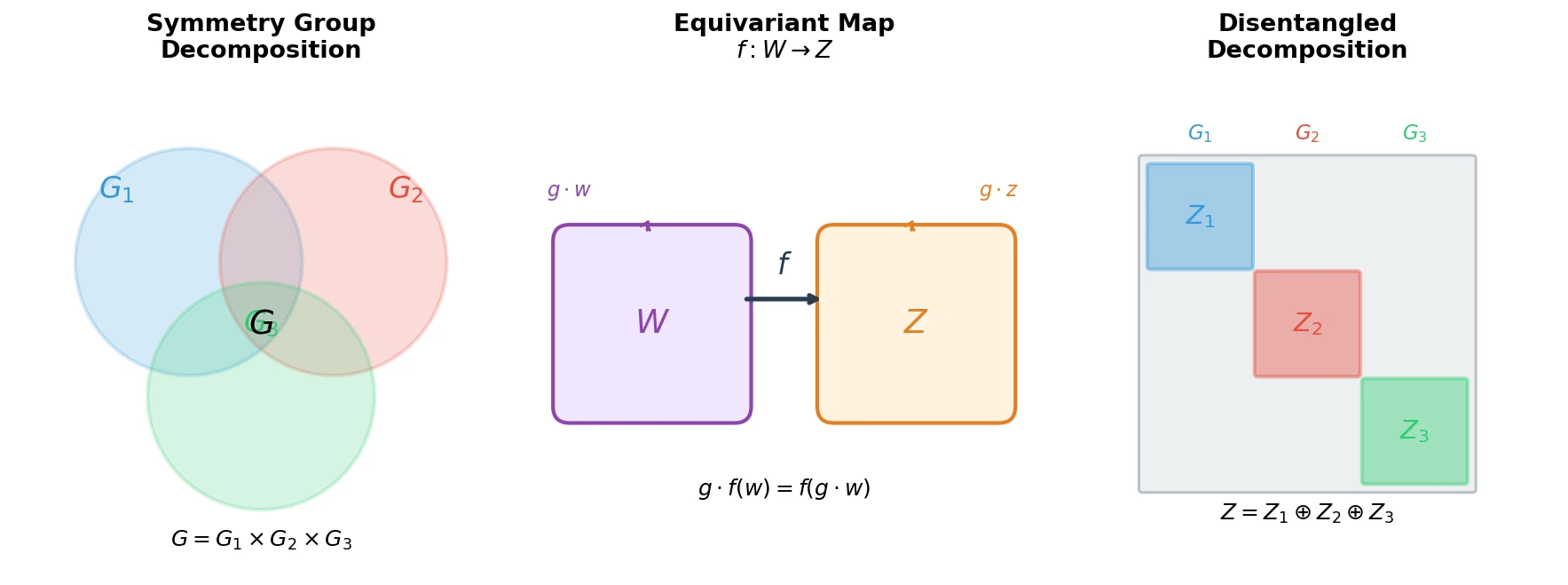
Proposes the first principled mathematical definition of disentangled representations by connecting symmetry group decompositions to independent subspaces in a representation’s vector space.
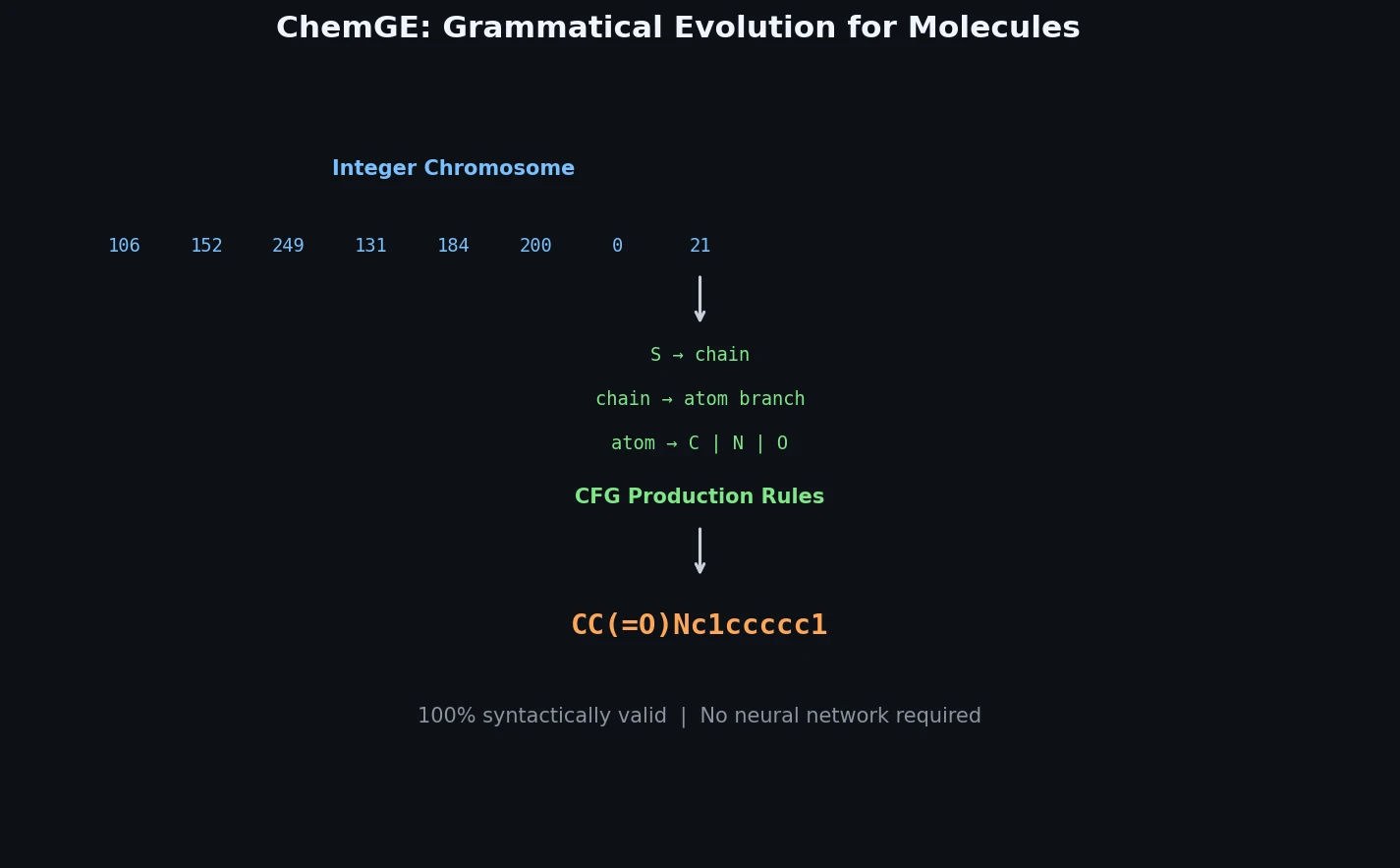
ChemGE uses grammatical evolution over SMILES context-free grammars to generate diverse drug-like molecules in parallel, outperforming deep learning baselines in throughput and molecular diversity.
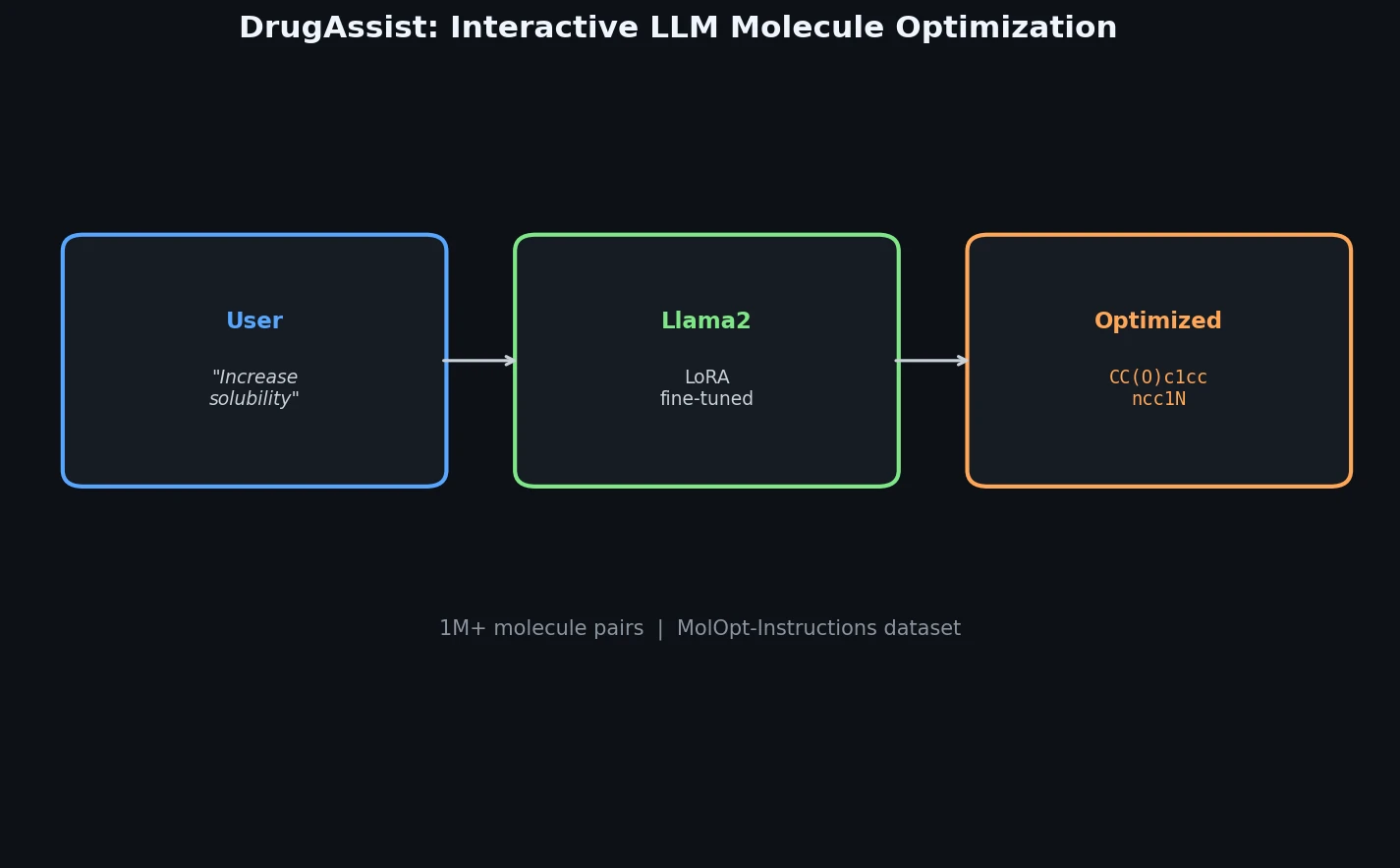
DrugAssist fine-tunes Llama2-7B-Chat on over one million molecule pairs for interactive, dialogue-based molecule optimization across six molecular properties.
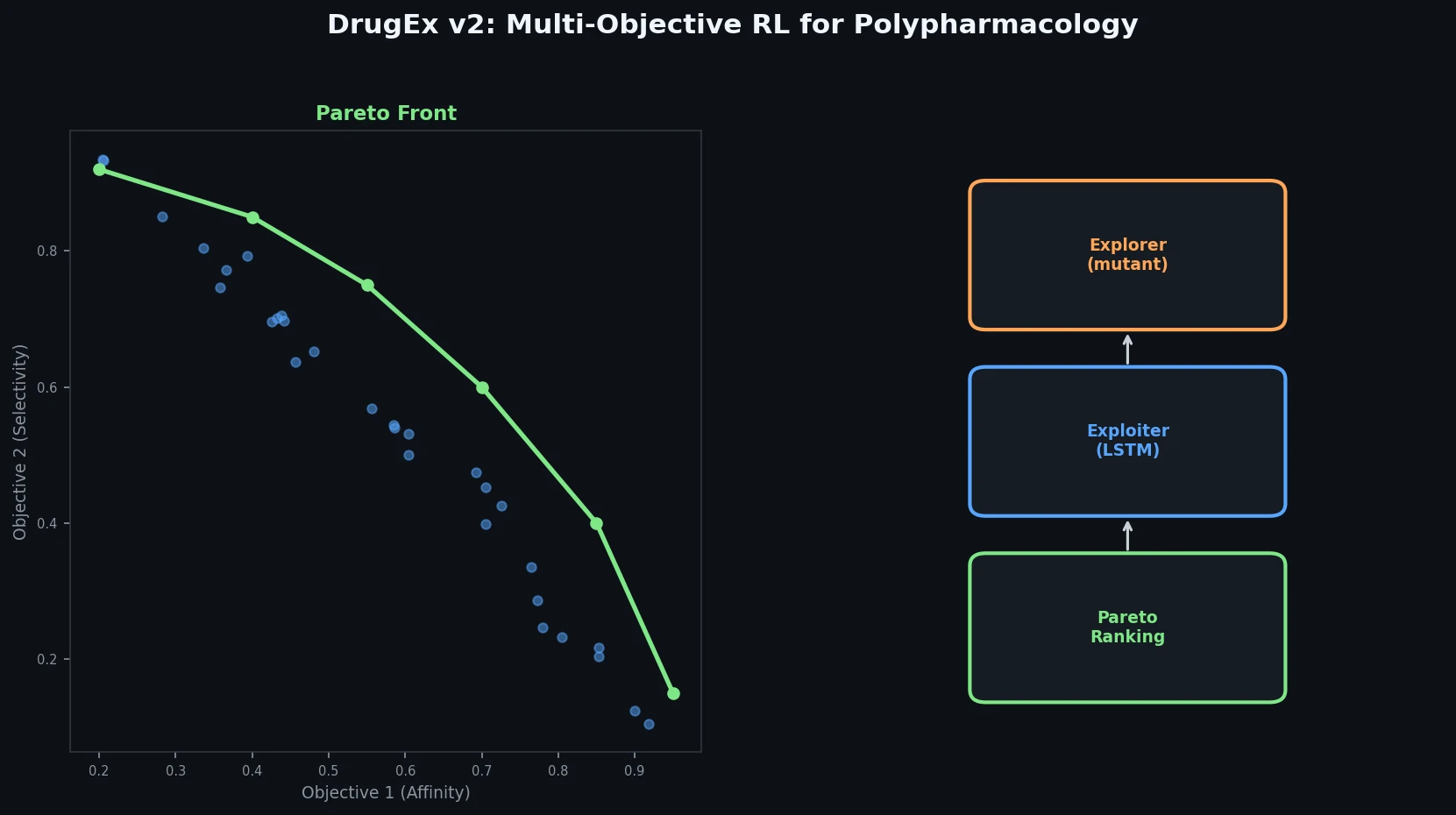
DrugEx v2 introduces Pareto-based multi-objective optimization and evolutionary exploration strategies into an RNN reinforcement learning framework for de novo drug design toward multiple protein targets.
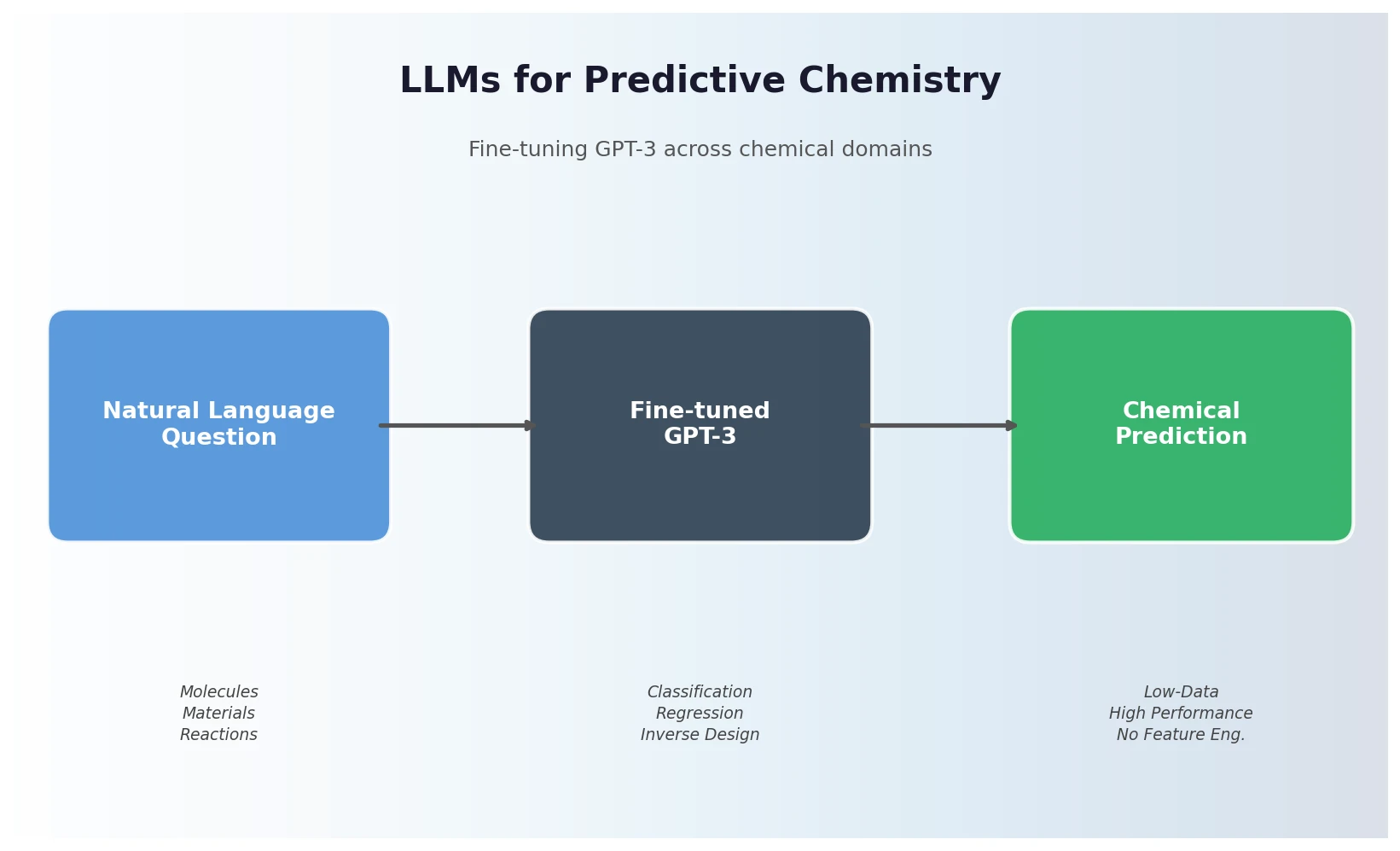
Jablonka et al. show that fine-tuning GPT-3 on natural language chemistry questions achieves competitive or superior performance to dedicated ML models across 15 benchmarks, with particular strength in low-data settings and inverse molecular design.
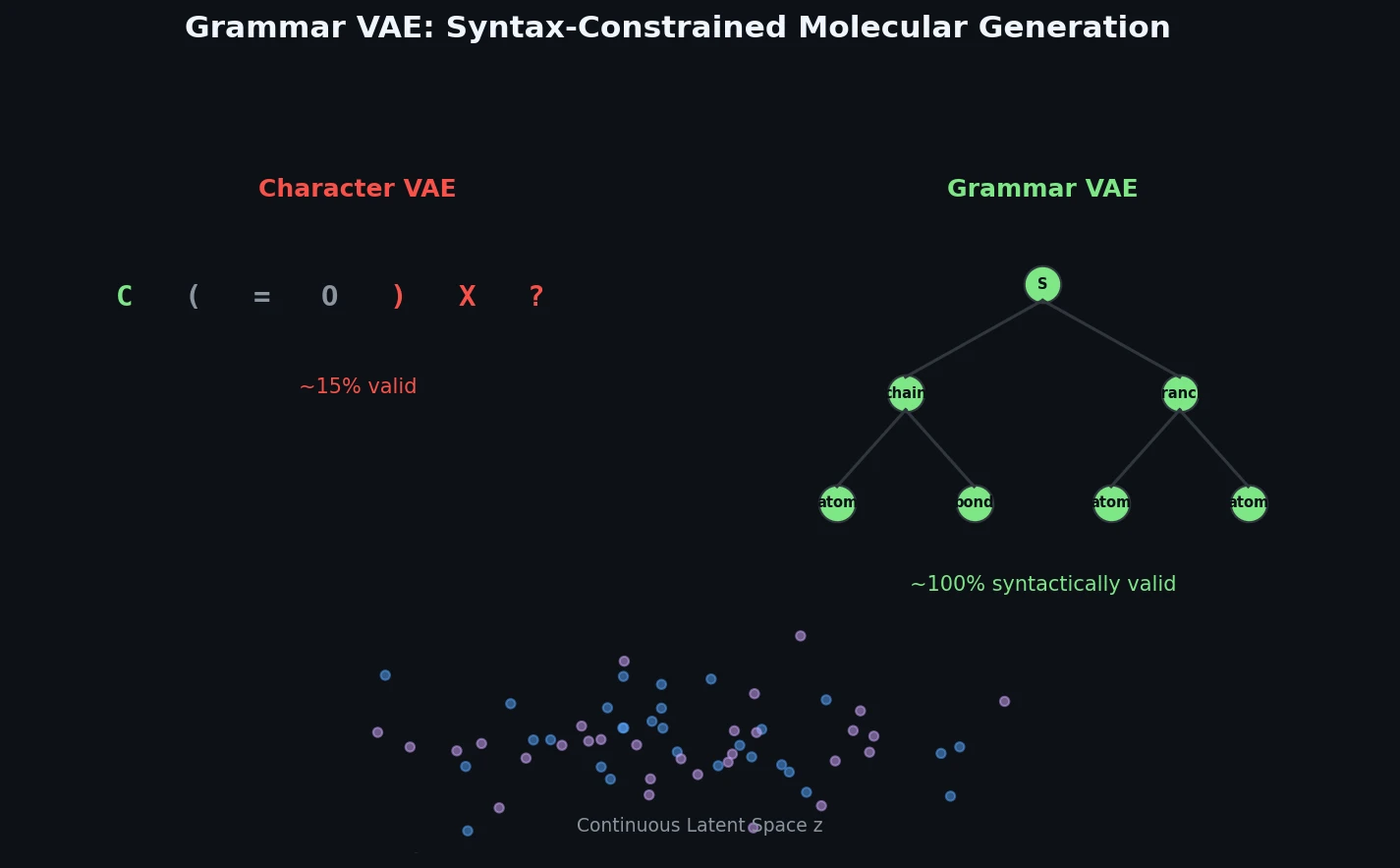
The Grammar VAE replaces character-level decoding with context-free grammar production rules, using a stack-based masking mechanism to guarantee that all generated SMILES strings are syntactically valid. Applied to molecular optimization and symbolic regression, it learns smoother latent spaces and finds better molecules than character-level baselines.
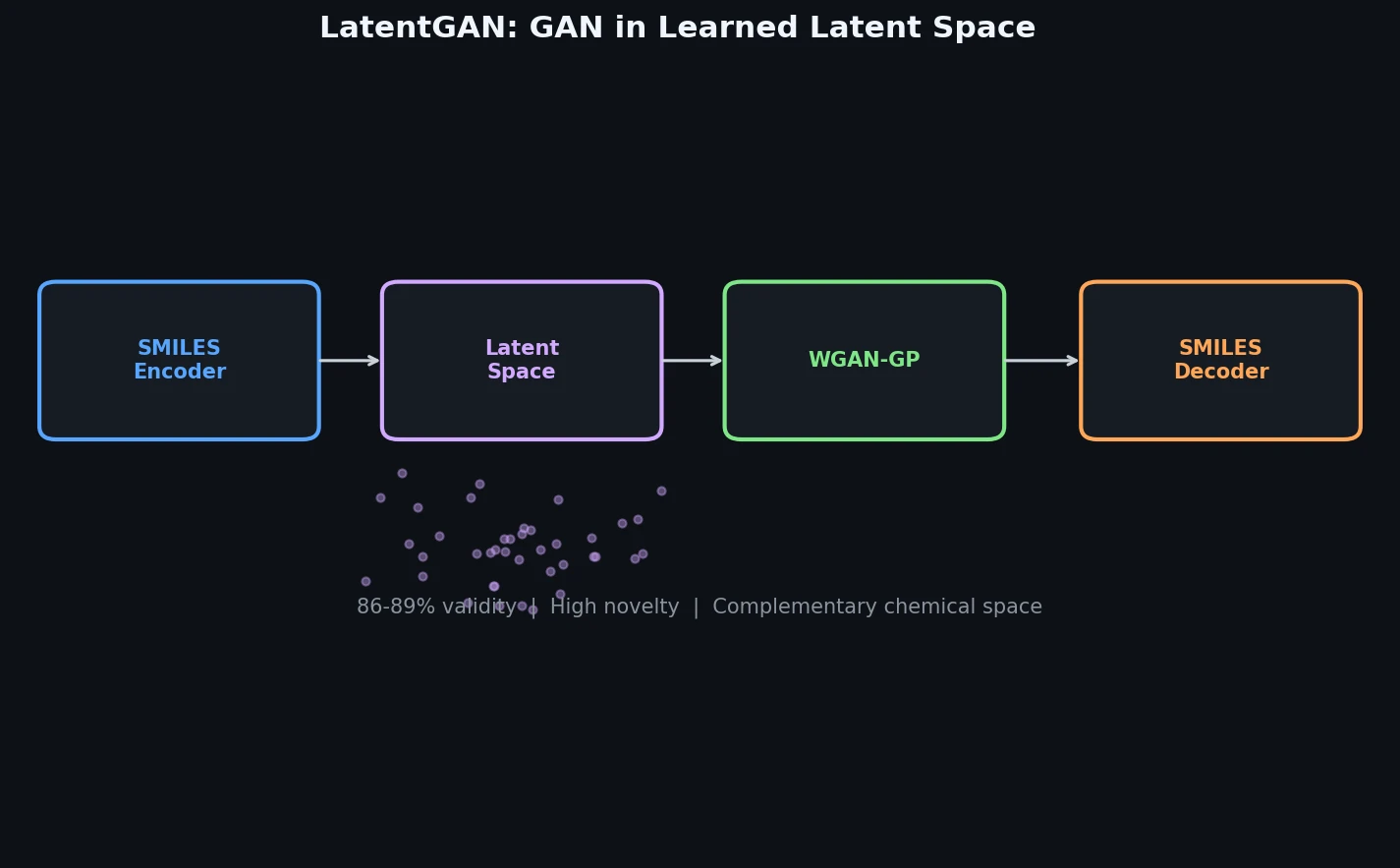
LatentGAN decouples molecular generation from SMILES syntax by training a Wasserstein GAN on latent vectors from a pretrained heteroencoder, enabling de novo design of drug-like and target-biased compounds.
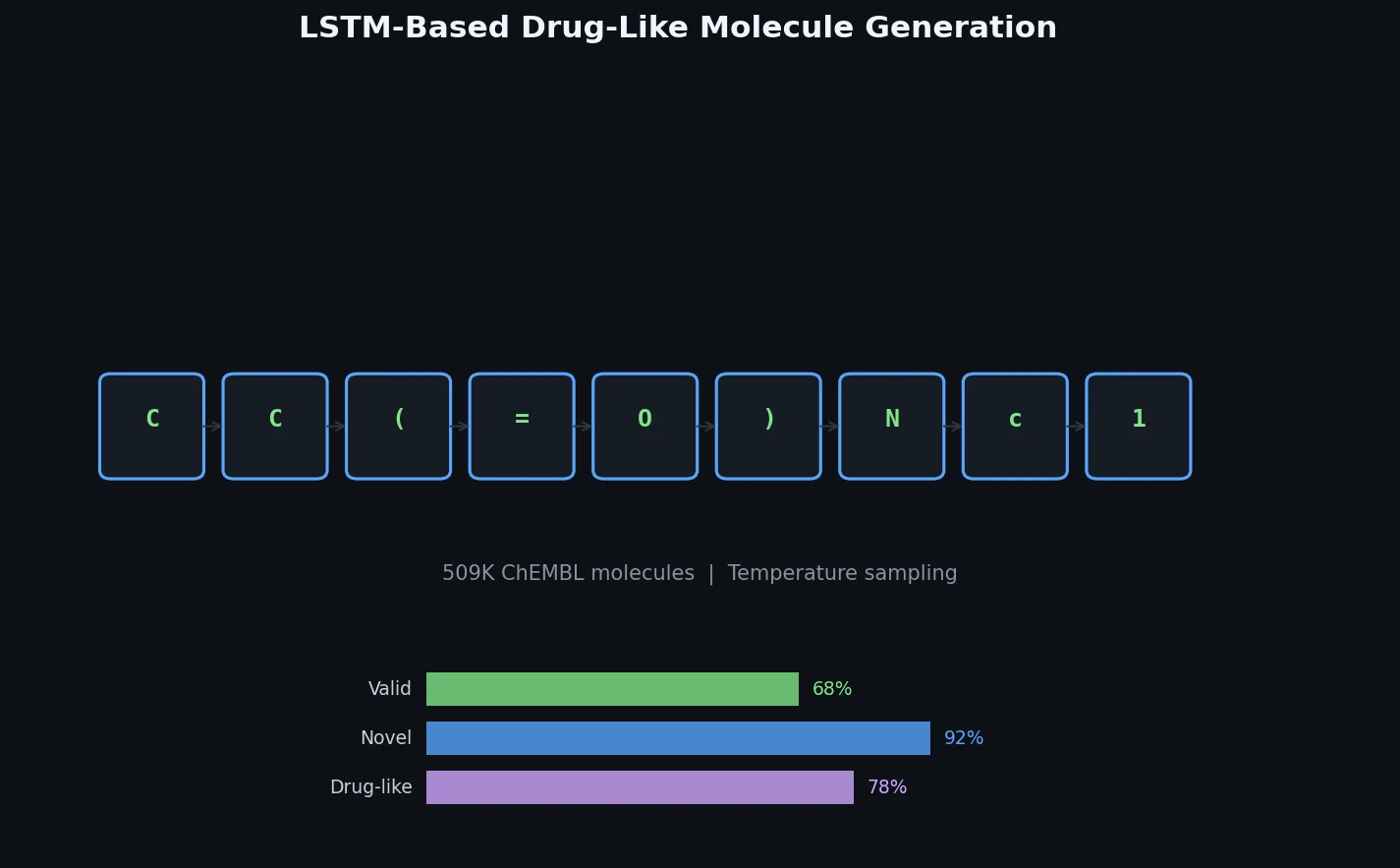
Ertl et al. train a character-level LSTM on 509K bioactive ChEMBL SMILES and generate one million novel, diverse molecules whose physicochemical properties, substructure features, and predicted bioactivity closely match the training distribution.
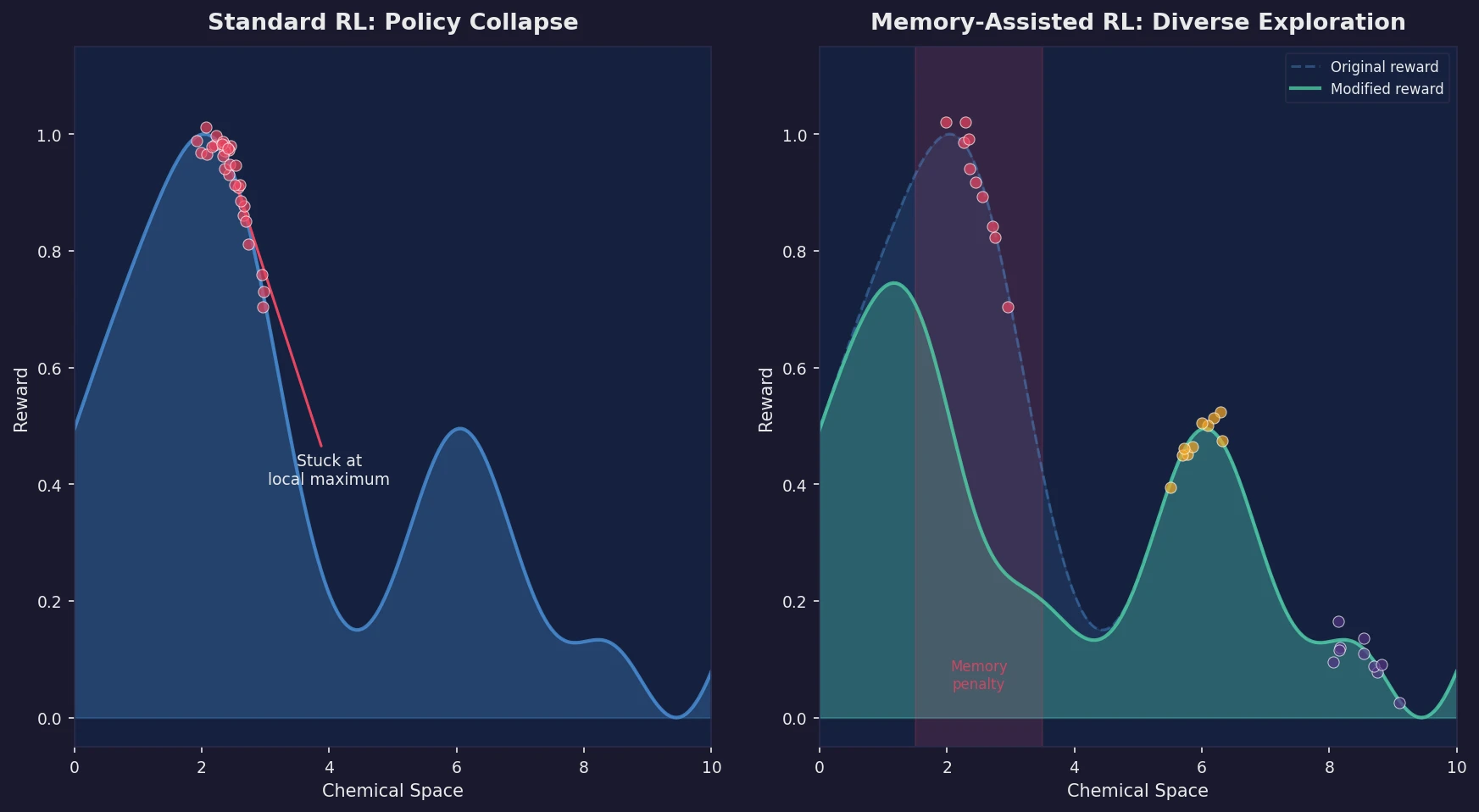
Introduces a memory unit that modifies the RL reward function to penalize previously explored chemical scaffolds, substantially increasing the diversity of generated molecules while maintaining relevance to known active ligands.
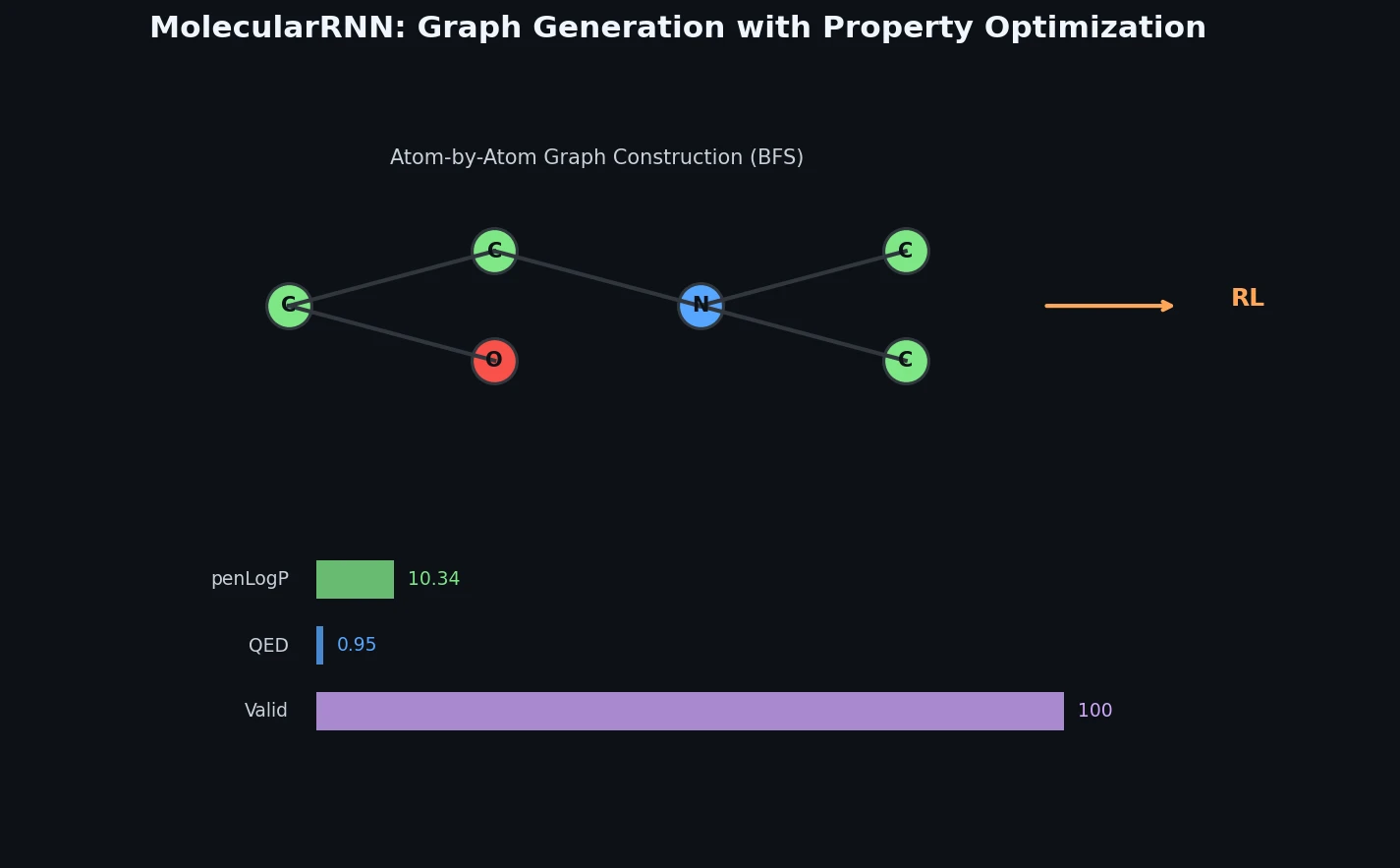
Proposes MolecularRNN, a graph recurrent model that generates molecular graphs atom-by-atom with 100% validity via valency-based rejection sampling, then shifts property distributions using policy gradient reinforcement learning.