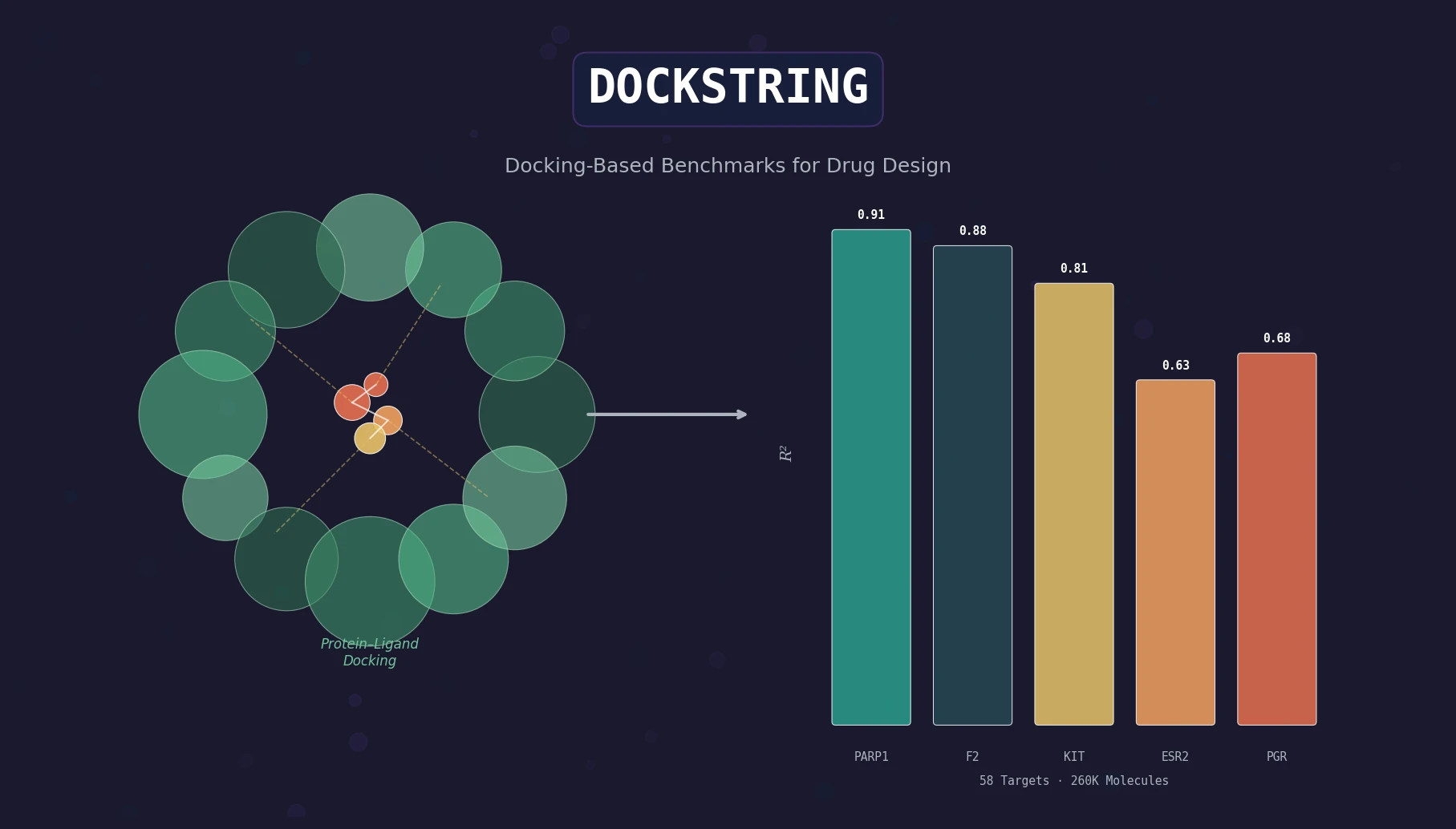
DOCKSTRING: Docking-Based Benchmarks for Drug Design
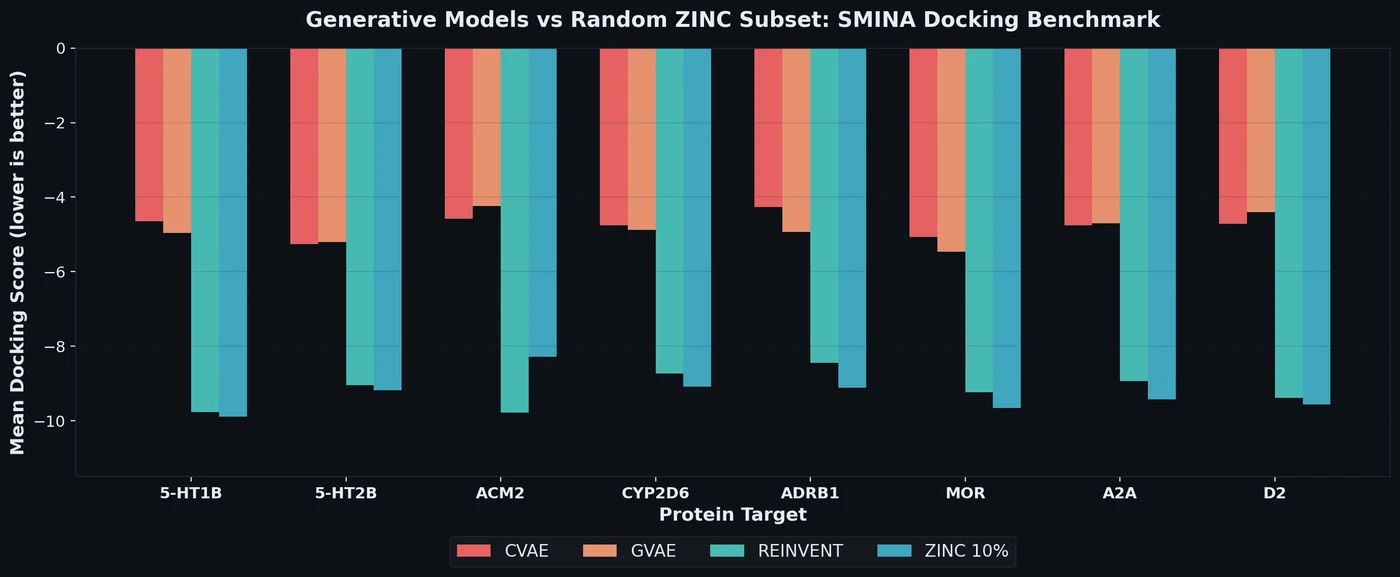
DOCKSTRING bundles an AutoDock Vina wrapper, a 260K-molecule docking dataset across 58 protein targets, and pharmaceutically relevant benchmarks for regression, virtual screening, and de novo design.