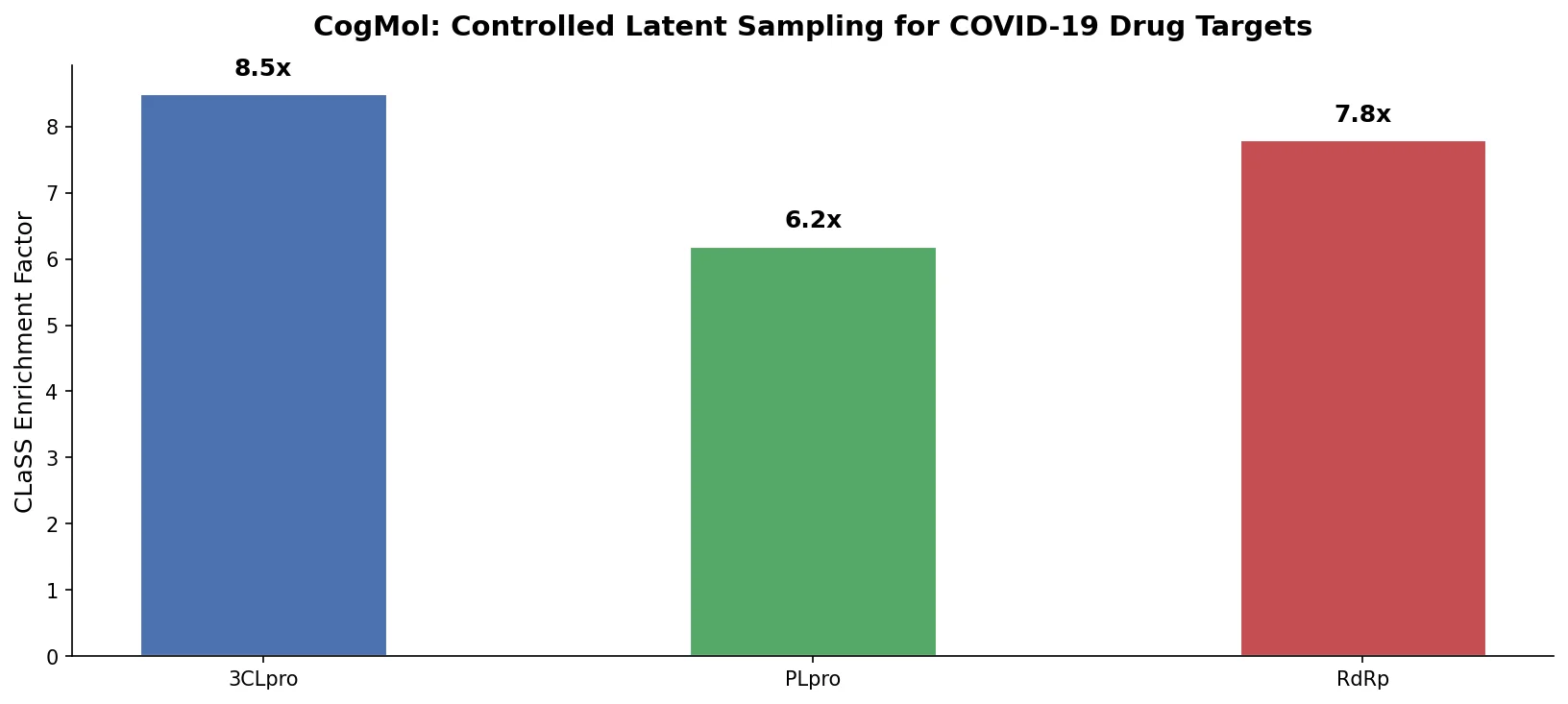
CogMol: Controlled Molecule Generation for COVID-19
CogMol uses a SMILES VAE and multi-attribute controlled sampling (CLaSS) to generate novel, target-specific drug molecules for unseen SARS-CoV-2 proteins without model retraining.

CogMol uses a SMILES VAE and multi-attribute controlled sampling (CLaSS) to generate novel, target-specific drug molecules for unseen SARS-CoV-2 proteins without model retraining.
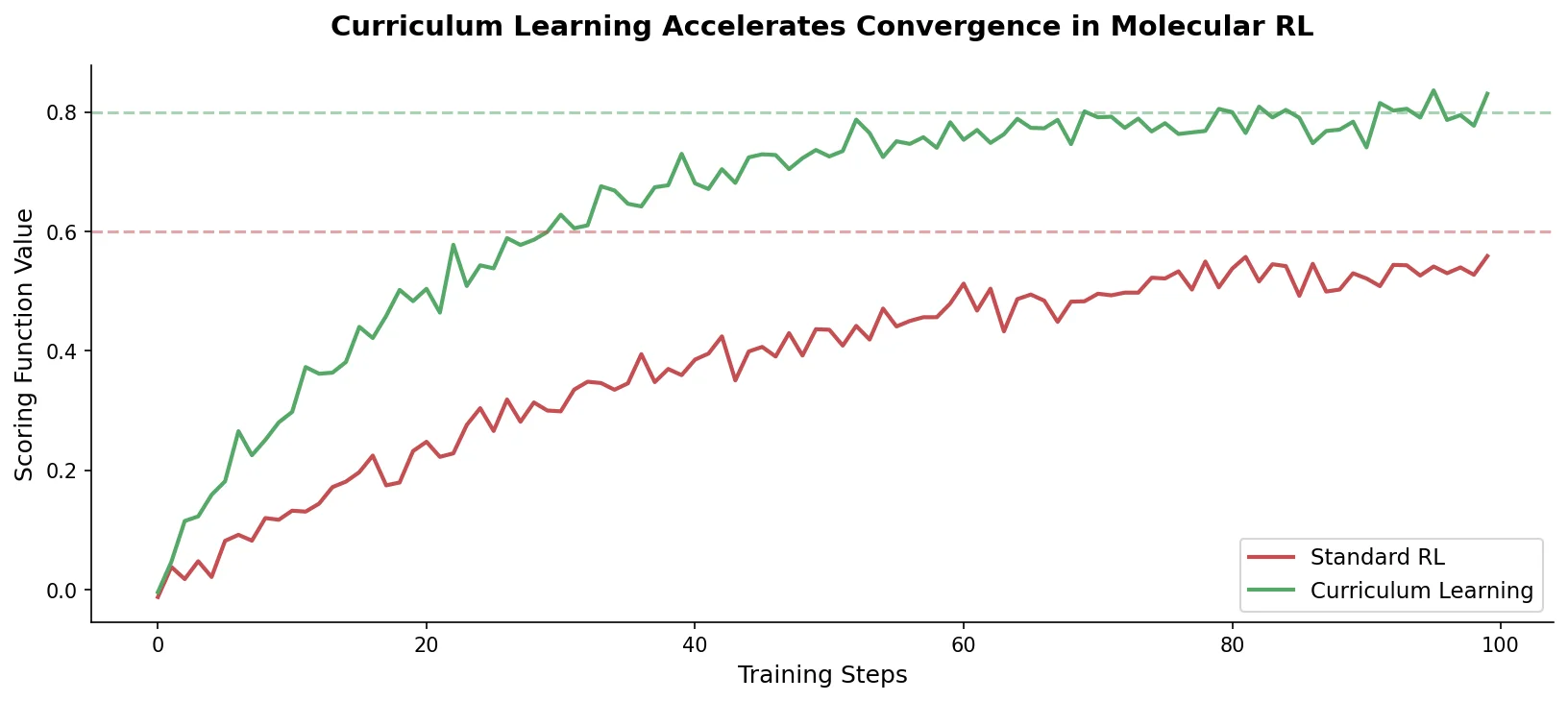
Introduces curriculum learning to the REINVENT de novo design platform, decomposing complex drug design objectives into simpler sequential tasks that accelerate agent convergence and improve output quality over standard reinforcement learning.
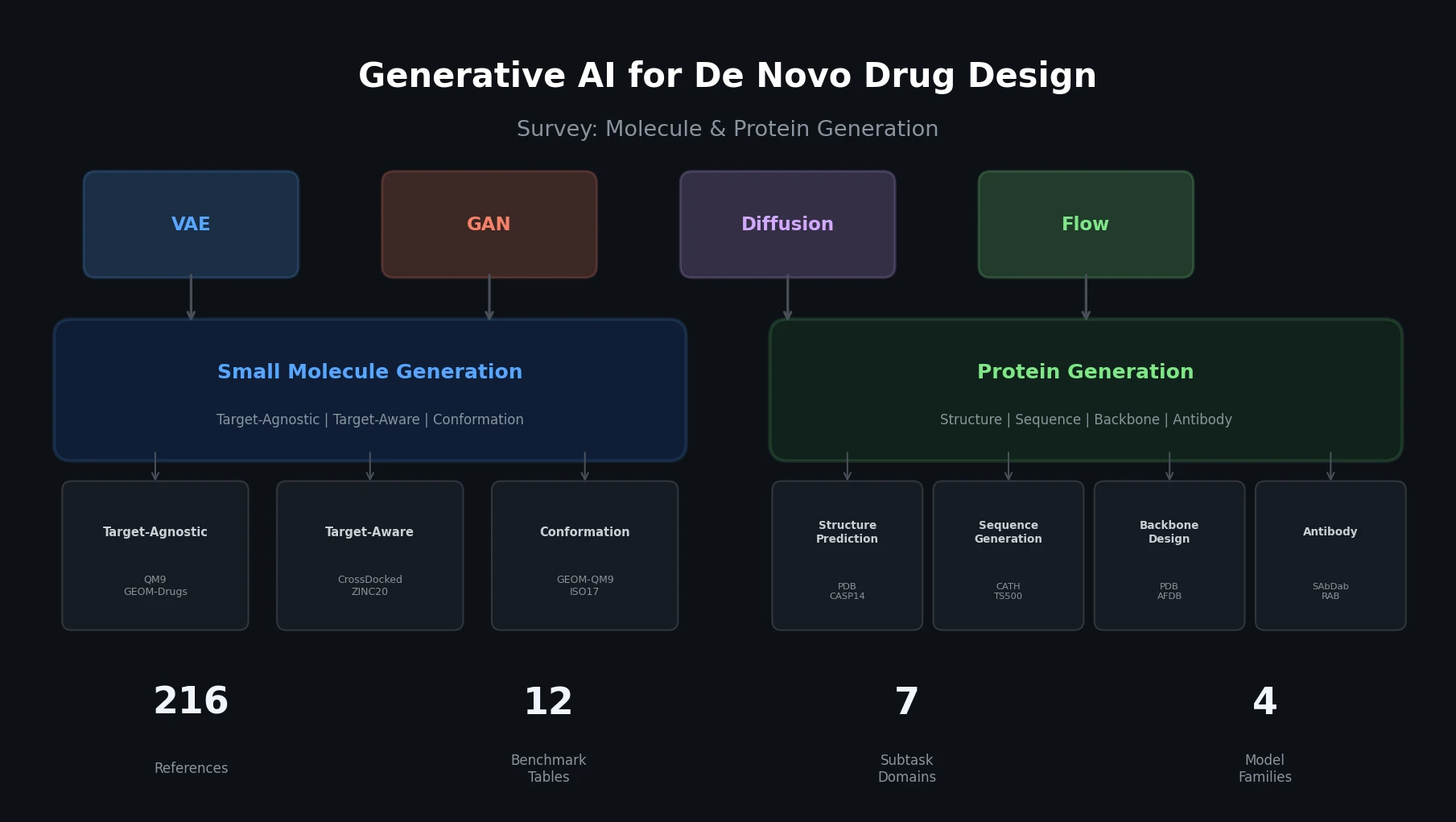
This survey organizes generative AI for de novo drug design into two themes: small molecule generation (target-agnostic, target-aware, conformation) and protein generation (structure prediction, sequence generation, backbone design, antibody, peptide). It covers four generative model families (VAEs, GANs, diffusion, flow-based), catalogs key datasets and benchmarks, and provides 12 comparative benchmark tables across all subtasks.
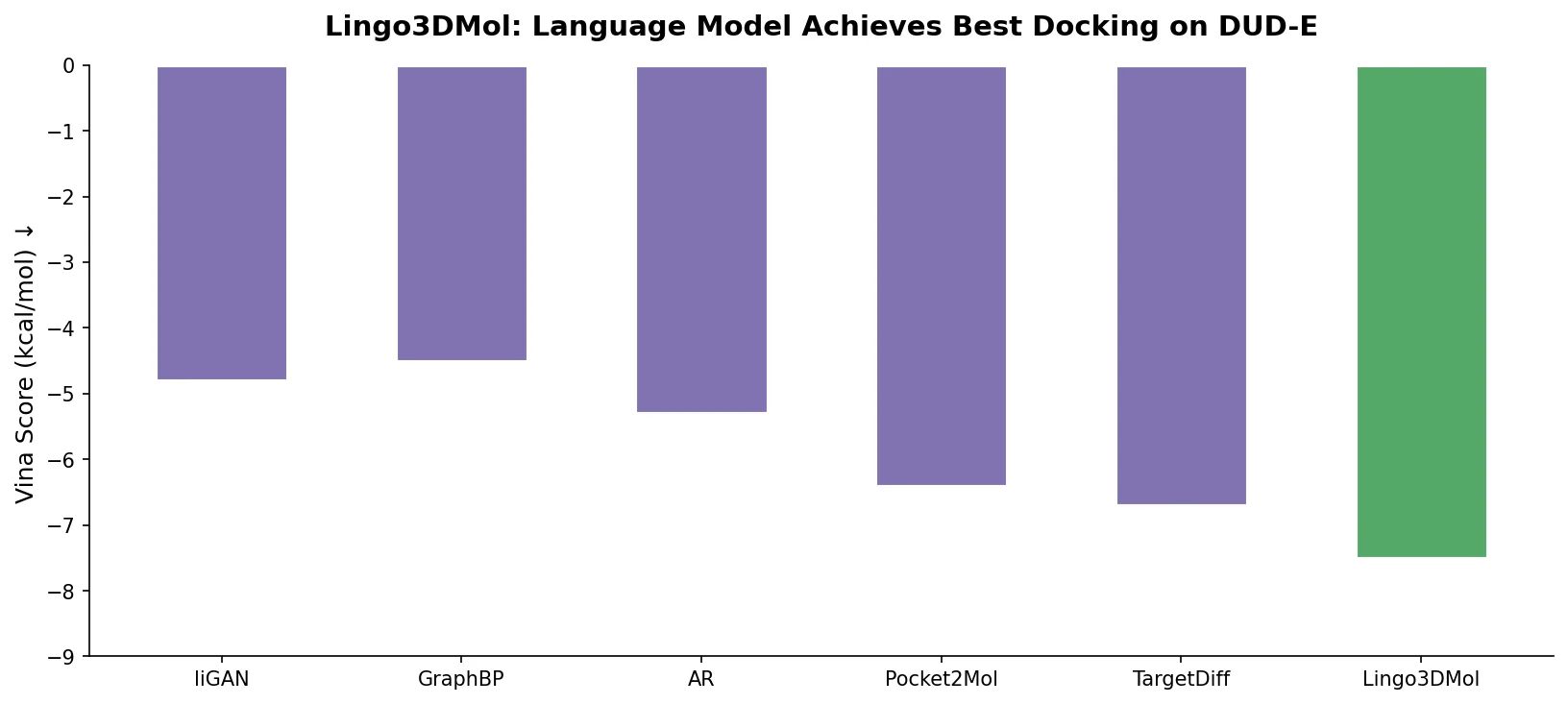
Lingo3DMol introduces FSMILES, a fragment-based SMILES representation with local and global coordinates, to generate drug-like 3D molecules in protein pockets via a transformer language model.
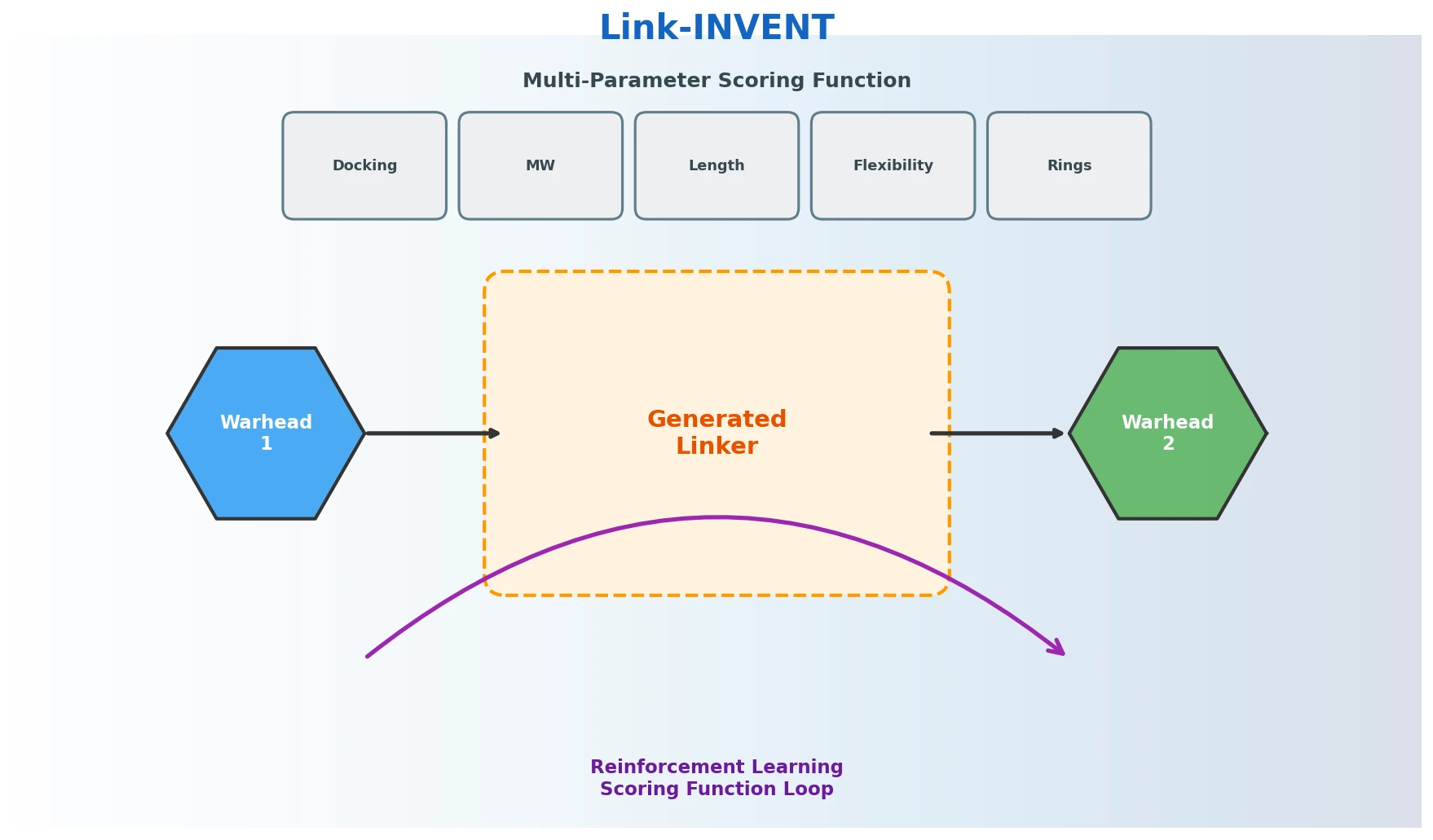
Link-INVENT is an RNN-based generative model for molecular linker design that uses reinforcement learning with a flexible scoring function, demonstrated on fragment linking, scaffold hopping, and PROTAC design.
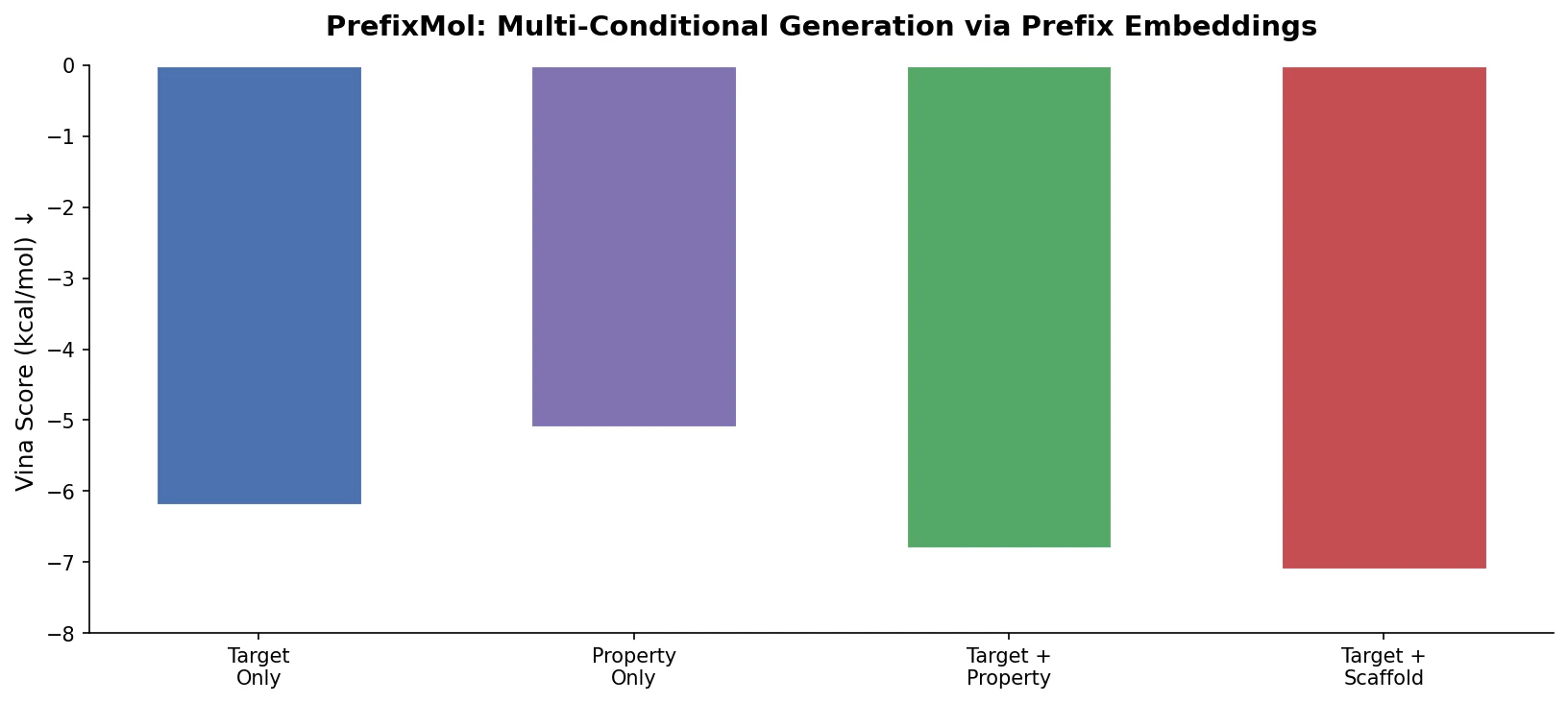
PrefixMol prepends learnable condition vectors to a GPT transformer for SMILES generation, enabling joint control over binding pocket targeting and chemical properties like QED, SA, and LogP.
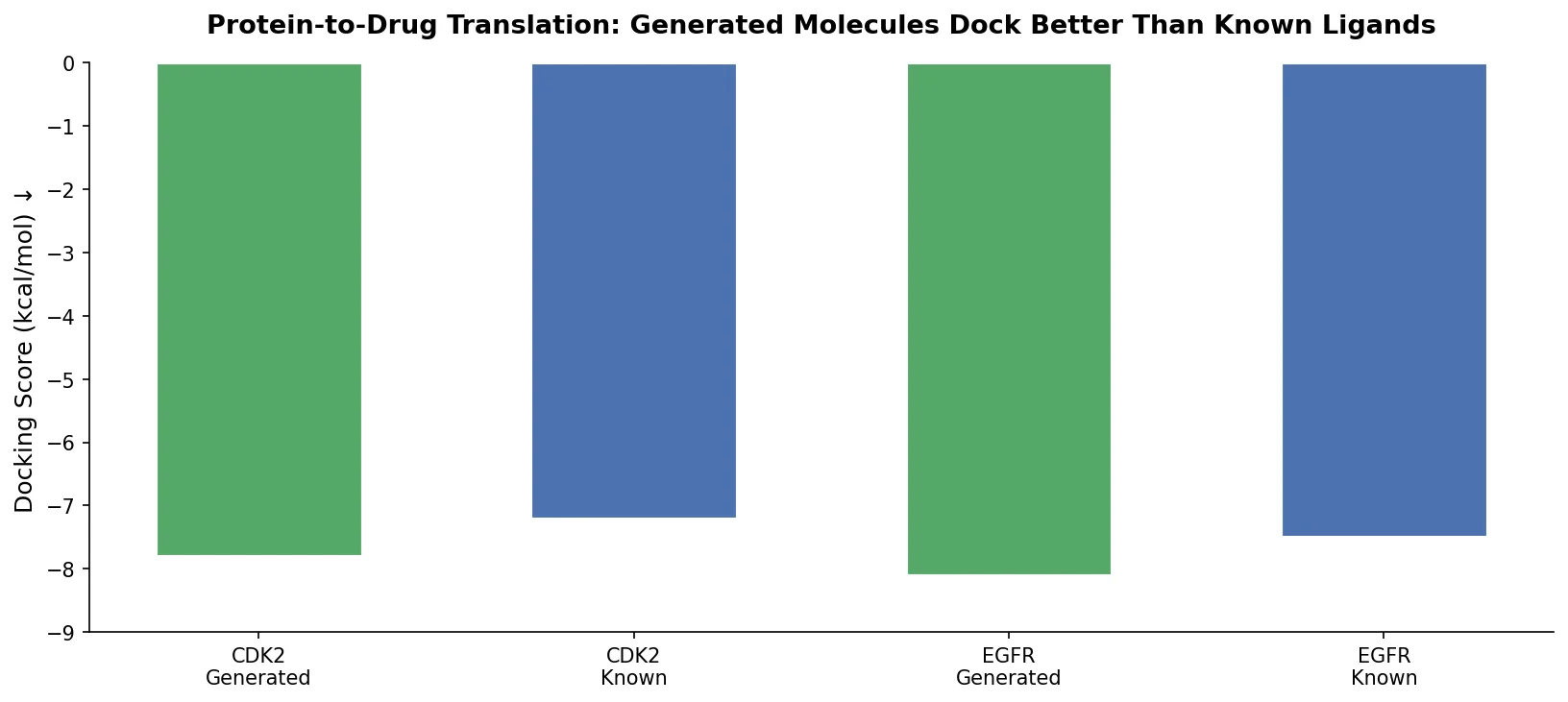
Applies the Transformer architecture to generate drug-like molecules conditioned on protein amino acid sequences, treating target-specific de novo drug design as a sequence-to-sequence translation problem.
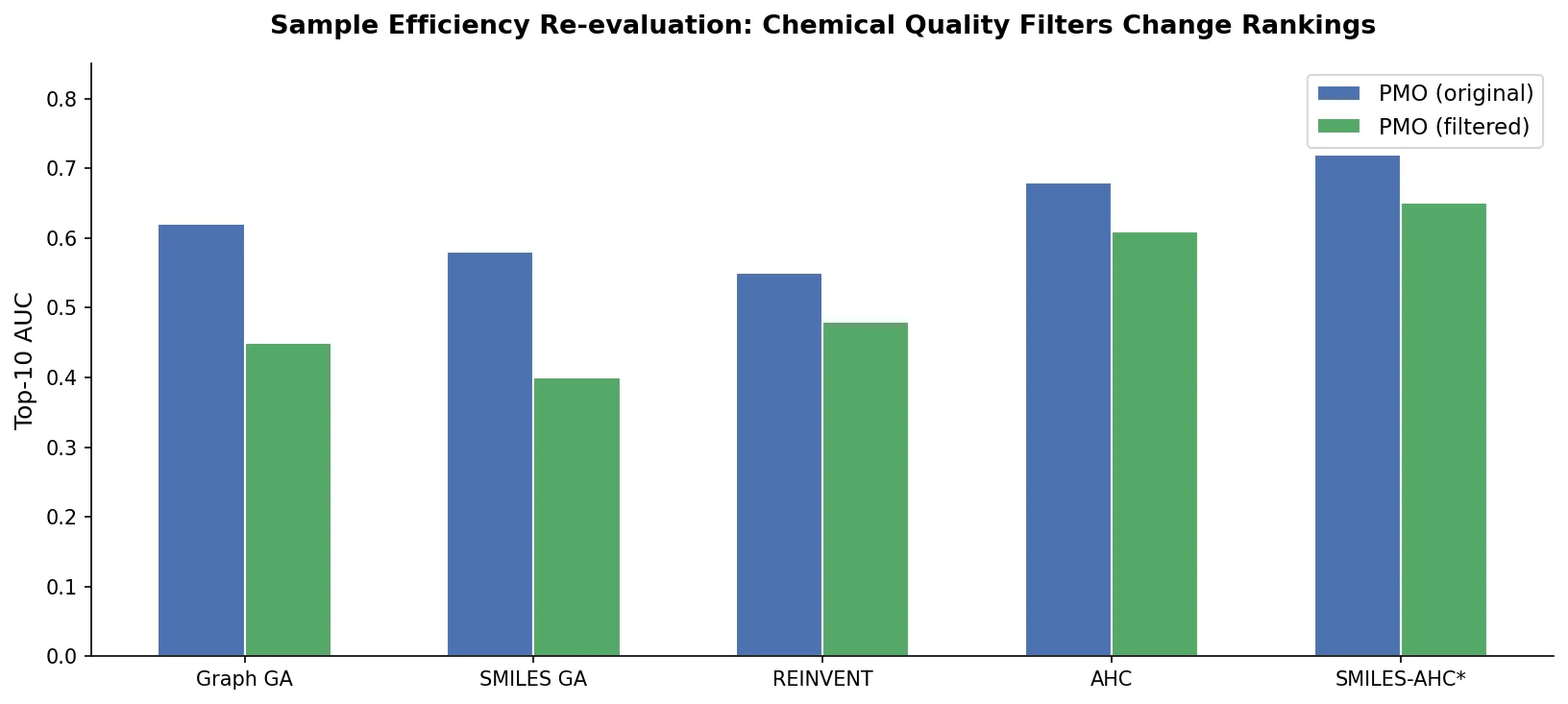
A critical reassessment of the PMO benchmark for de novo molecule generation, showing that adding molecular weight, LogP, and diversity filters substantially re-ranks generative models, with Augmented Hill-Climb emerging as the top method.
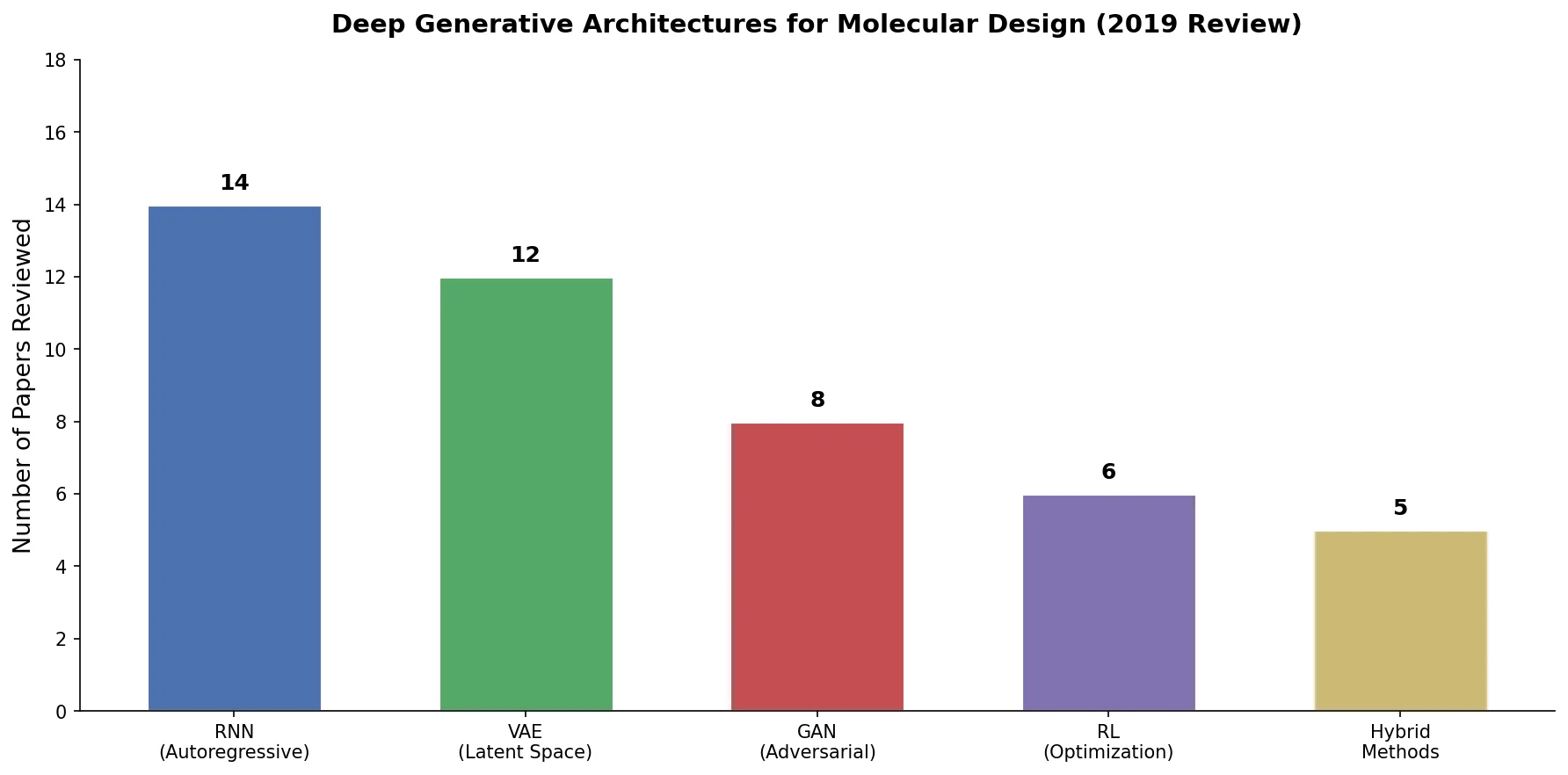
An early and influential review cataloging 45 papers on deep generative modeling for molecules, comparing RNN, VAE, GAN, and reinforcement learning architectures across SMILES and graph-based representations.
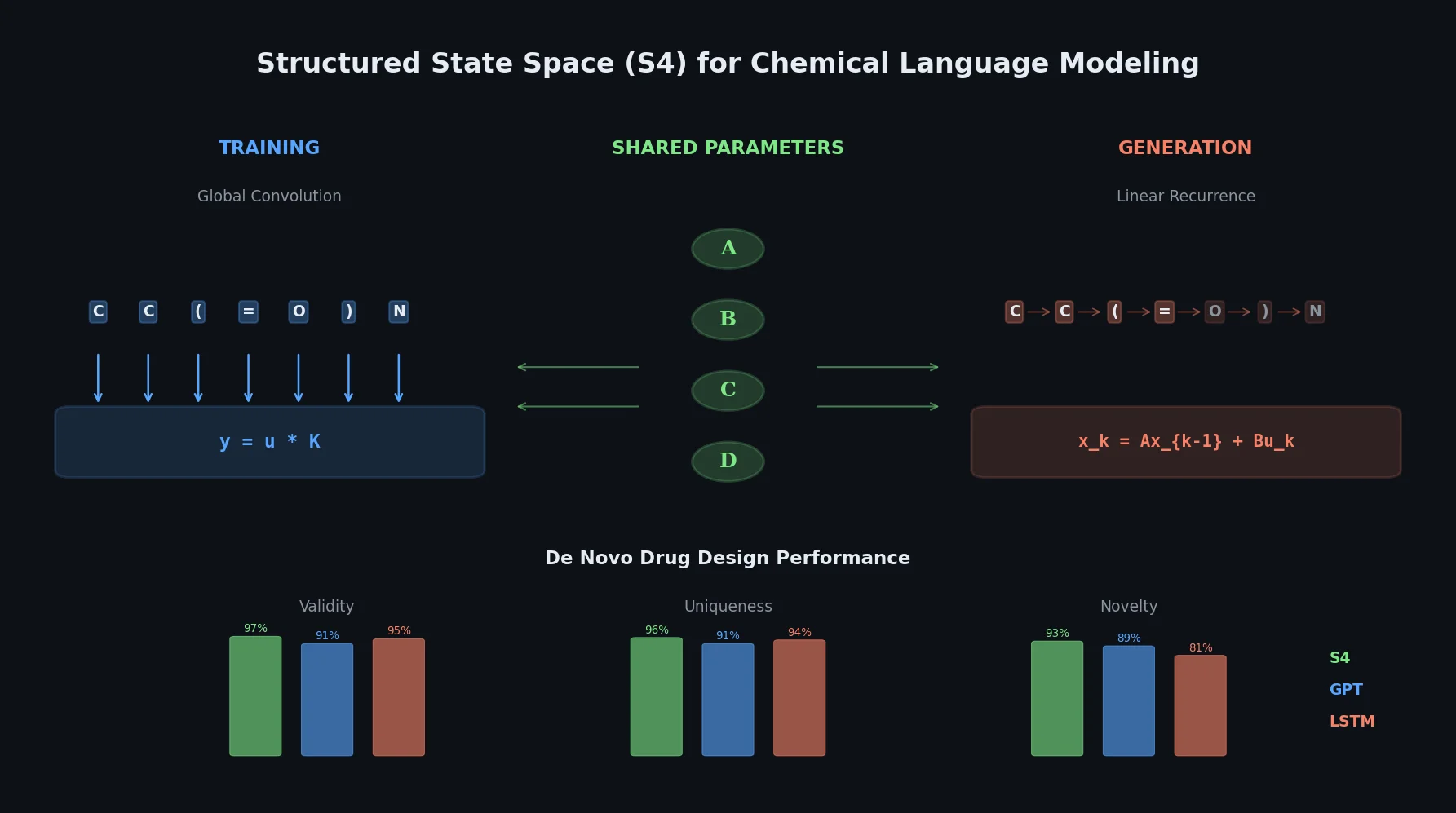
This paper introduces structured state space sequence (S4) models to chemical language modeling, showing they combine the strengths of LSTMs (efficient recurrent generation) and GPTs (holistic sequence learning) for de novo molecular design.
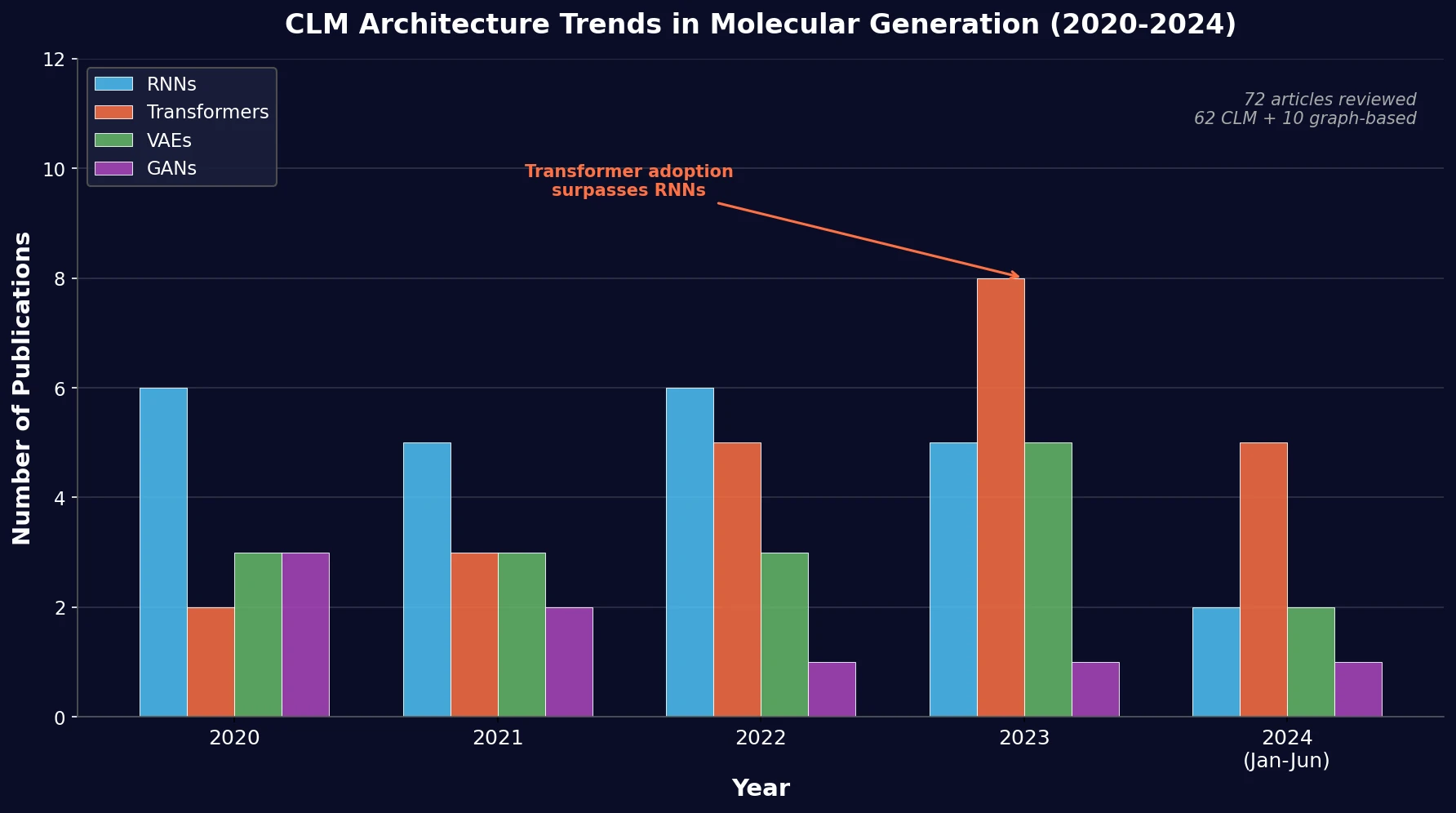
PRISMA-based systematic review of 72 papers on chemical language models for molecular generation, comparing architectures and biased methods using MOSES metrics.
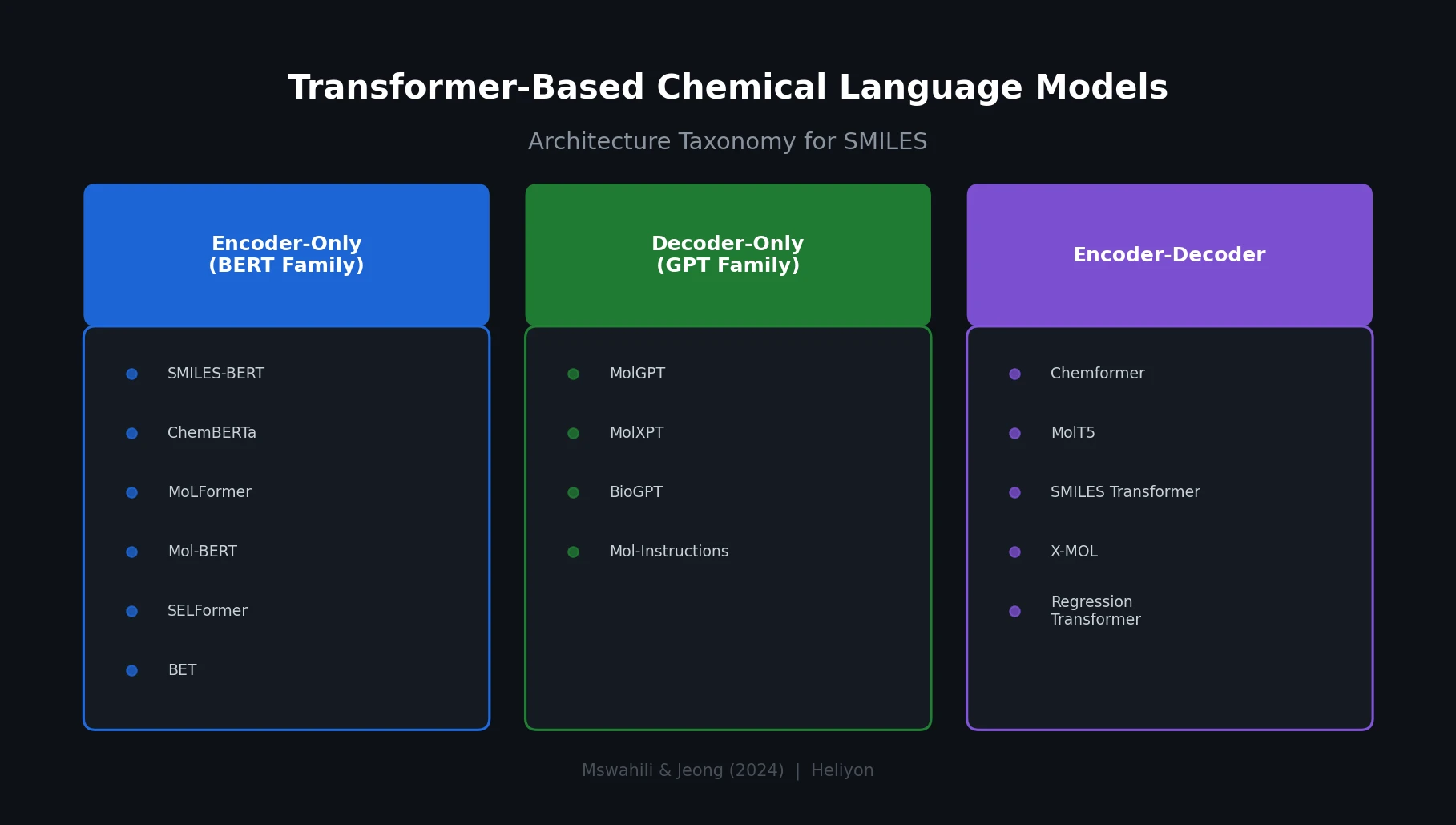
A comprehensive review of transformer-based chemical language models operating on SMILES, categorizing encoder-only (BERT variants), decoder-only (GPT variants), and encoder-decoder models with analysis of tokenization strategies, pre-training approaches, and future directions.