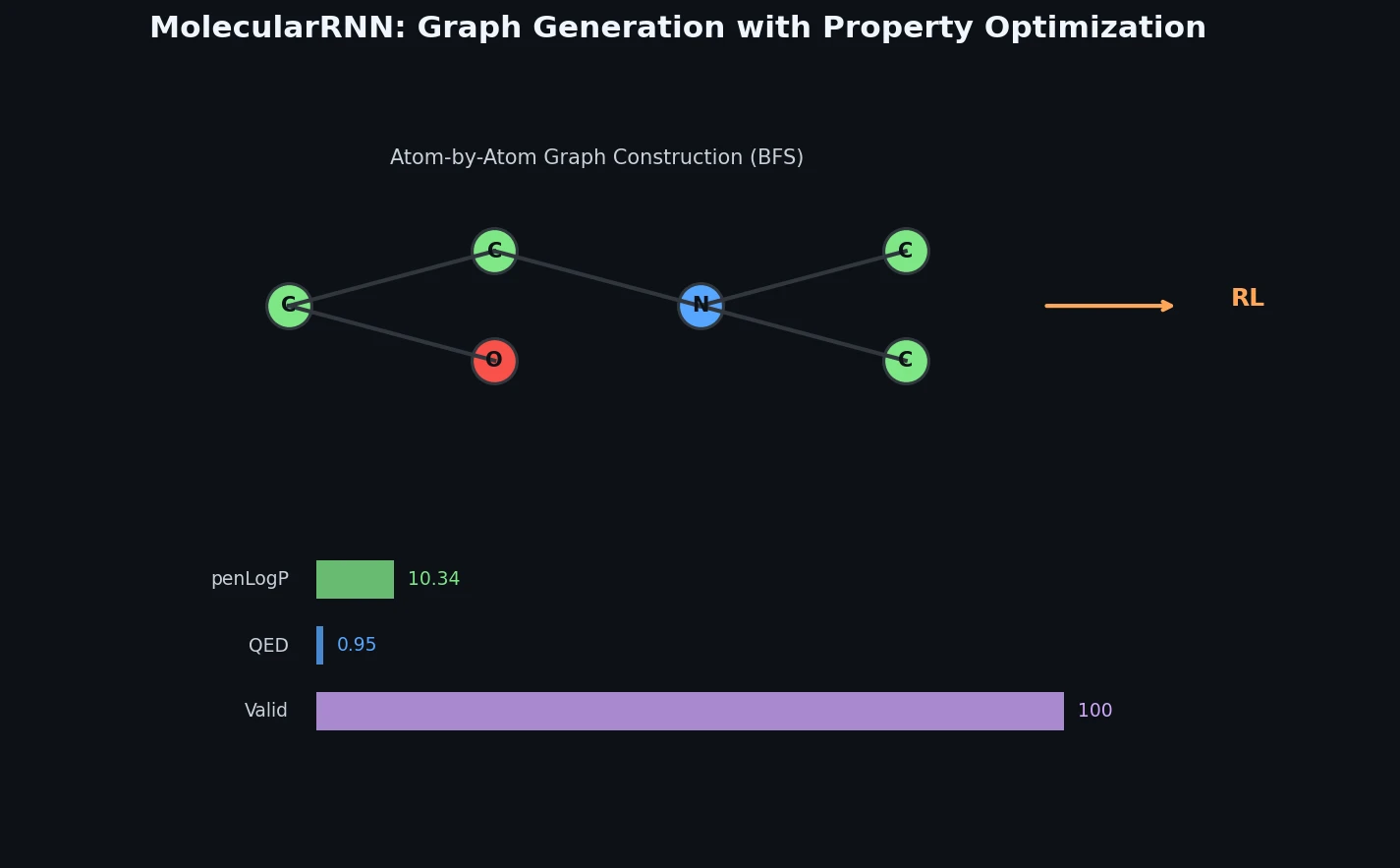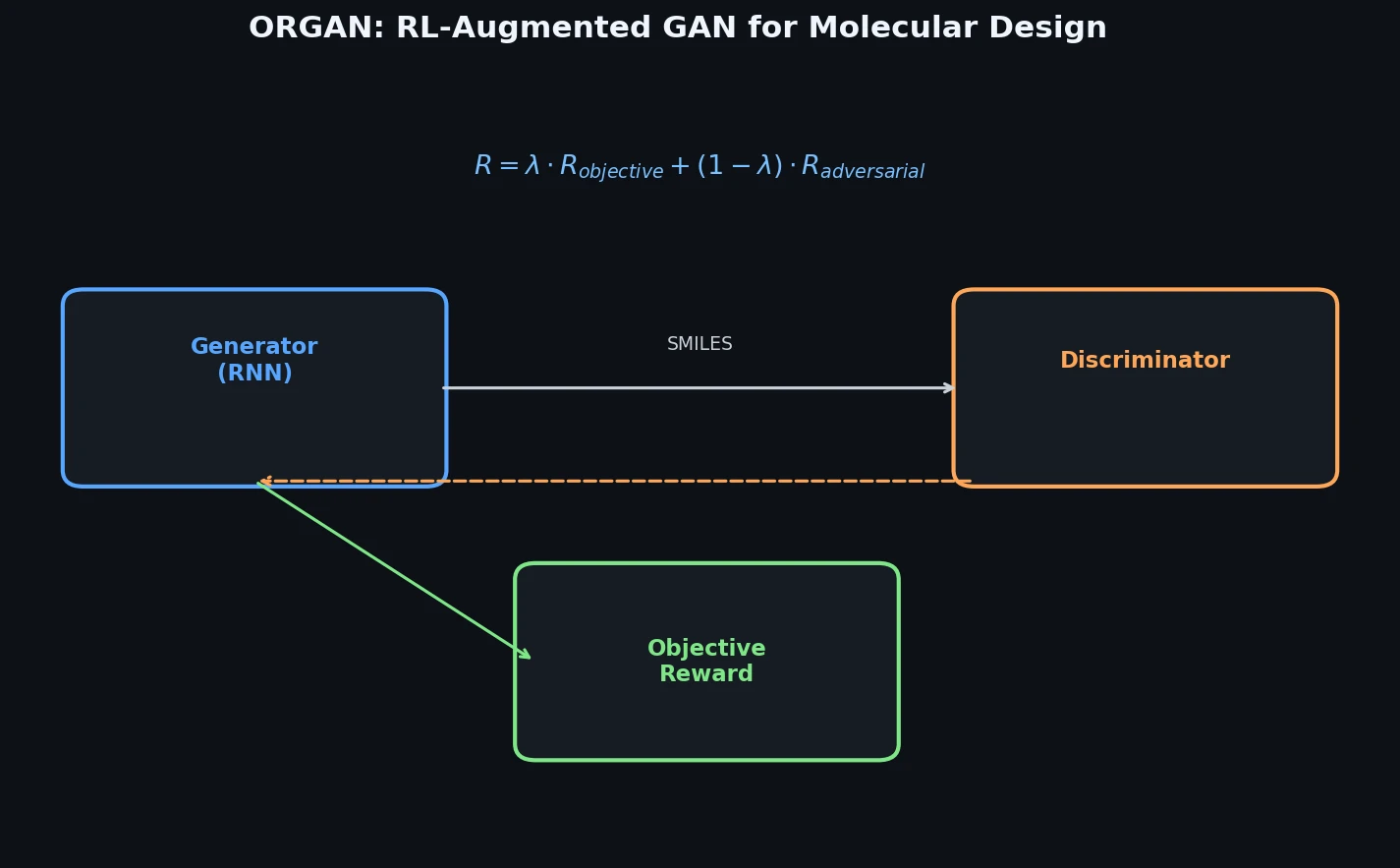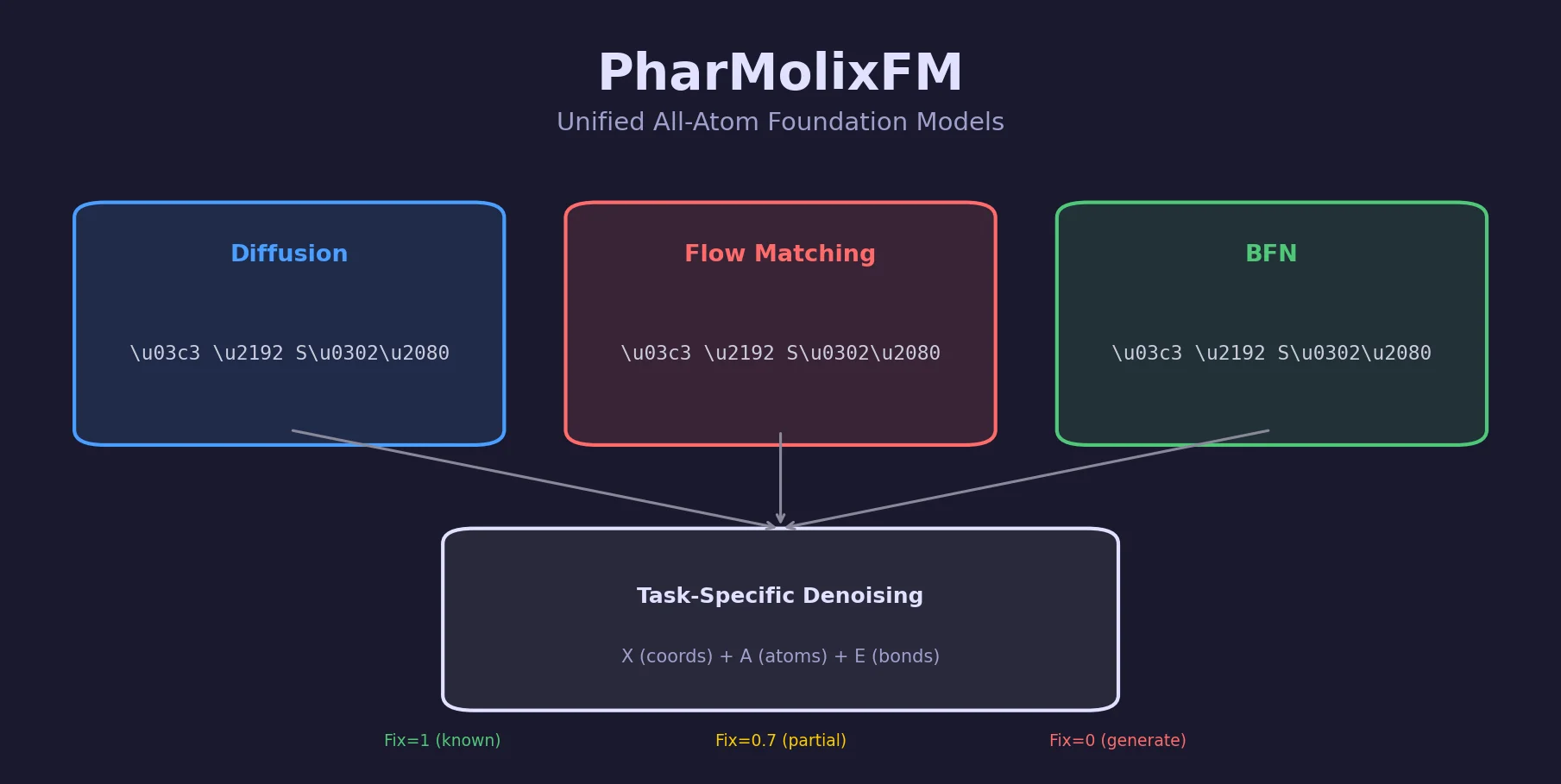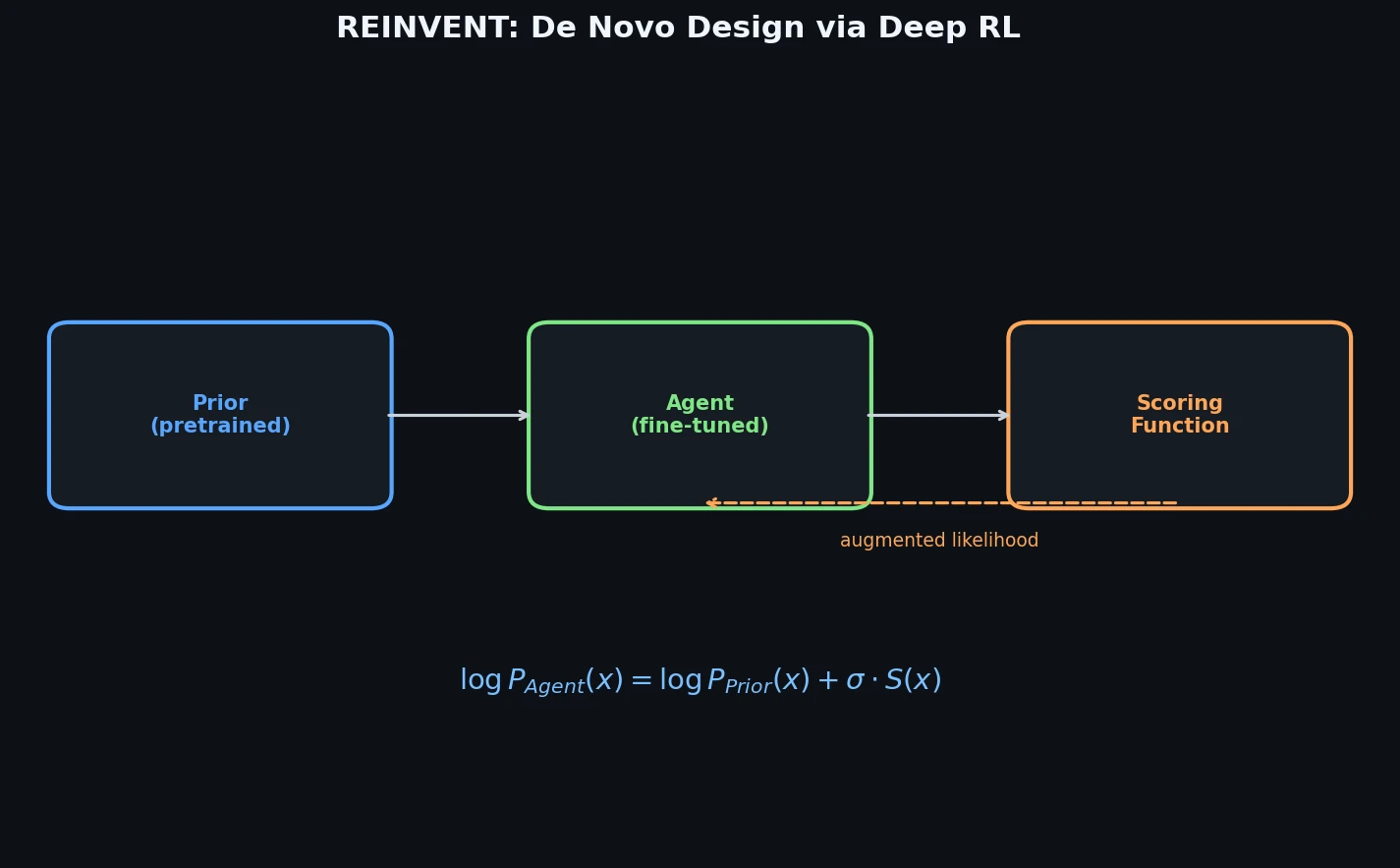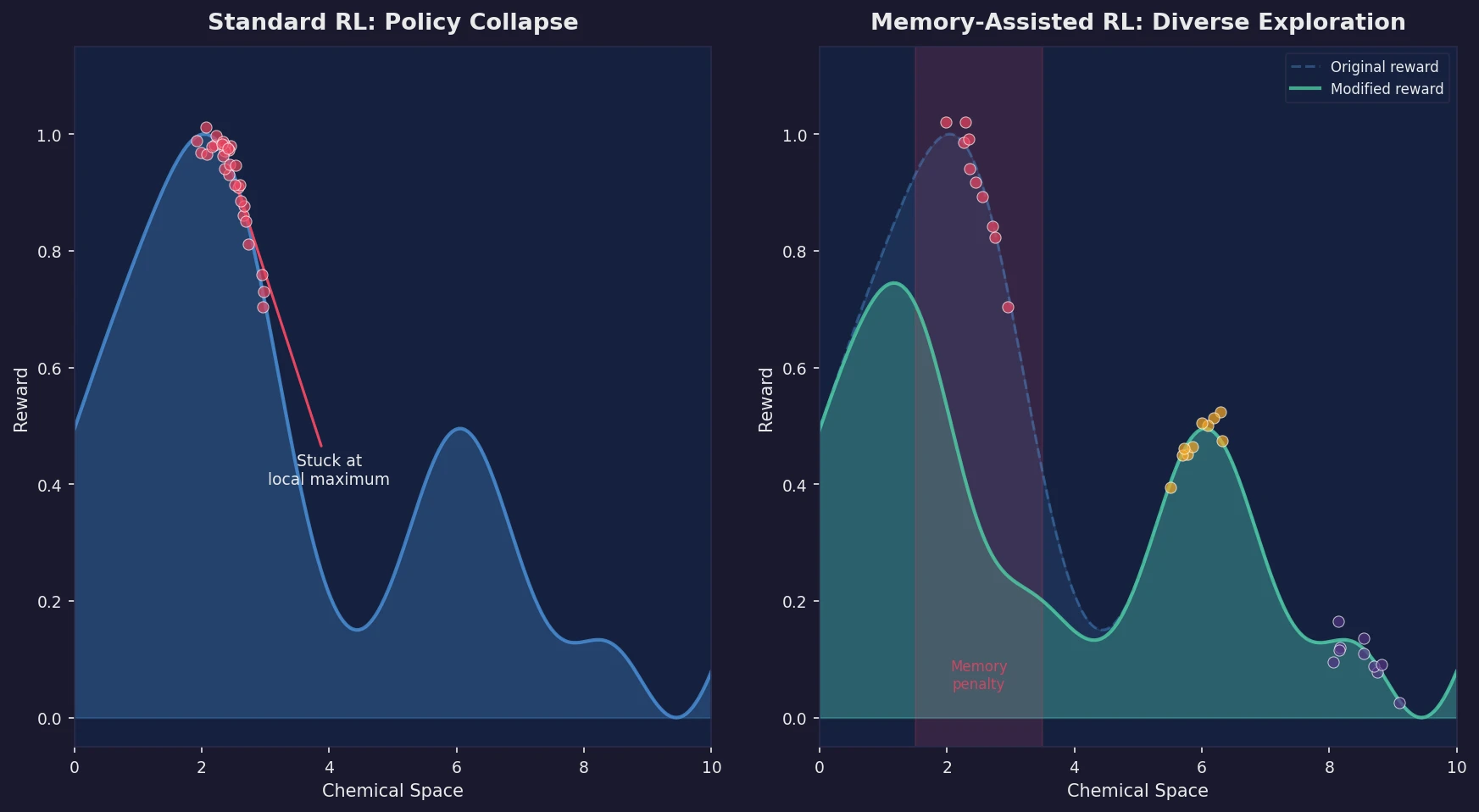
Memory-Assisted RL for Diverse De Novo Mol. Design
Introduces a memory unit that modifies the RL reward function to penalize previously explored chemical scaffolds, substantially increasing the diversity of generated molecules while maintaining relevance to known active ligands.