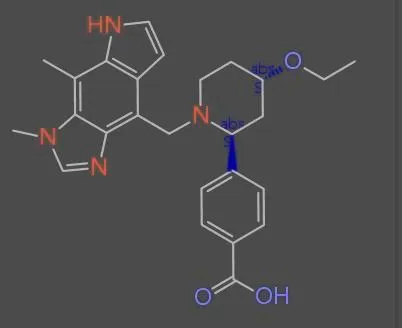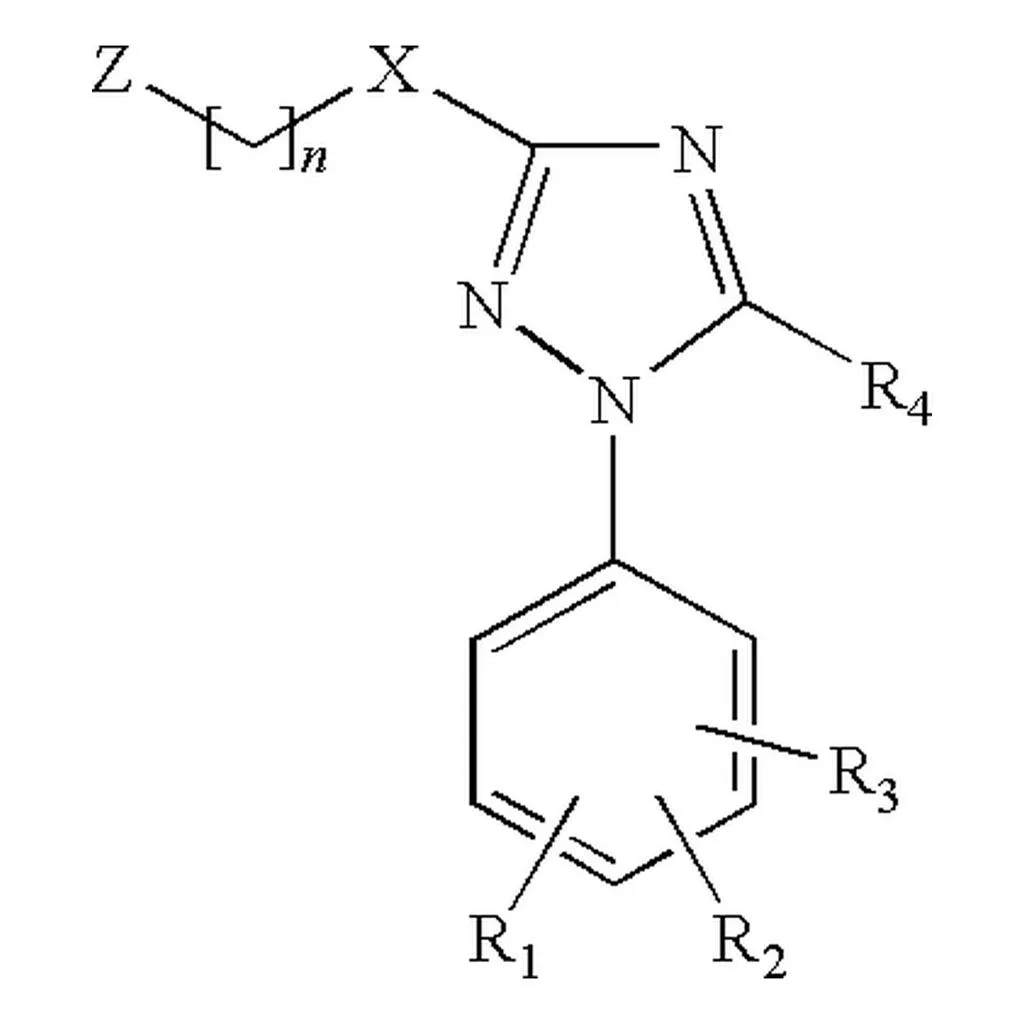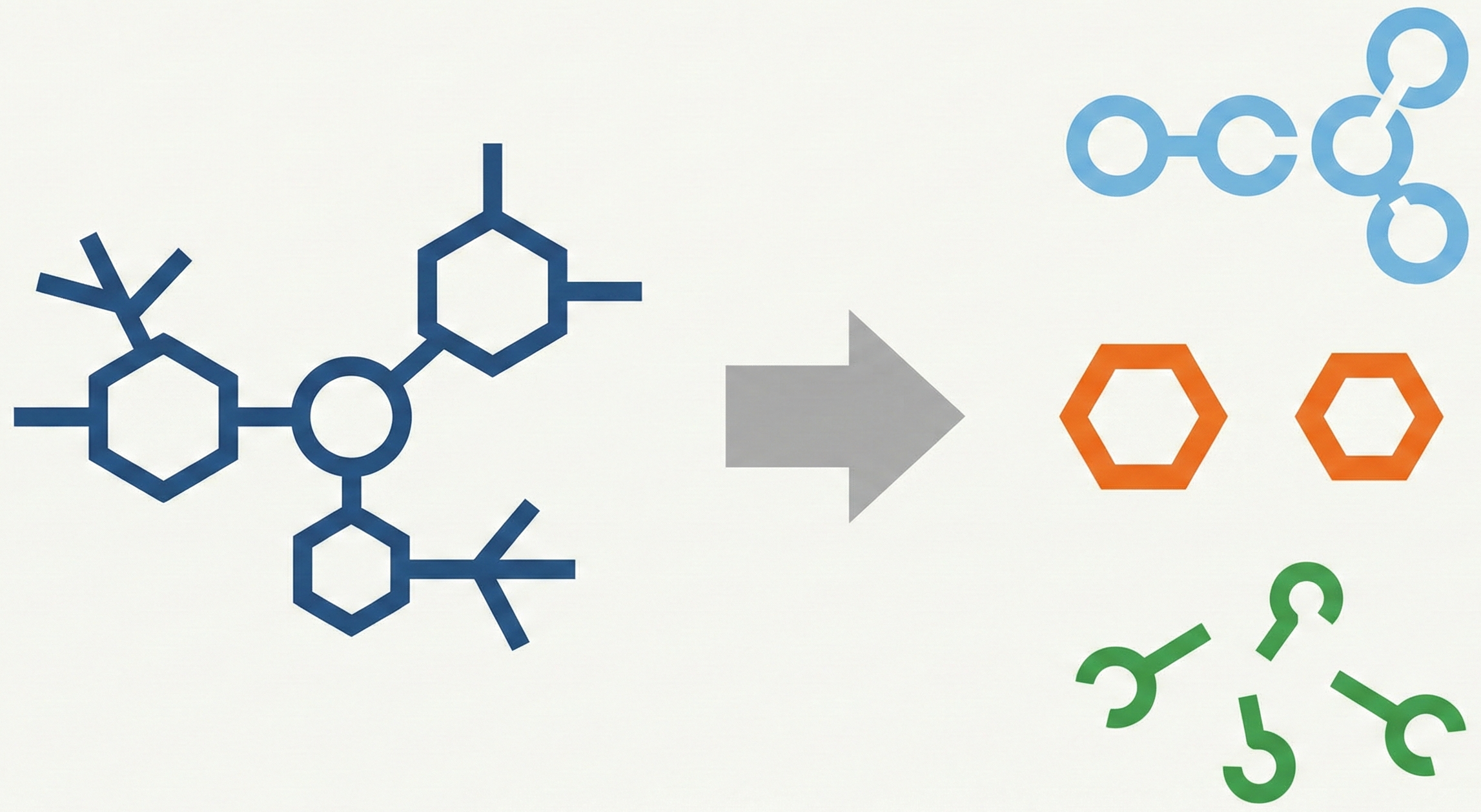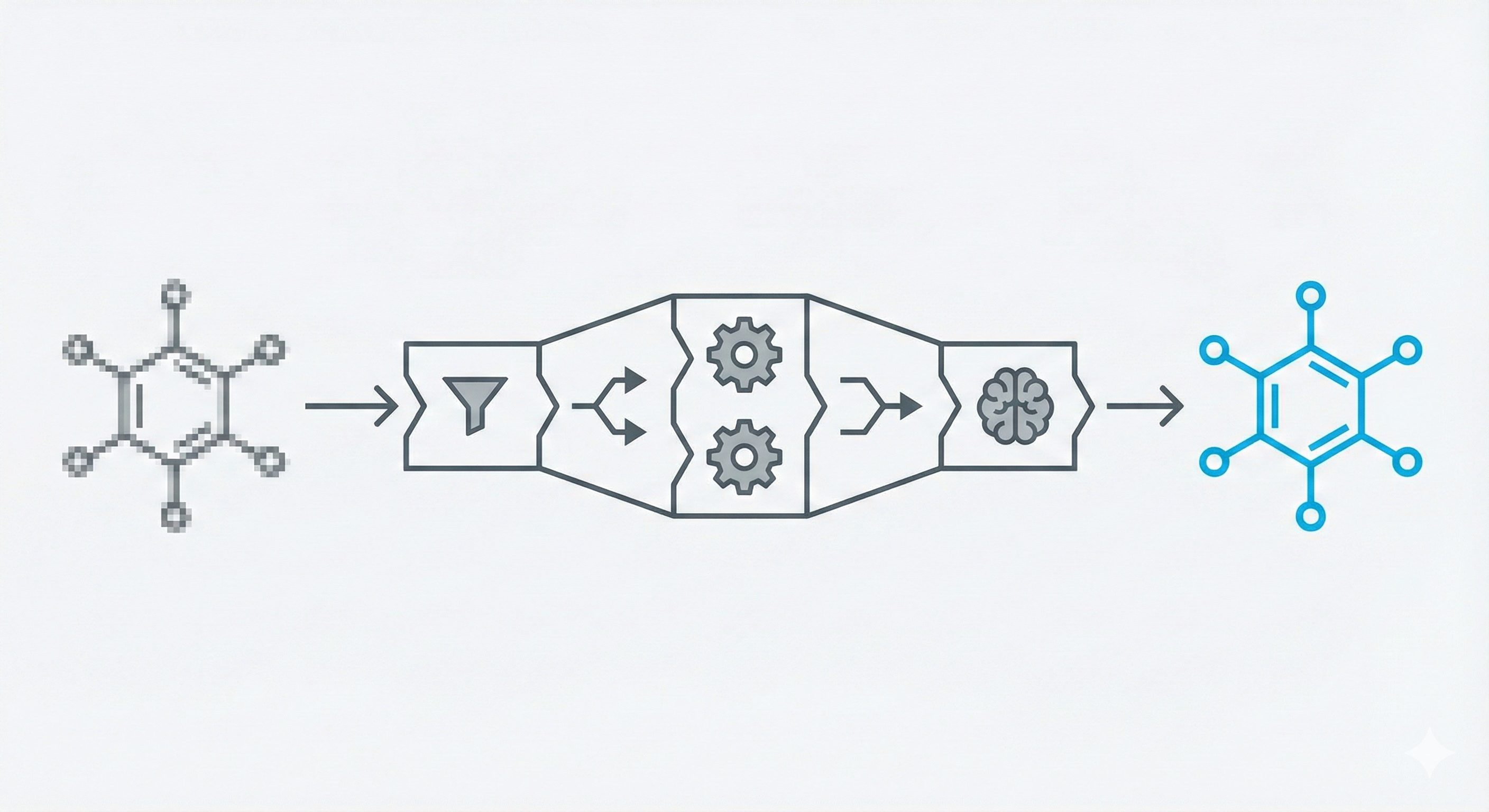
Reconstruction of Chemical Molecules from Images
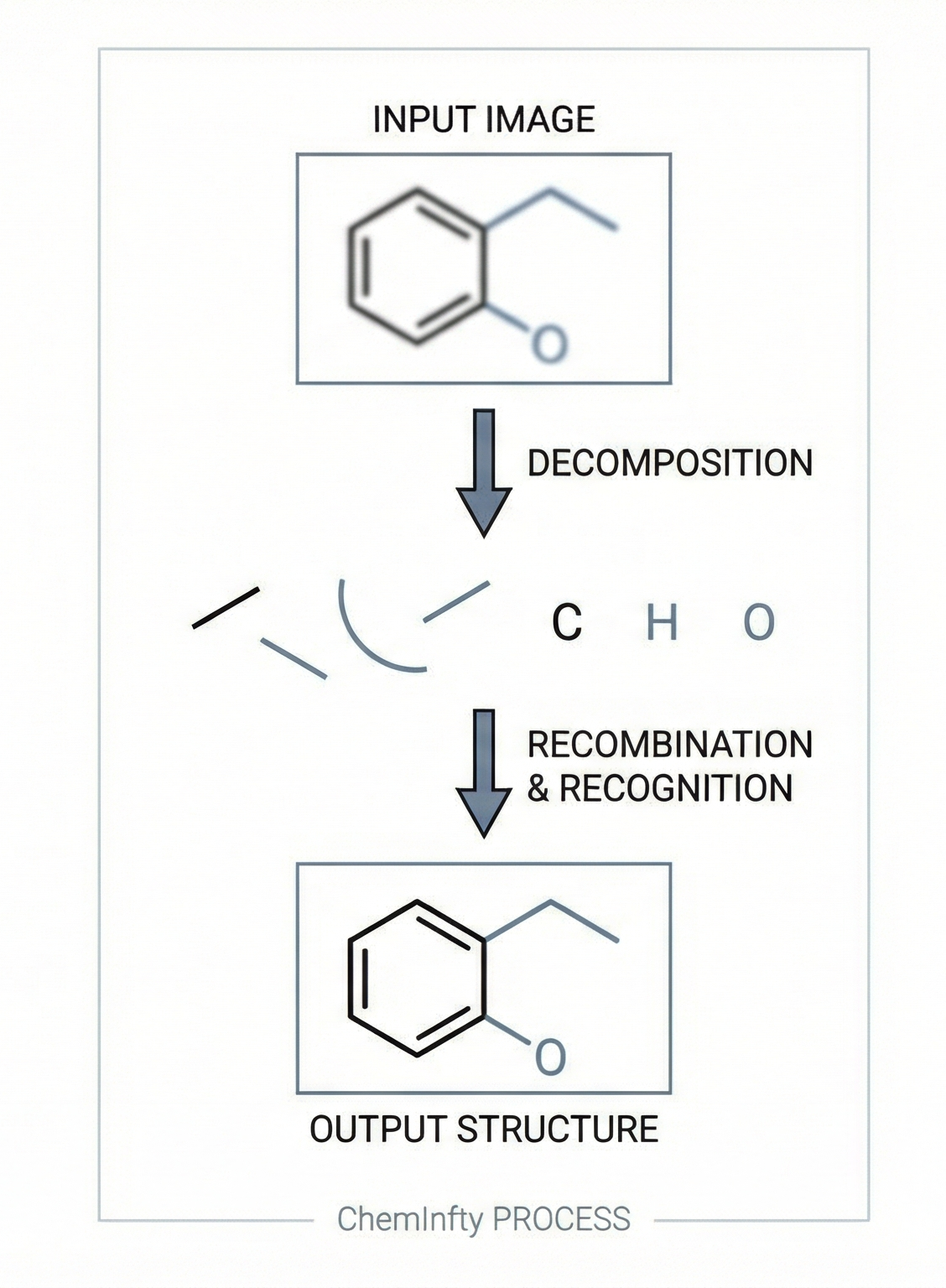
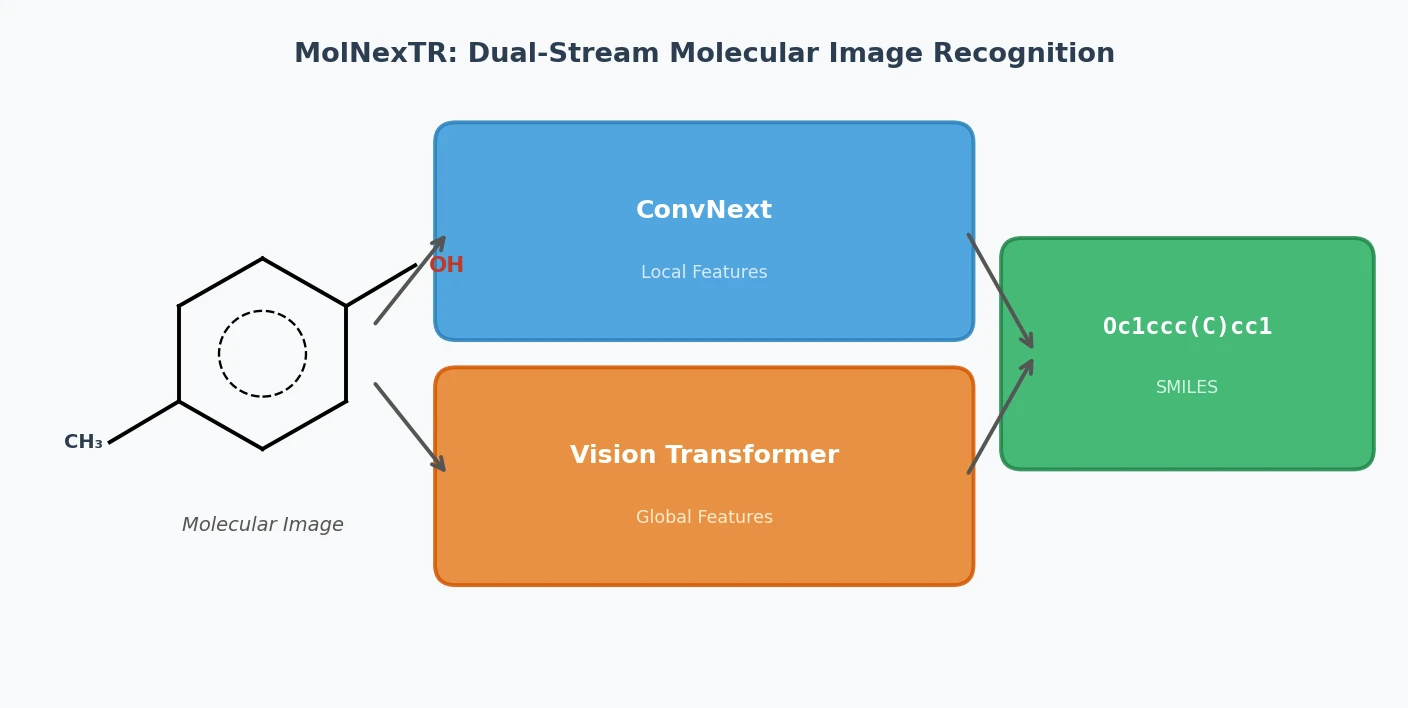
This methodological paper proposes a comprehensive pipeline to digitize chemical structure images. It achieves 97% reconstruction accuracy on benchmarks by combining a topology-preserving vectorizer with a chemical knowledge validation module.