CLiDE Pro: Optical Chemical Structure Recognition Tool
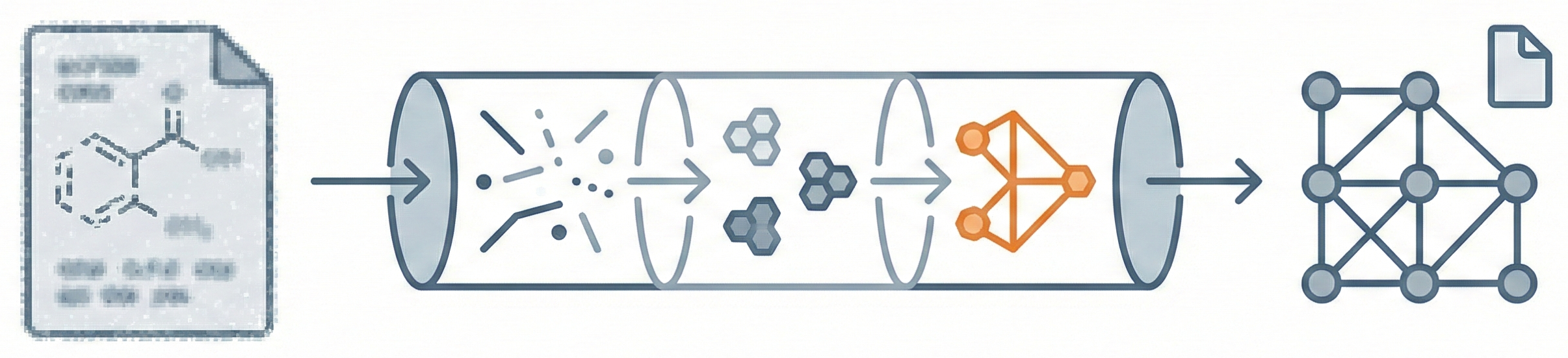
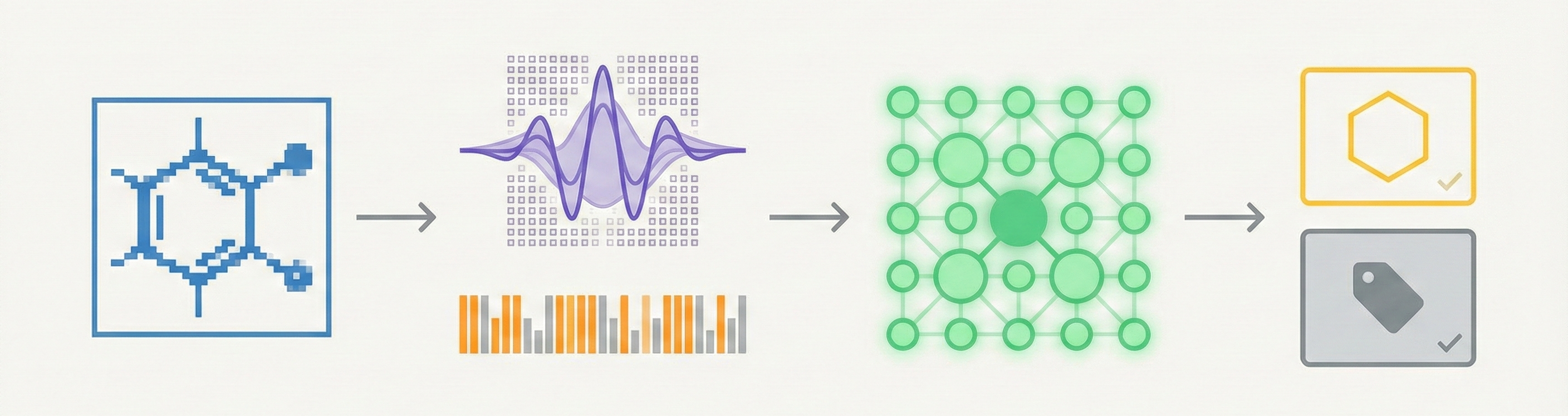
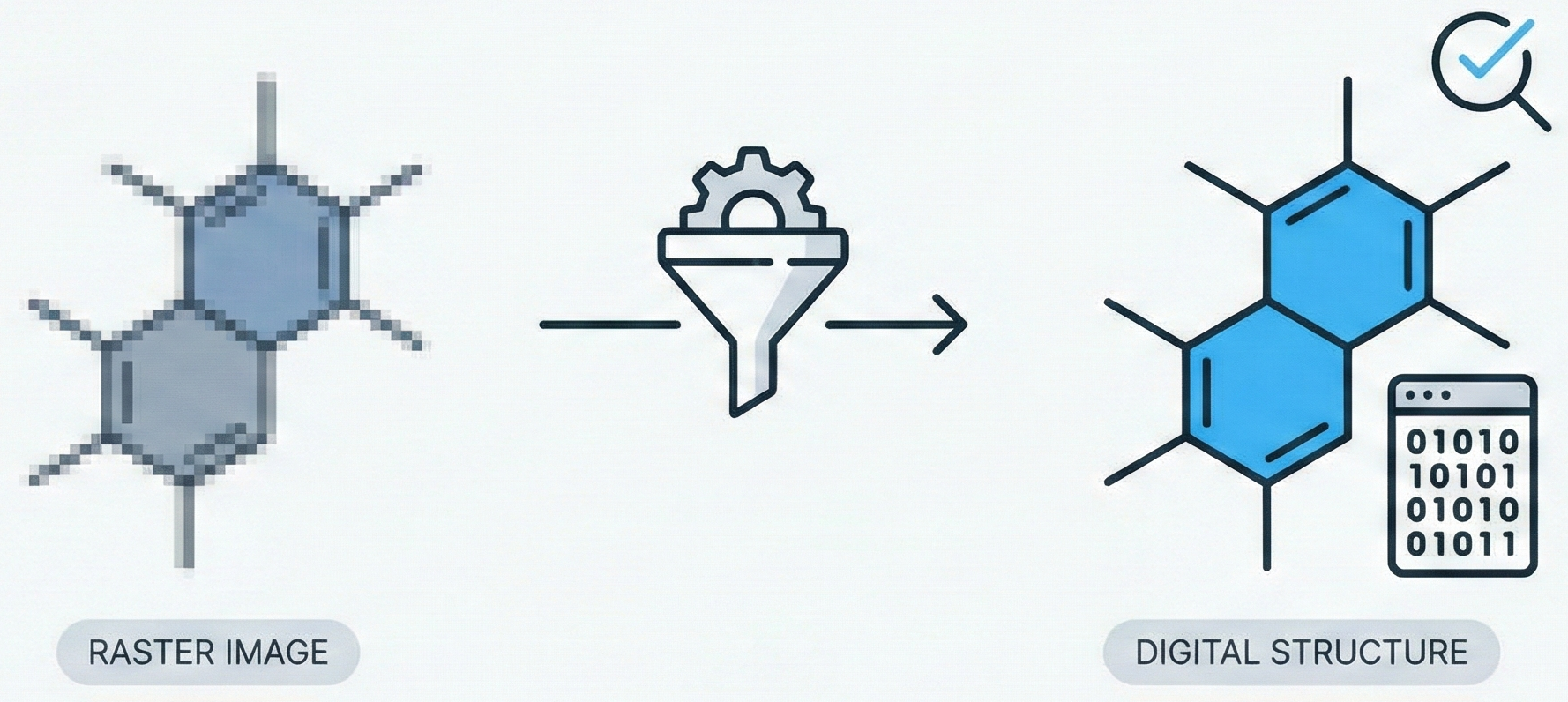
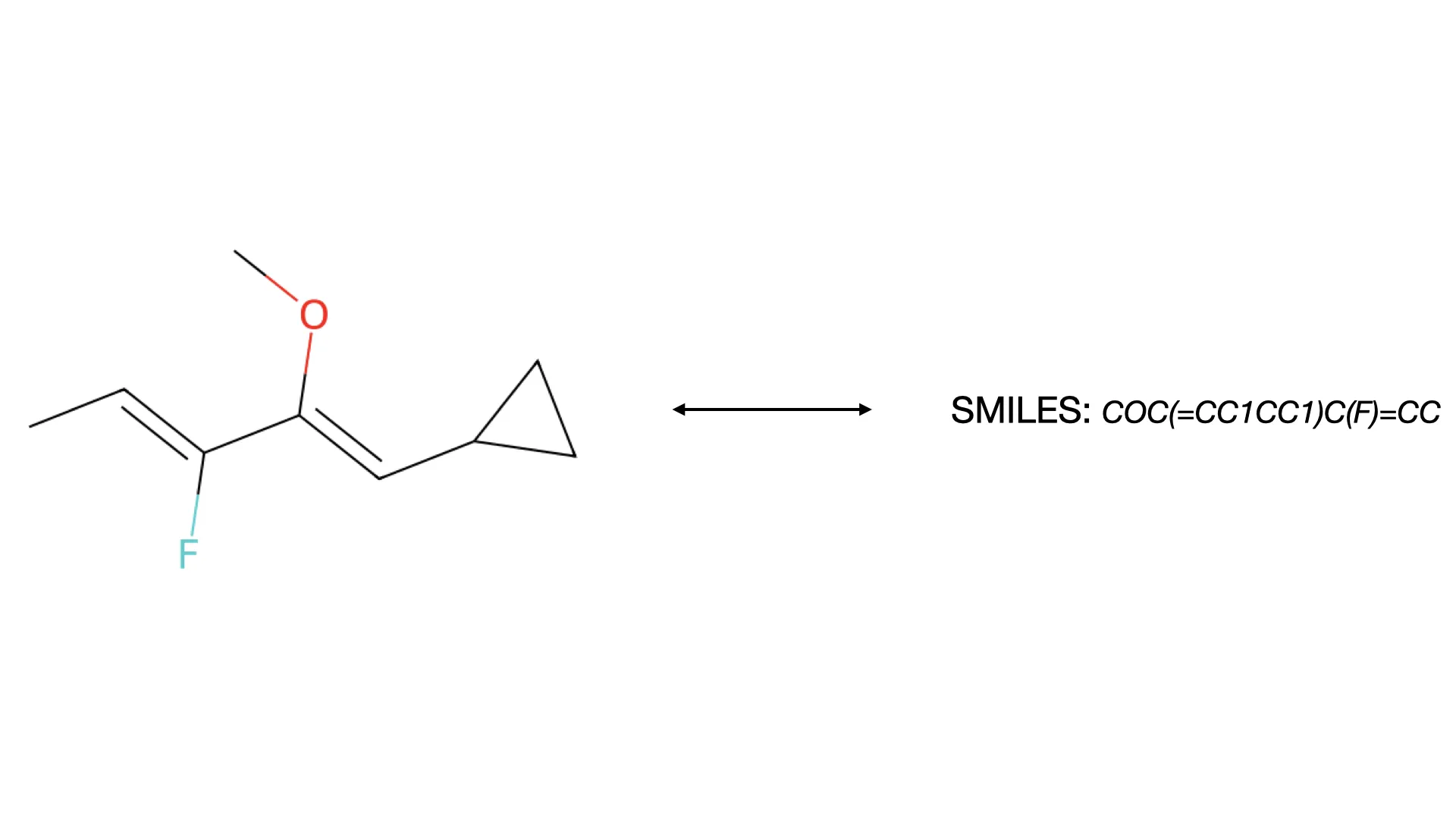
This paper introduces CLiDE Pro, an advanced OCSR system that segments document images and reconstructs chemical connection tables. It features novel handling for crossing bonds and generic structures, validating performance on a publicly released benchmark of 454 scanned images.