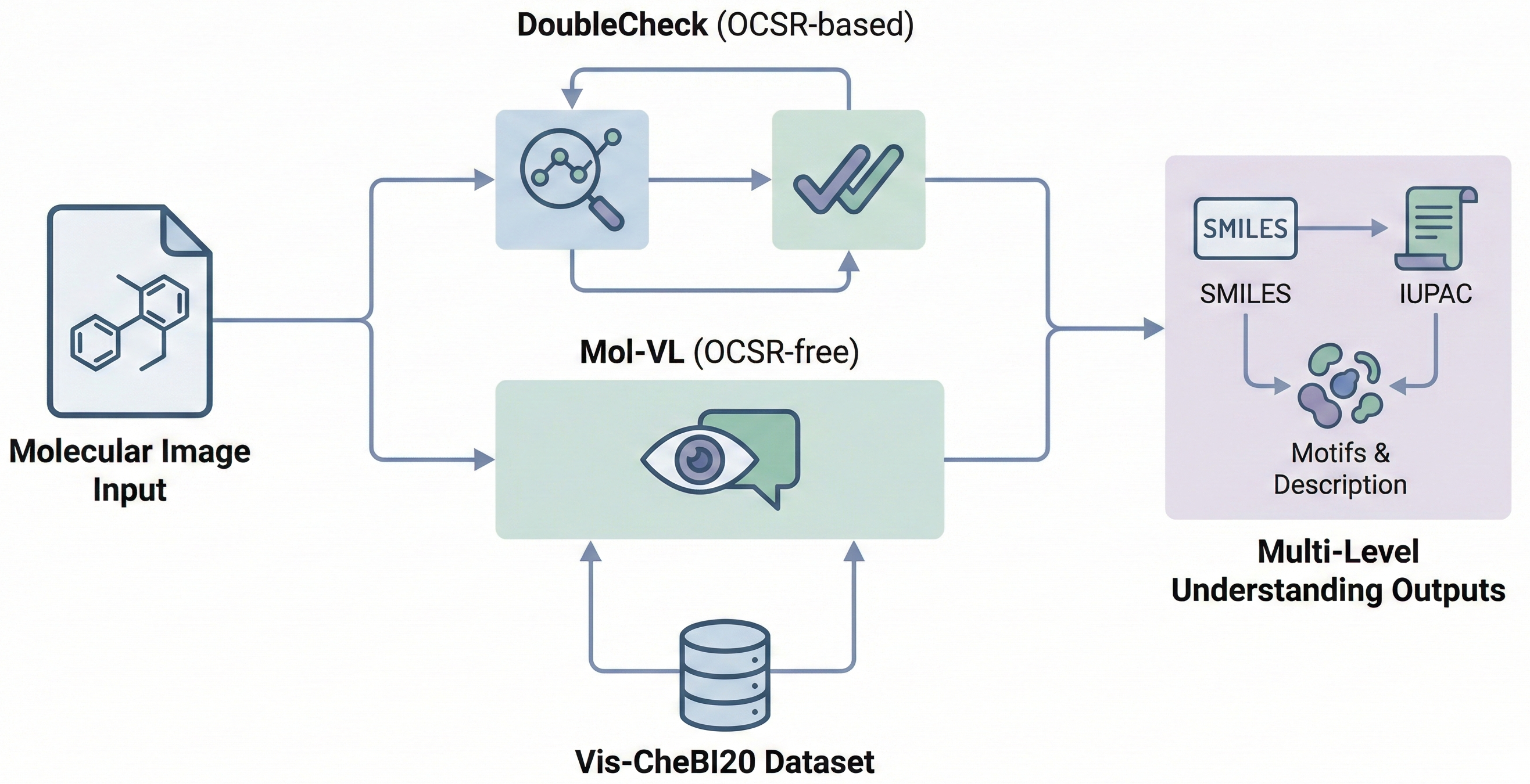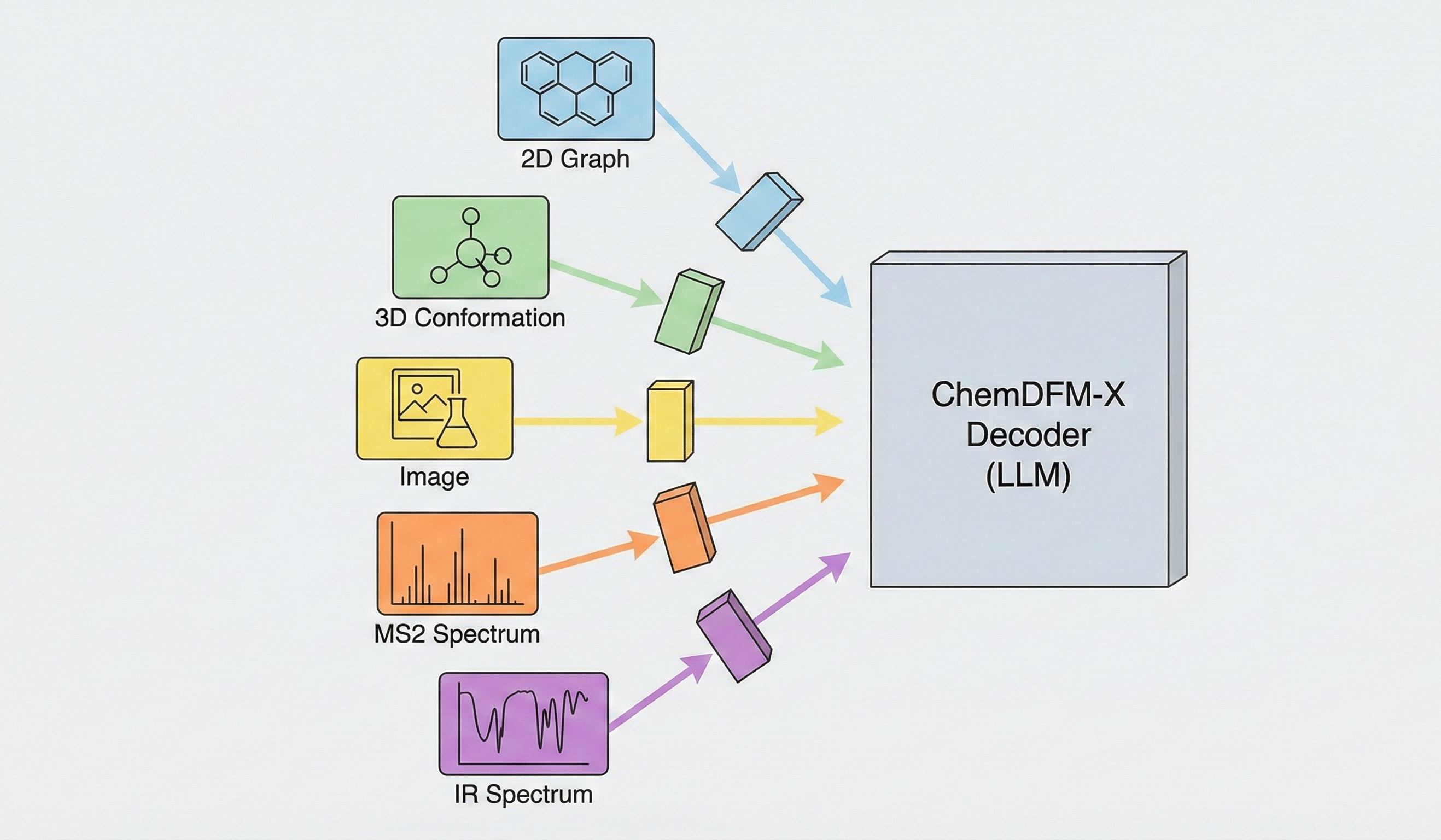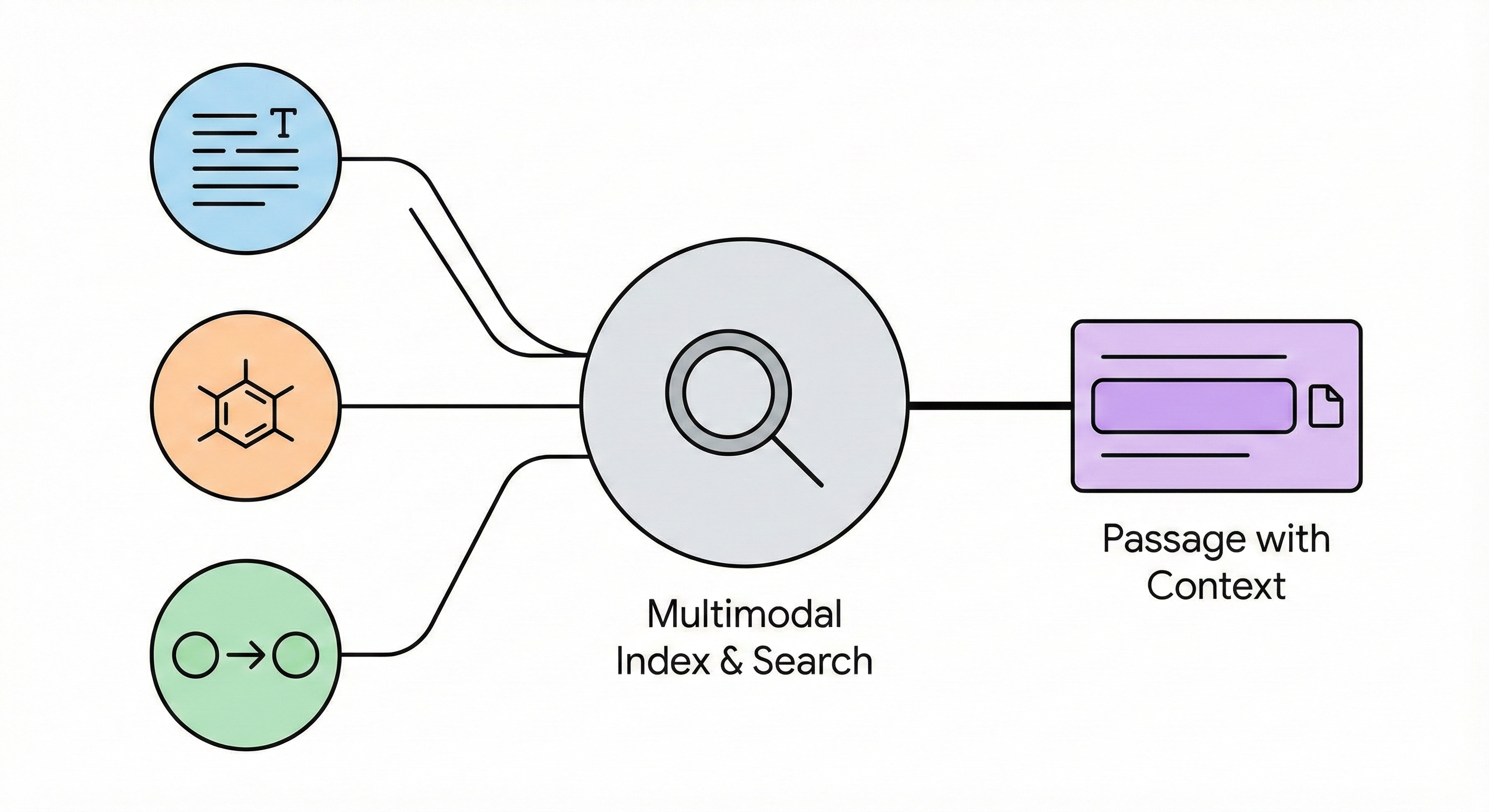
GTR-CoT: Graph Traversal Chain-of-Thought for Molecules
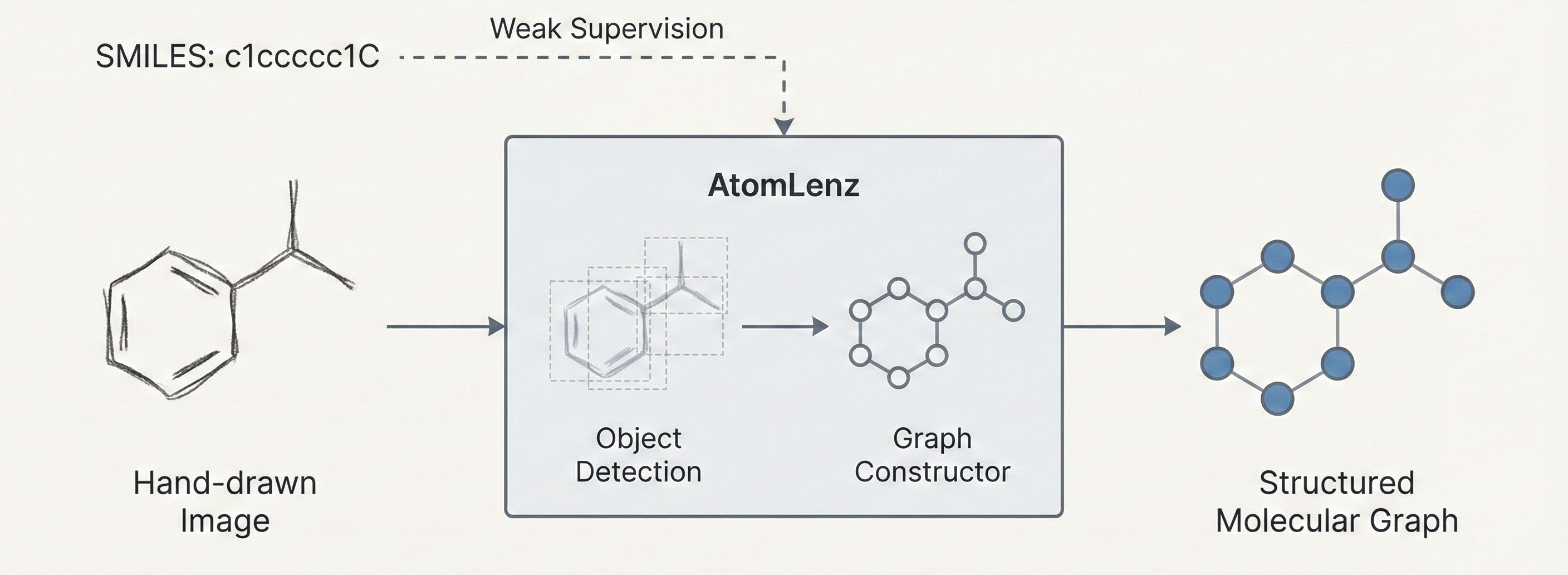
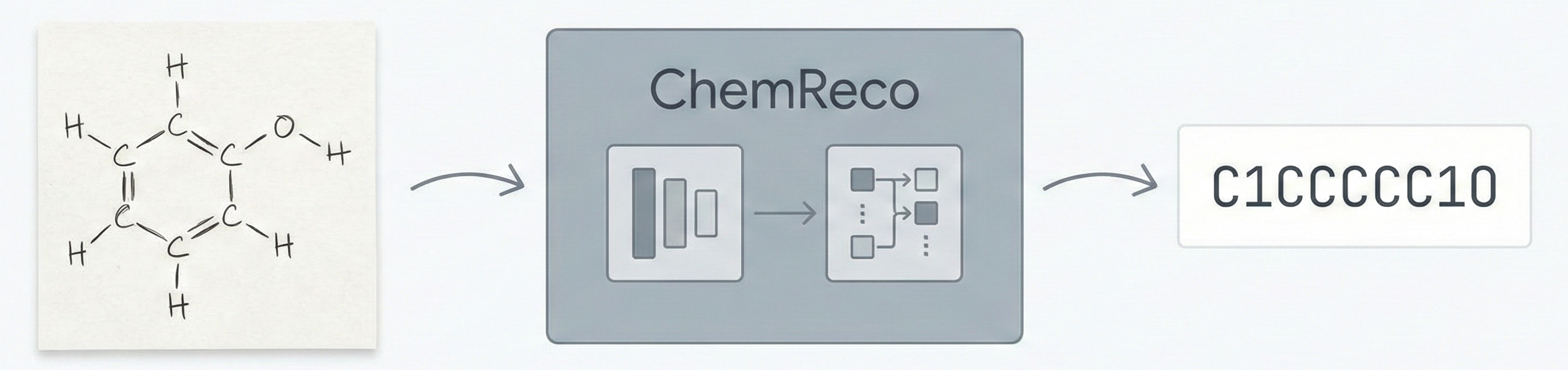
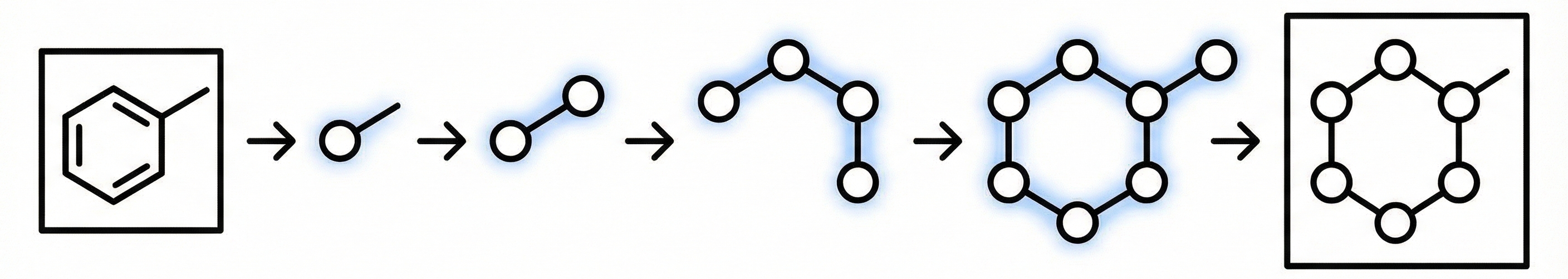
A 2025 Vision-Language Model for OCSR that uses graph traversal chain-of-thought reasoning and a two-stage SFT plus GRPO training scheme to handle both printed molecules (including chemical abbreviations like Ph and Et) and hand-drawn structures, achieving strong performance on the new MolRec-Bench benchmark.