
GutenOCR: A Grounded Vision-Language Front-End for Documents
GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.

GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.
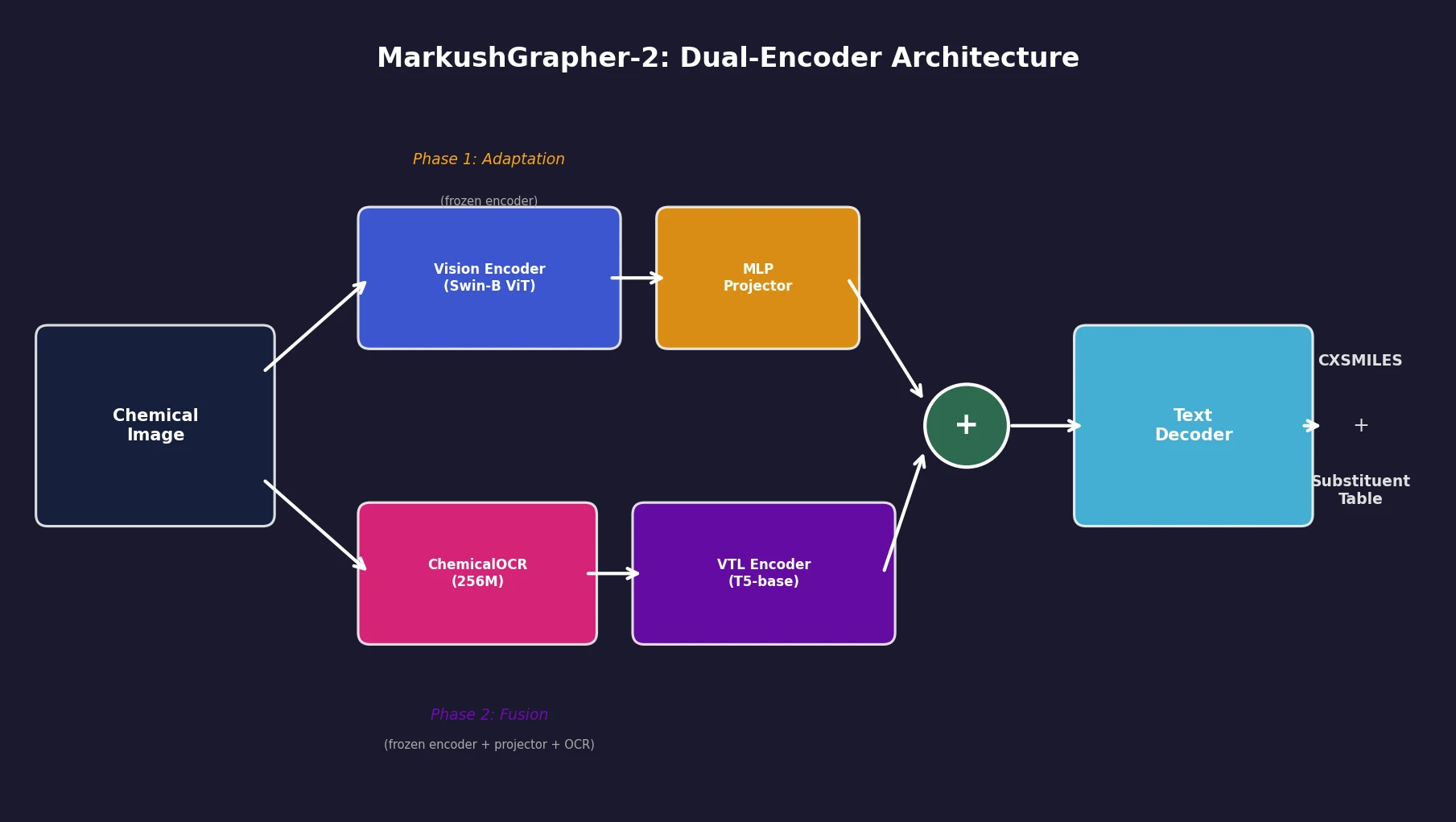
An 831M-parameter encoder-decoder model that jointly encodes image, OCR text, and layout information through a two-stage training strategy, achieving state-of-the-art multimodal Markush structure recognition while remaining competitive on standard molecular structure recognition.
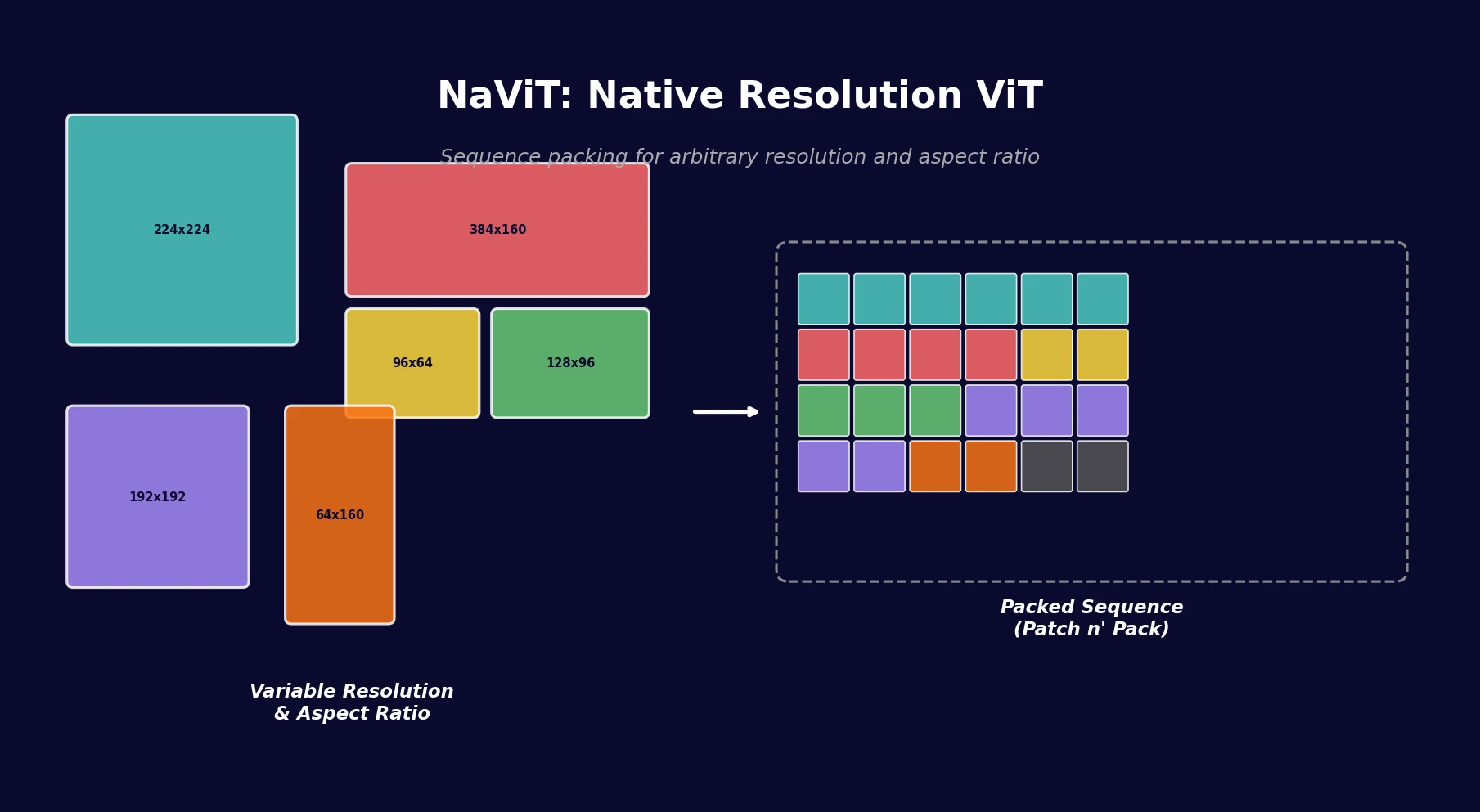
NaViT applies sequence packing (Patch n’ Pack) to Vision Transformers, enabling training on images of arbitrary resolution and aspect ratio while improving training efficiency by up to 4x over standard ViT.
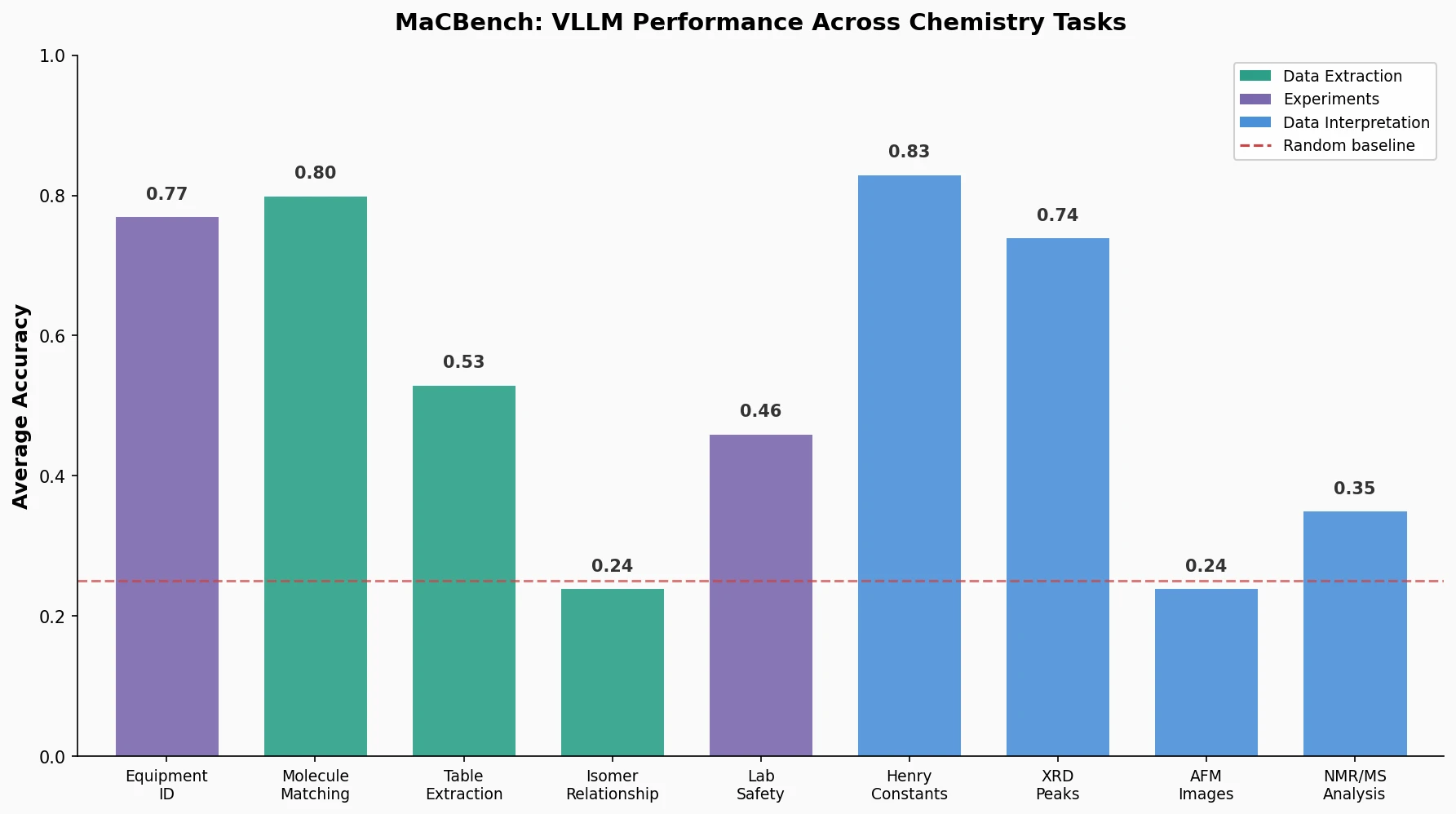
MaCBench evaluates frontier vision language models across 1,153 chemistry and materials science tasks spanning data extraction, experimental execution, and data interpretation, uncovering fundamental limitations in spatial reasoning and cross-modal integration.
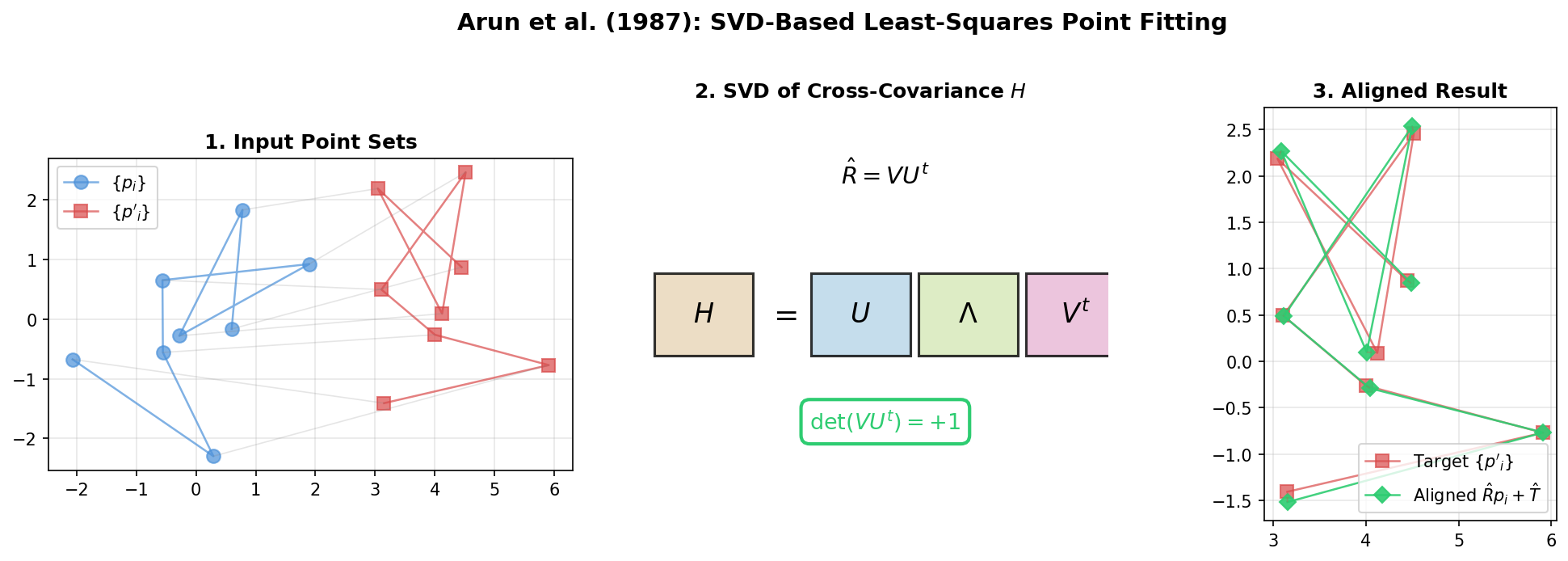
Presents a concise SVD-based algorithm for finding the optimal rotation and translation between two 3D point sets, with analysis of the degenerate reflection case that Umeyama later corrected.
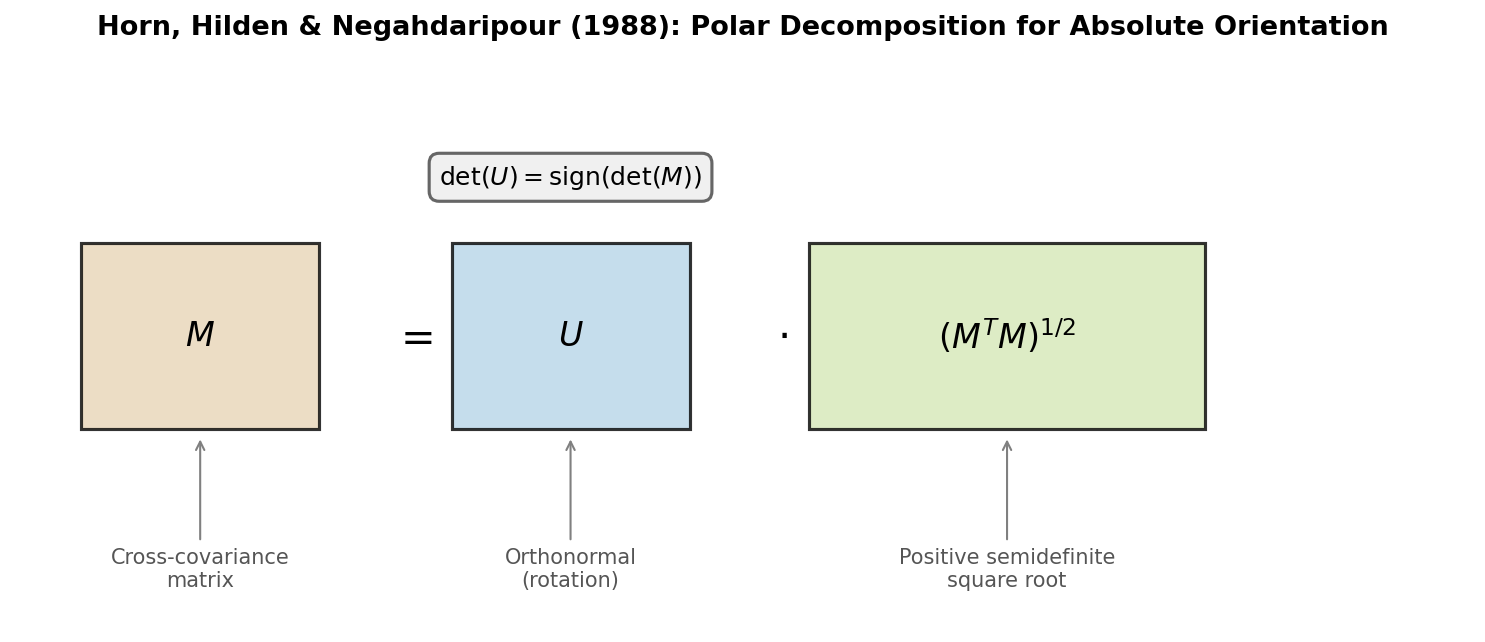
The matrix-based companion to Horn’s 1987 quaternion method, deriving the optimal rotation as the orthonormal factor in the polar decomposition of the cross-covariance matrix via eigendecomposition of a 3x3 symmetric matrix.
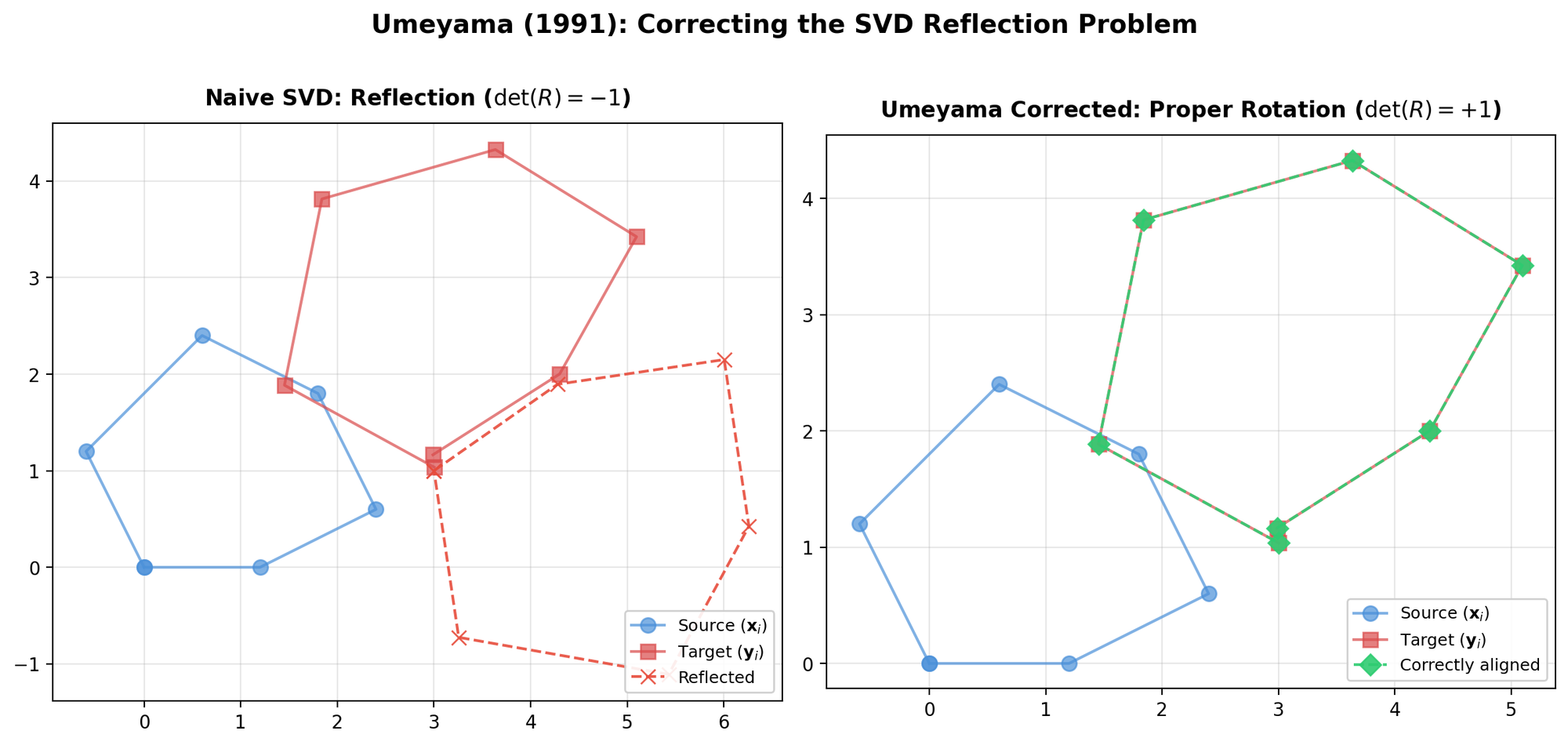
Corrects a flaw in prior SVD-based alignment methods (Arun et al., Horn et al.) that could produce reflections instead of rotations under noisy data, and provides a complete closed-form solution for similarity transformations in arbitrary dimensions.
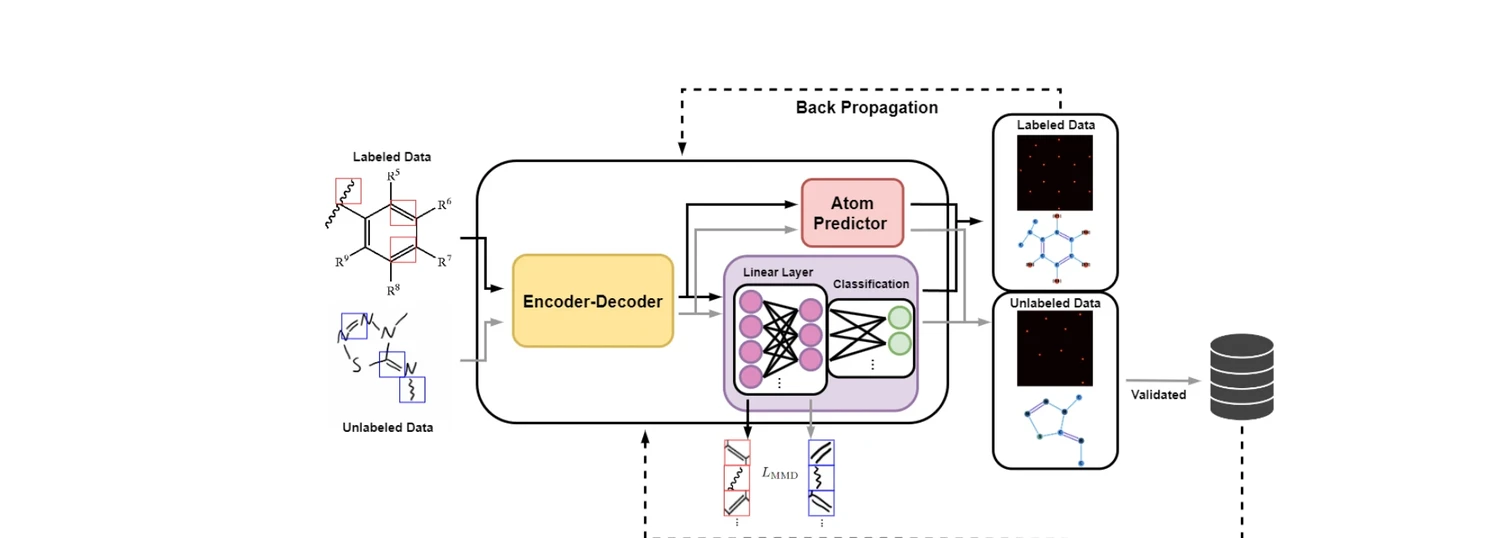
AdaptMol combines an end-to-end graph reconstruction model with unsupervised domain adaptation via class-conditional MMD on bond features and SMILES-validated self-training. Achieves 82.6% accuracy on hand-drawn molecules (10.7 points above prior best) while maintaining state-of-the-art results on four literature benchmarks, using only 4,080 real hand-drawn images for adaptation.
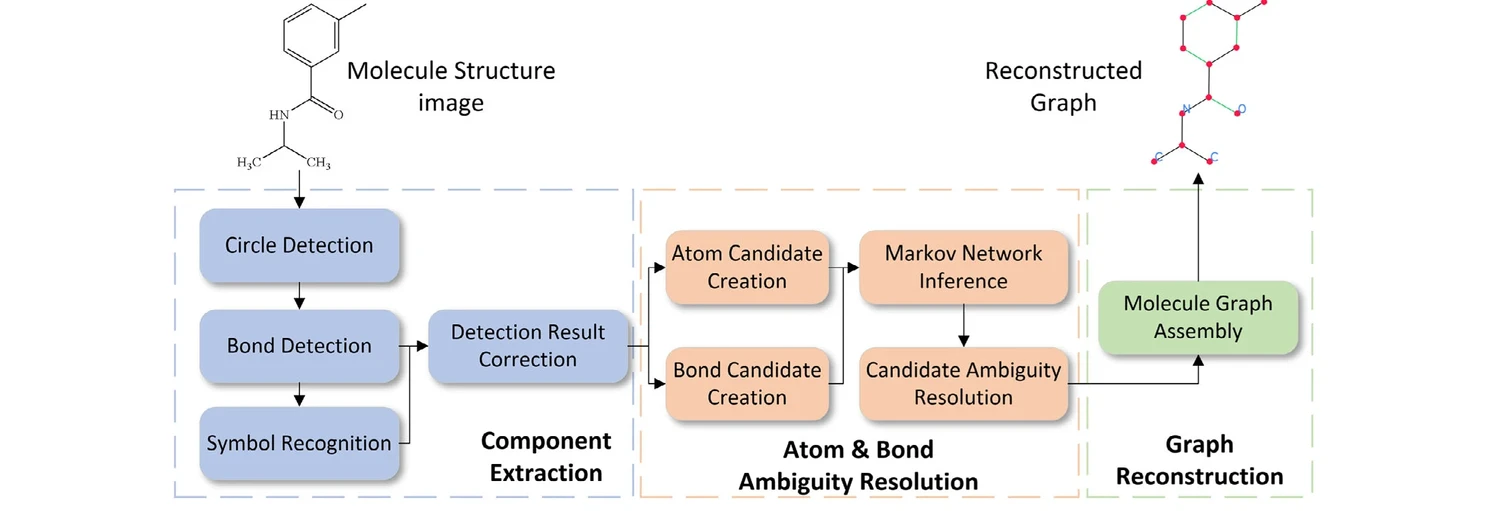
GraphReco presents a rule-based OCSR system with two key innovations: a Fragment Merging line detection algorithm for precise bond identification and a Markov network for probabilistic resolution of atom/bond ambiguity during graph assembly. Achieves 94.2% accuracy on USPTO-10K, outperforming both traditional rule-based and some ML-based methods.
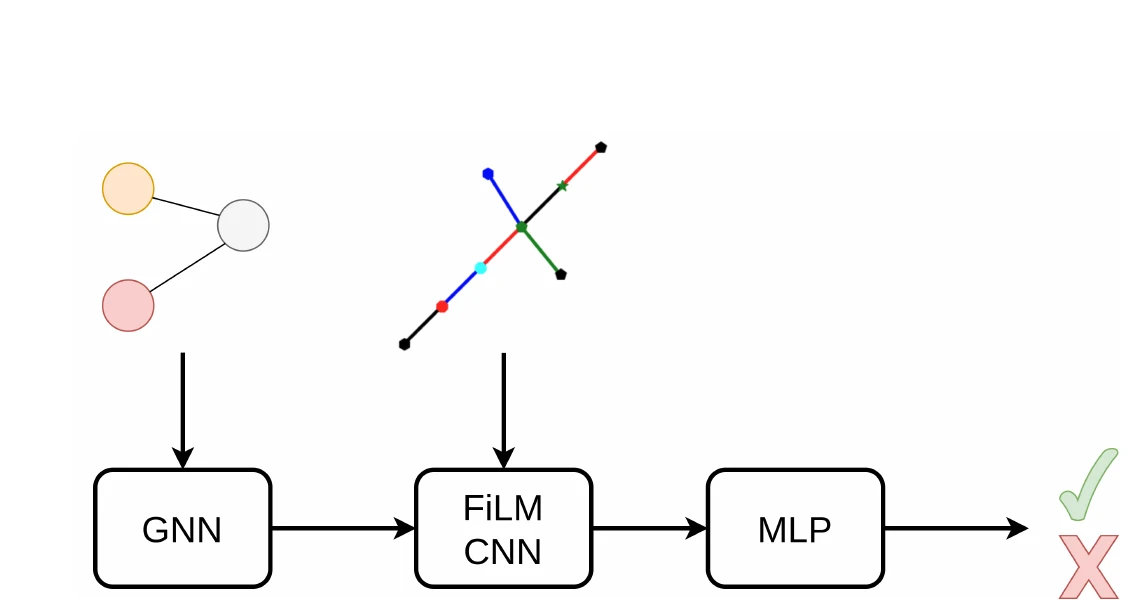
GraSP introduces a general framework for recognizing graphs in images by framing it as sequential subgraph prediction with a binary classifier. A GNN conditions a CNN via FiLM layers to predict whether a candidate graph is a subgraph of the target. Applied to OCSR on QM9, GraSP achieves 67.5% accuracy with no domain-specific modifications.
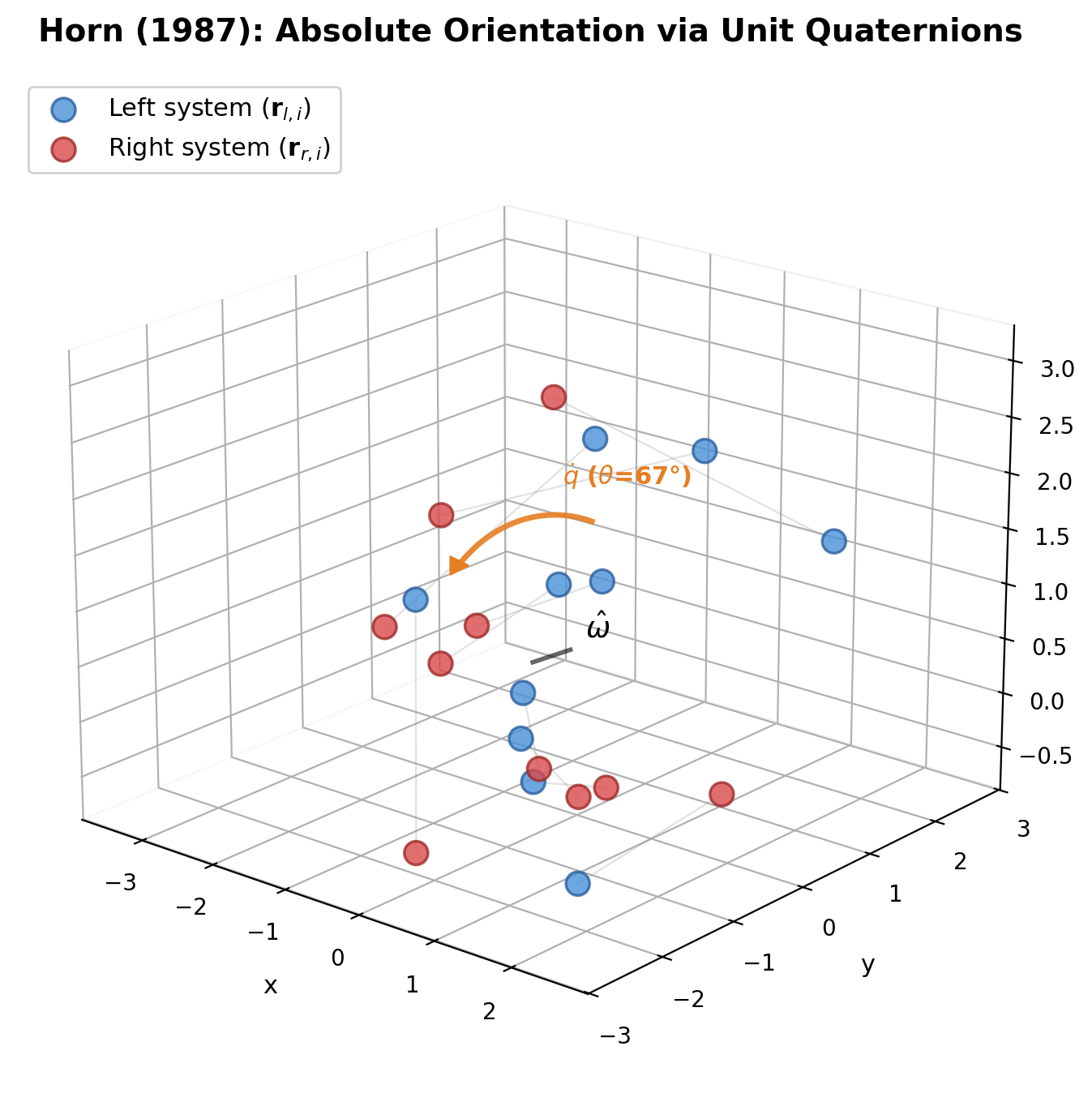
Derives the optimal rotation between two 3D point sets as the eigenvector of a 4x4 symmetric matrix built from cross-covariance sums, using unit quaternions to enforce the orthogonality constraint.
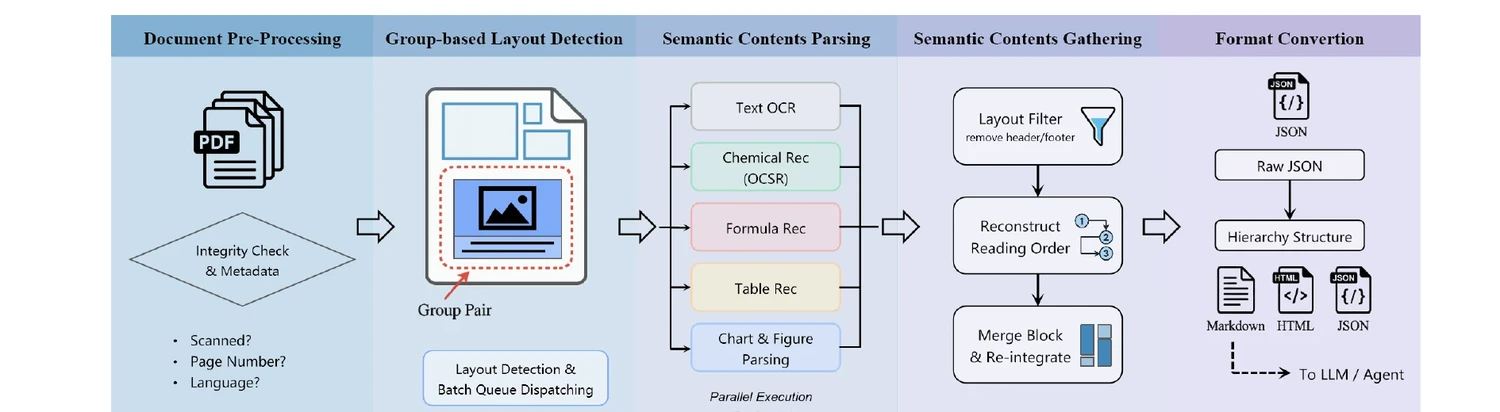
Technical report on Uni-Parser, an industrial-grade document parsing engine that uses a modular multi-expert architecture to parse scientific PDFs into structured representations. Integrates MolParser 1.5 for OCSR, achieving 88.6% accuracy on chemical structures while processing up to 20 pages per second.