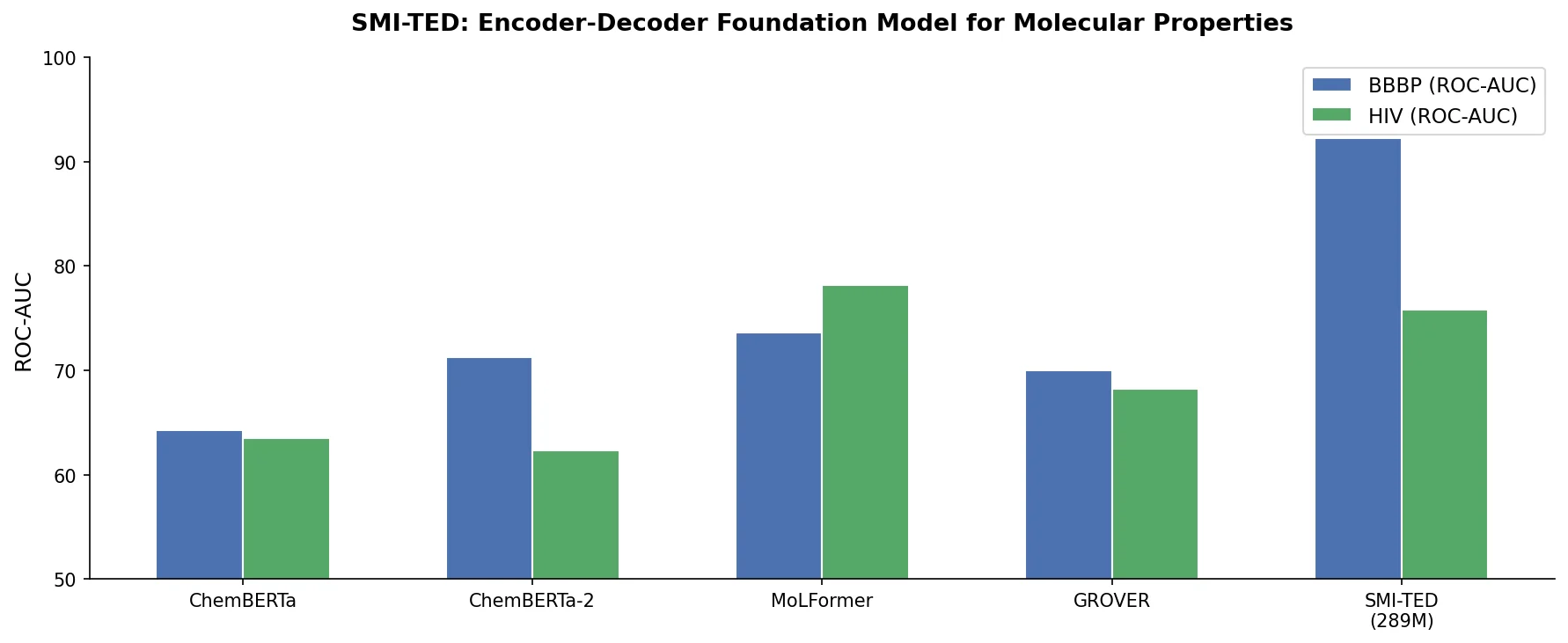
SMI-TED: Encoder-Decoder Foundation Models for Chemistry
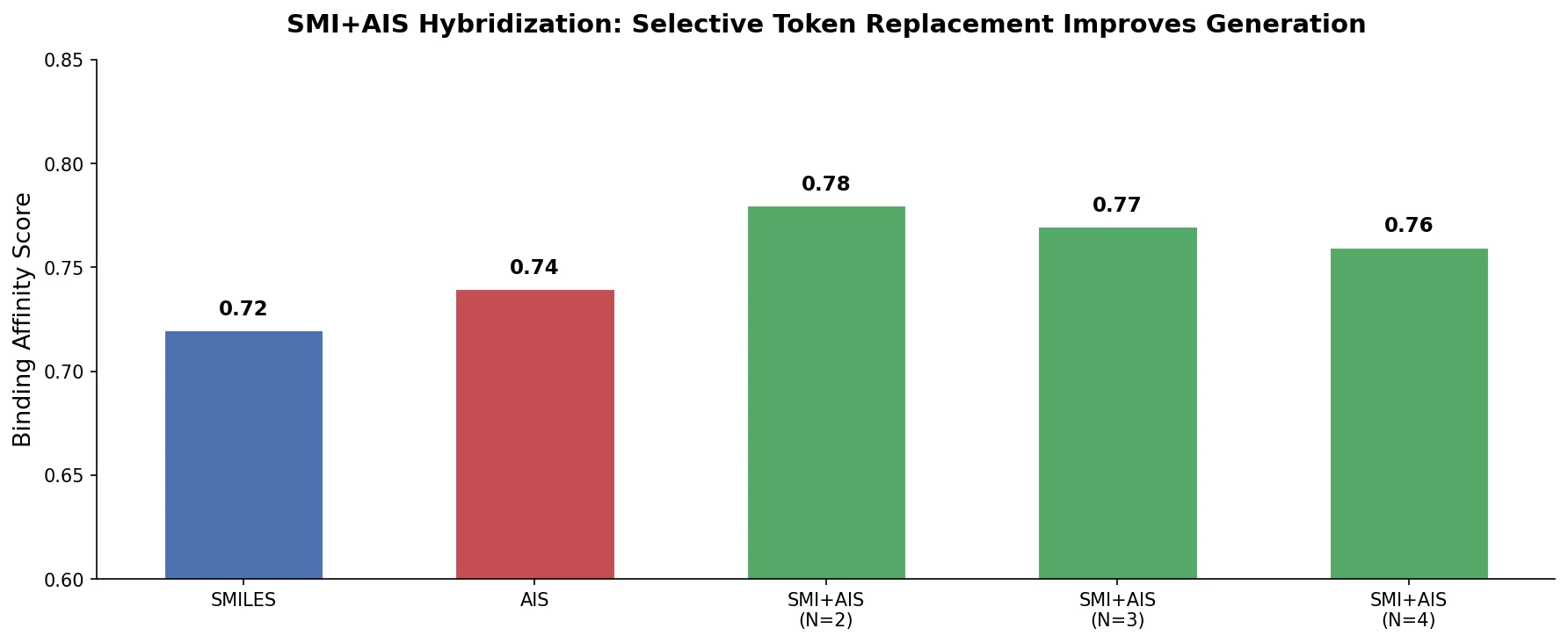
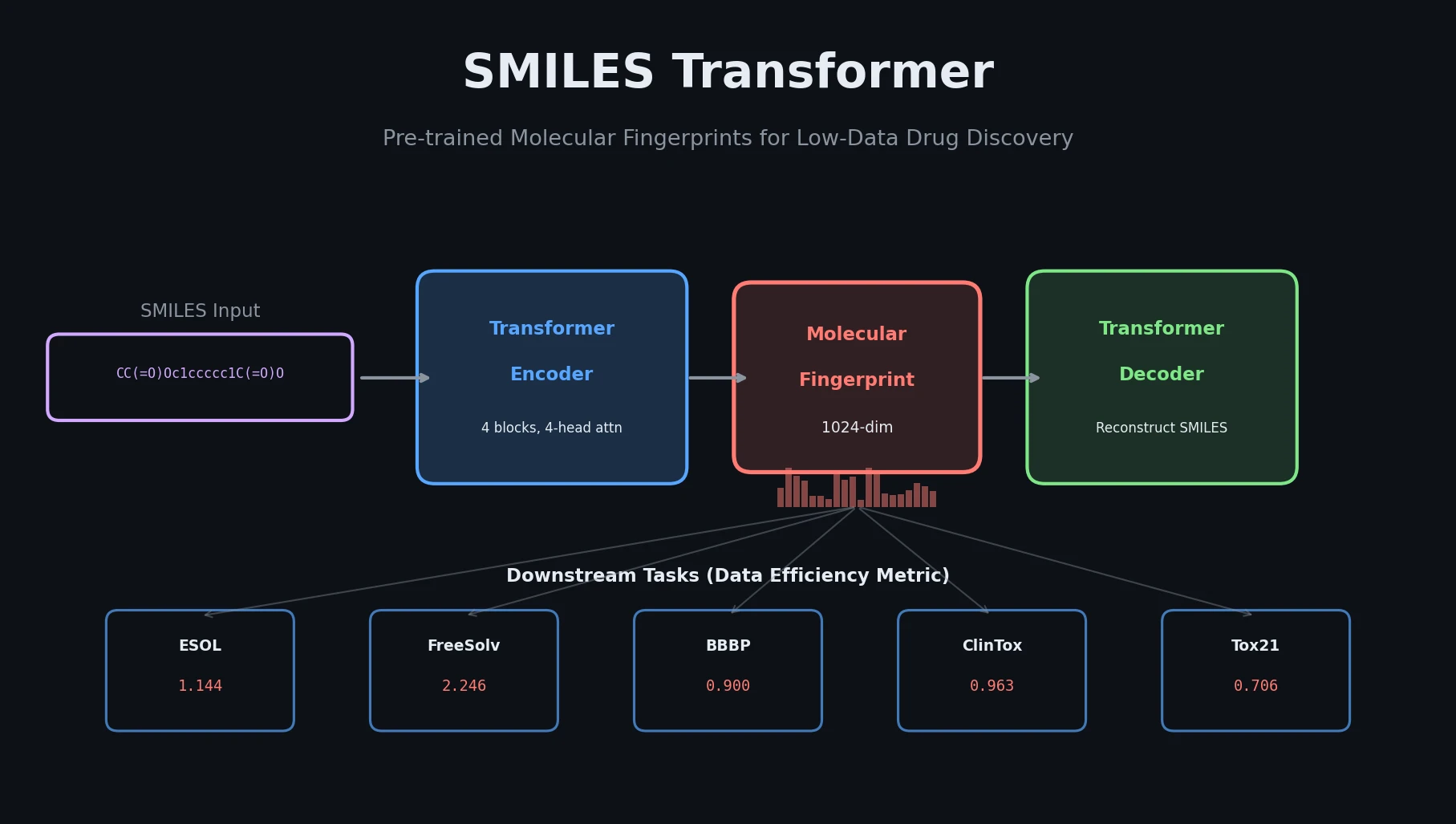
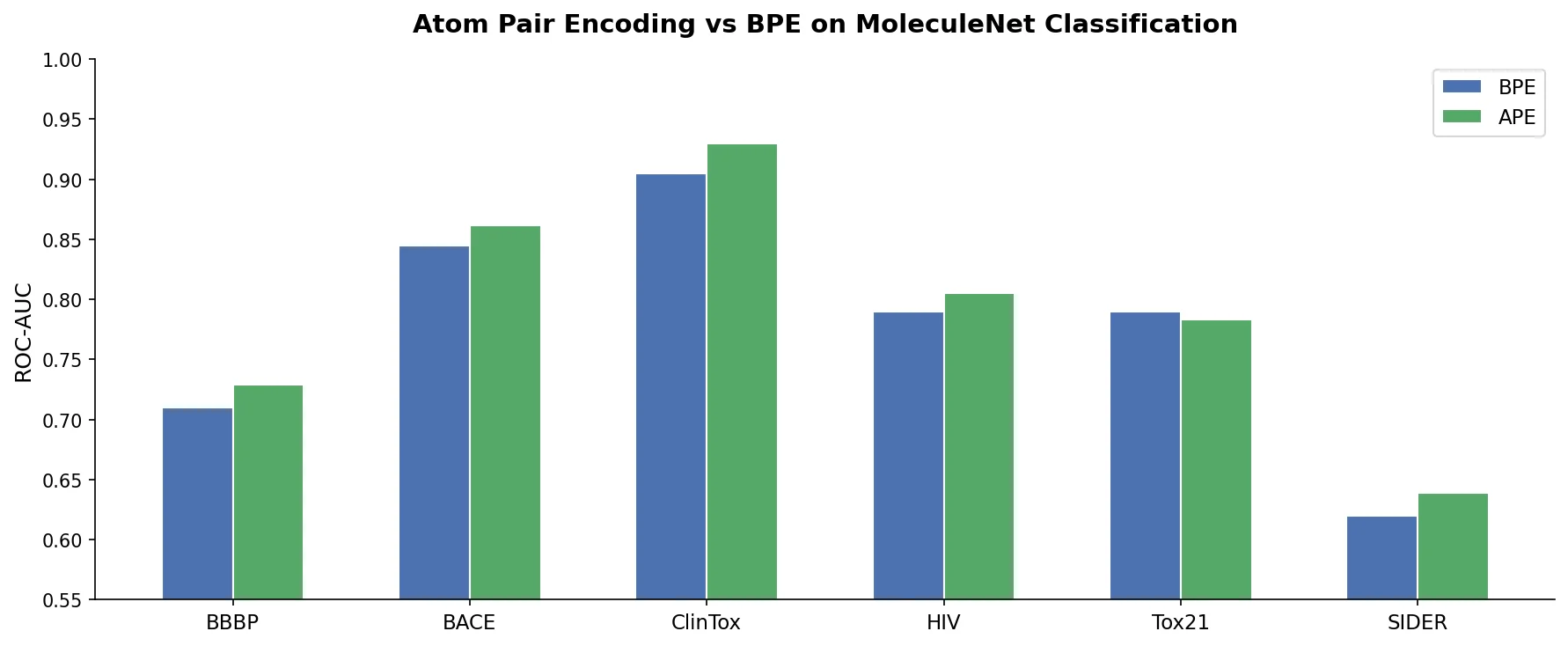
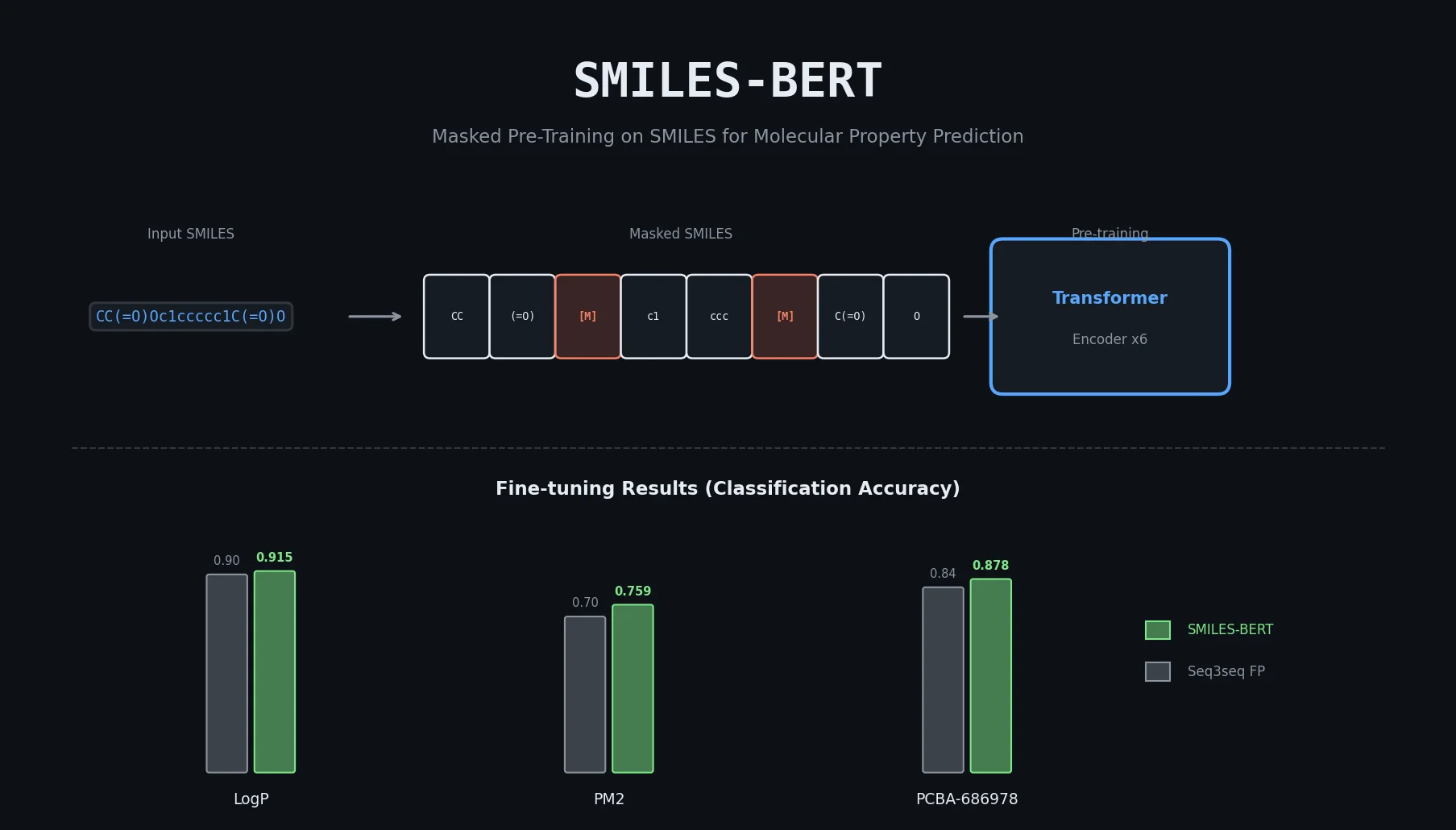
SMI-TED introduces encoder-decoder chemical foundation models (289M parameters) pre-trained on 91 million PubChem molecules, achieving strong results across property prediction, reaction yield, and molecule generation benchmarks.