Foundation Models in Chemistry: A 2025 Perspective
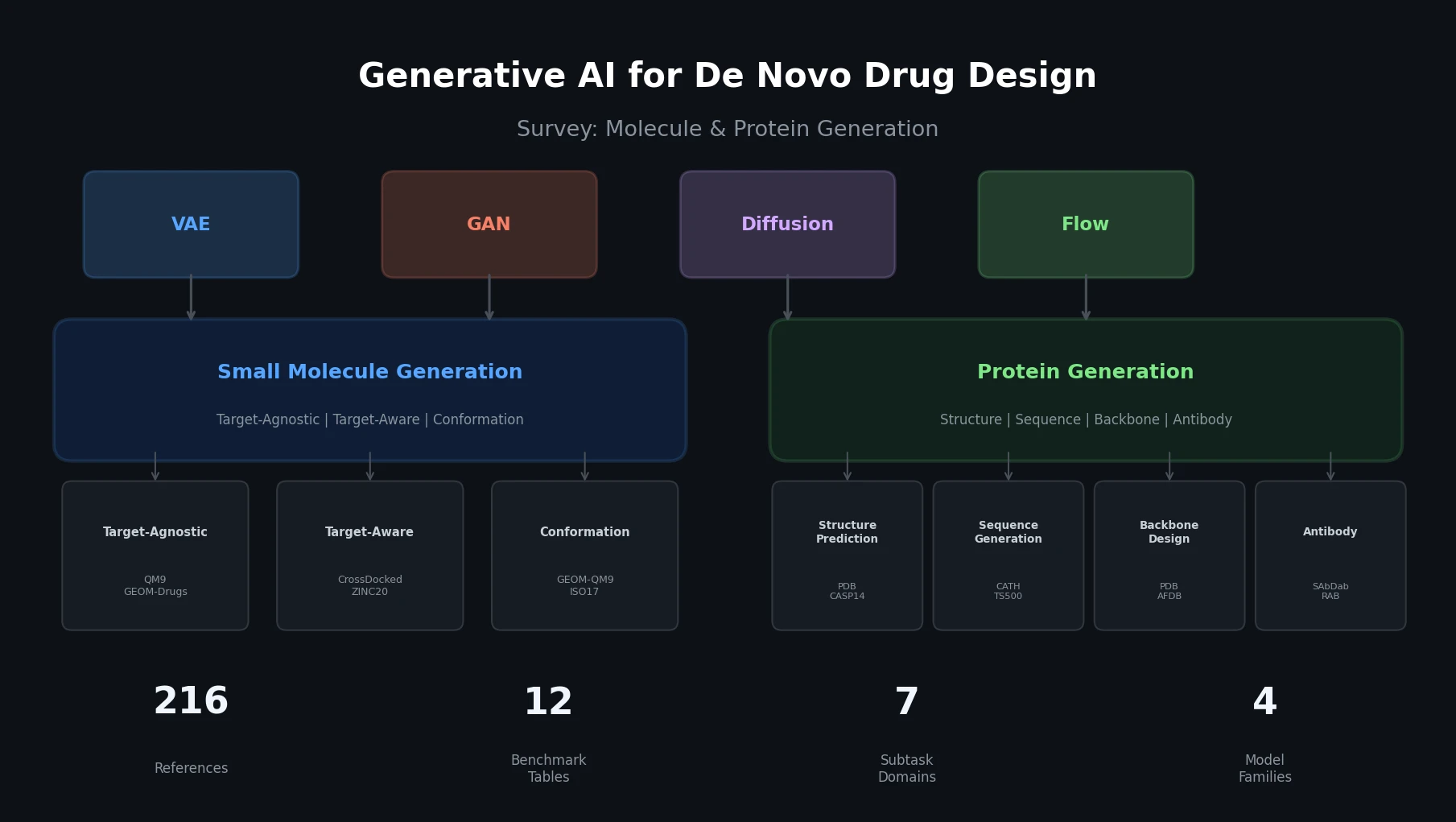
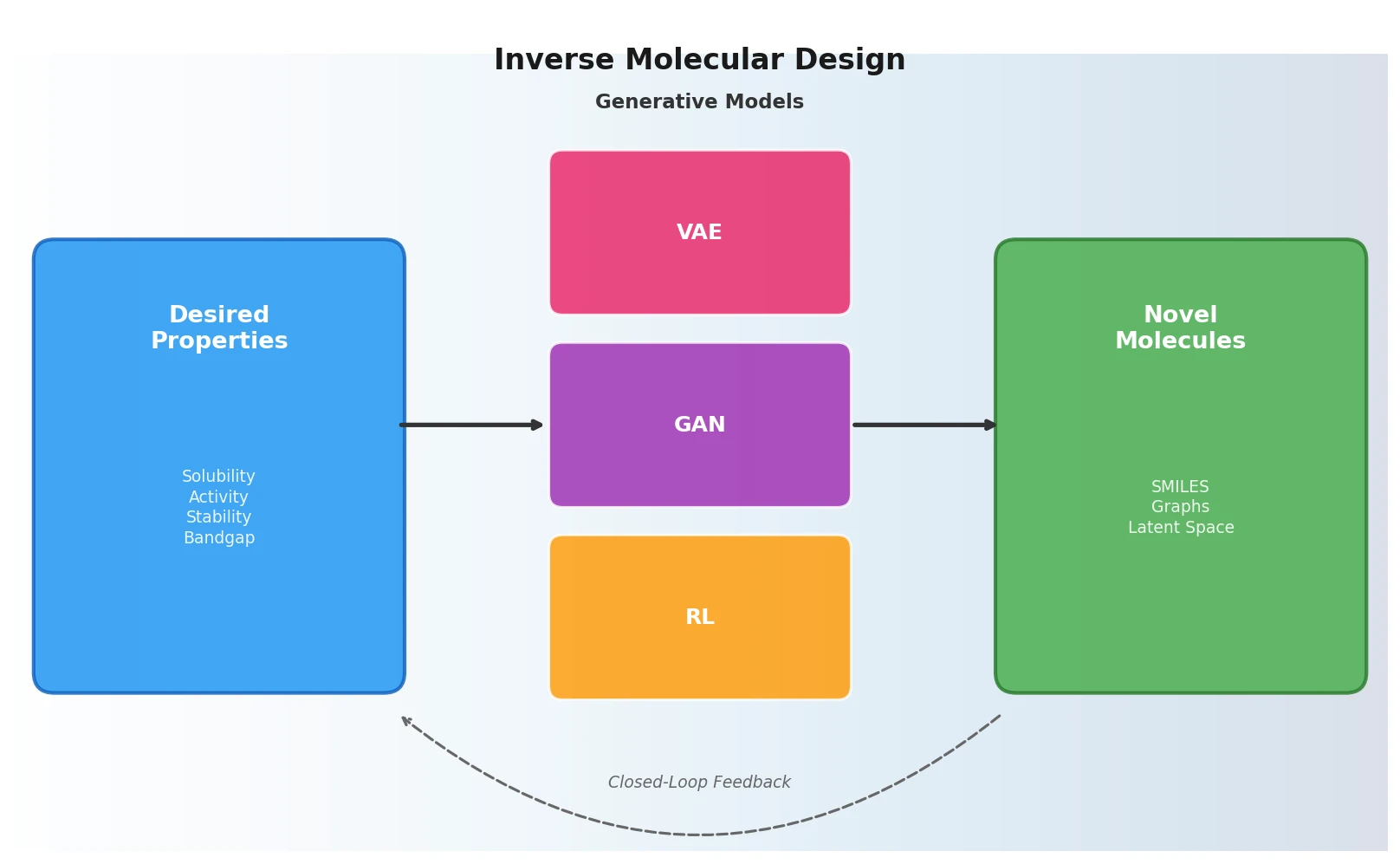
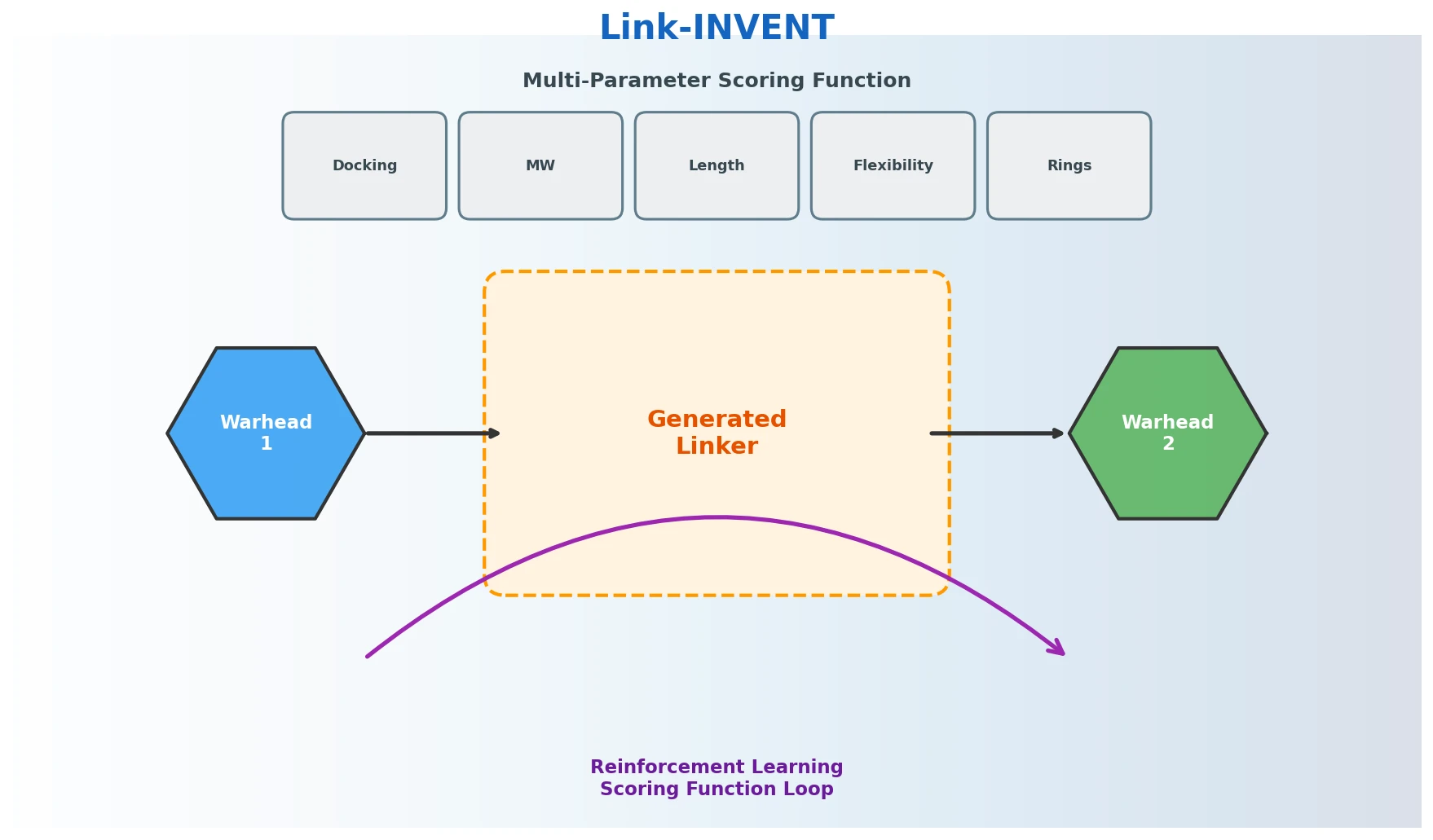
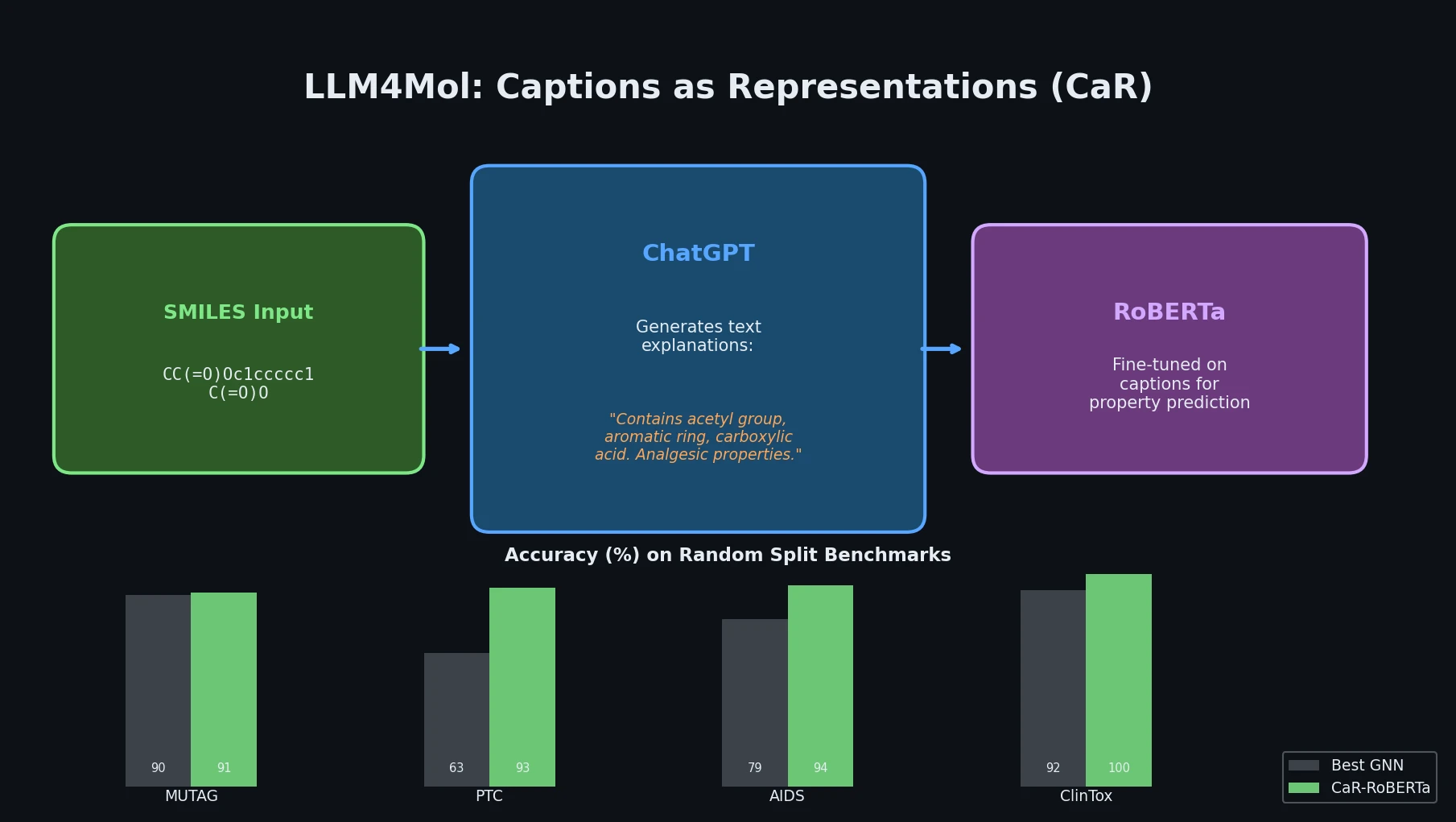
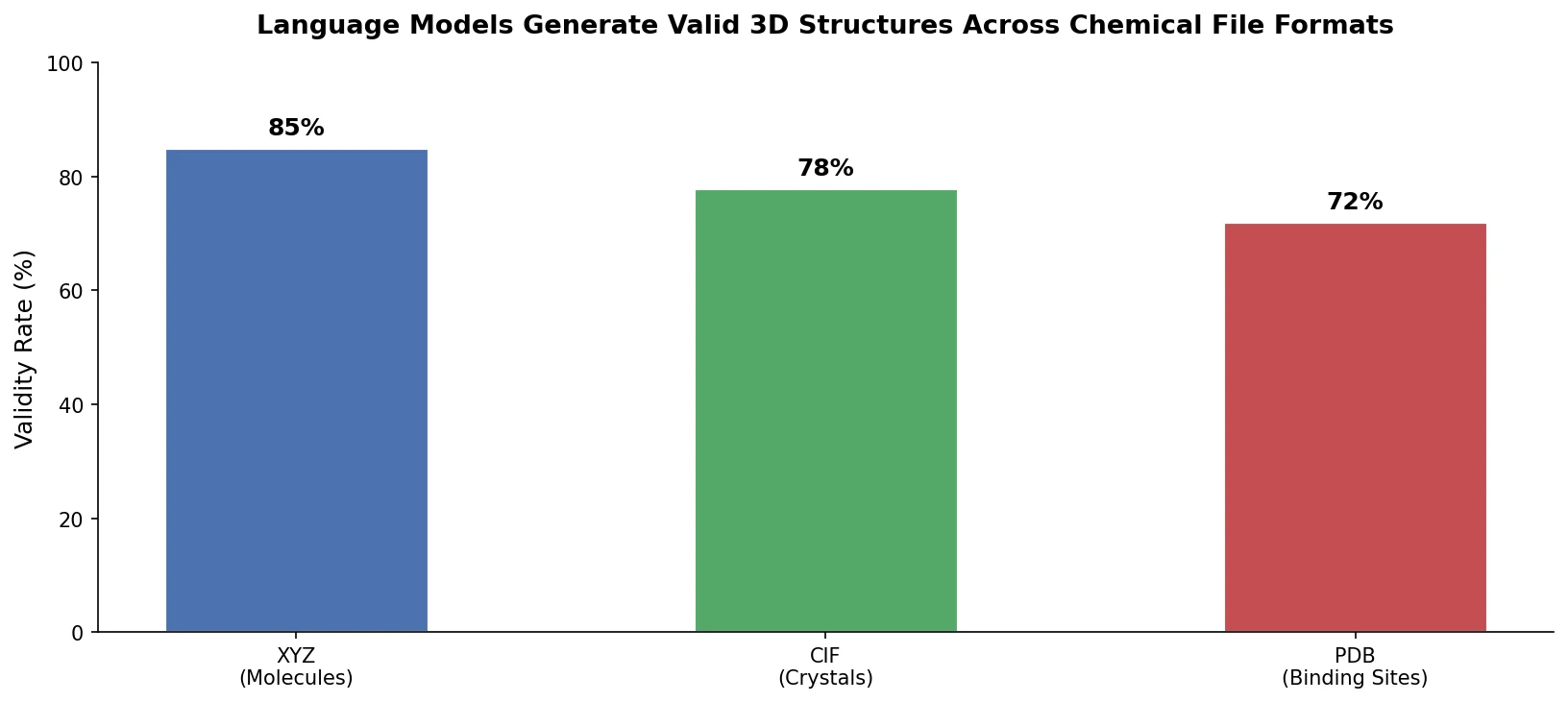
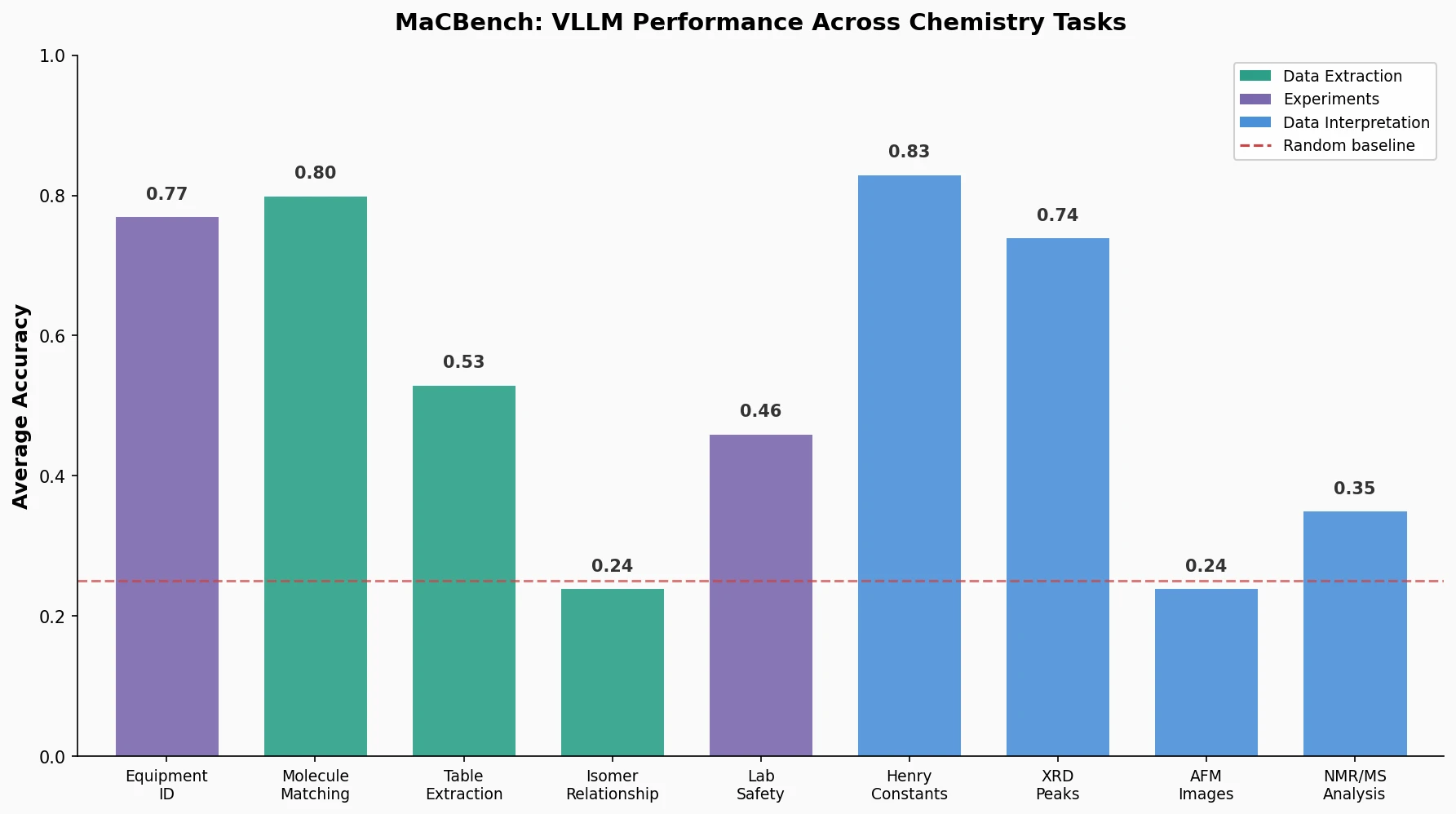
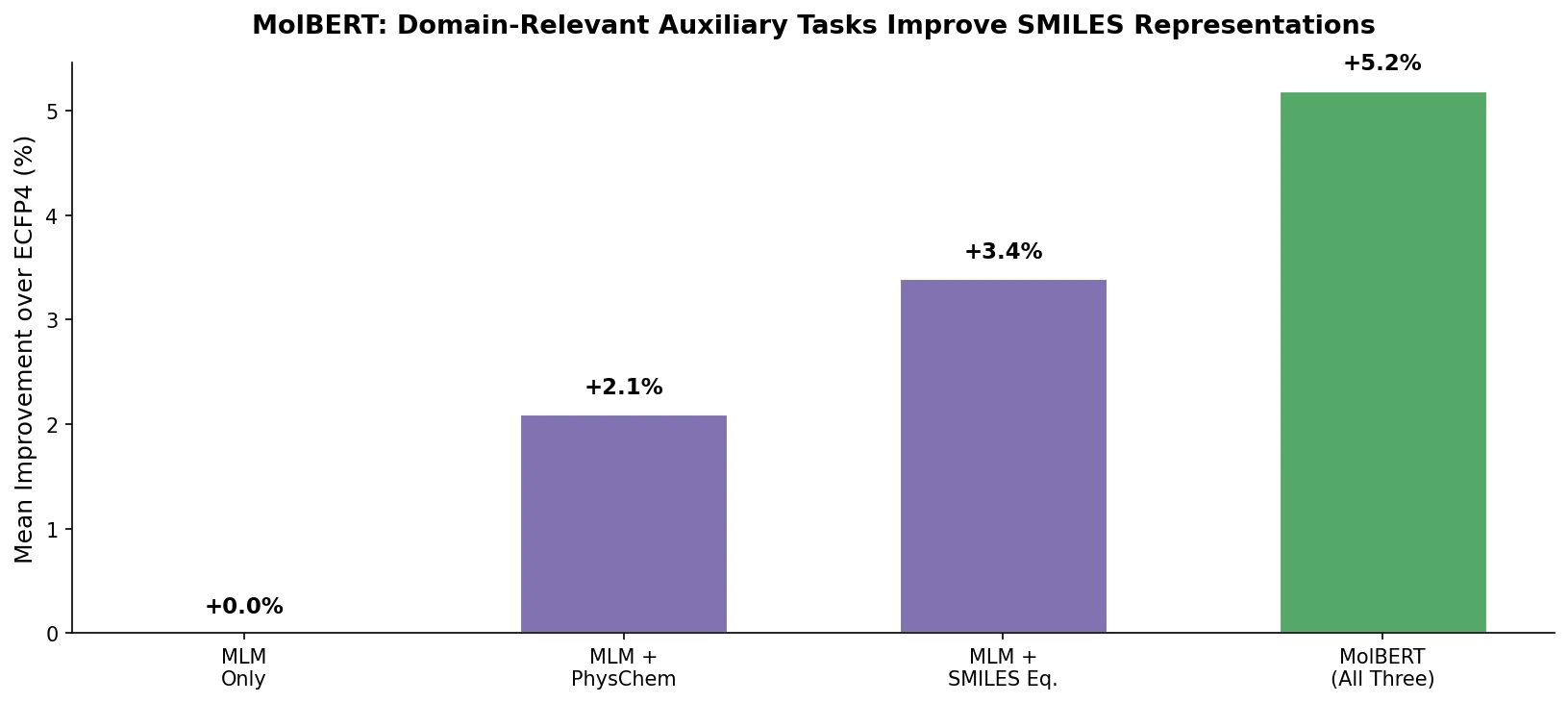
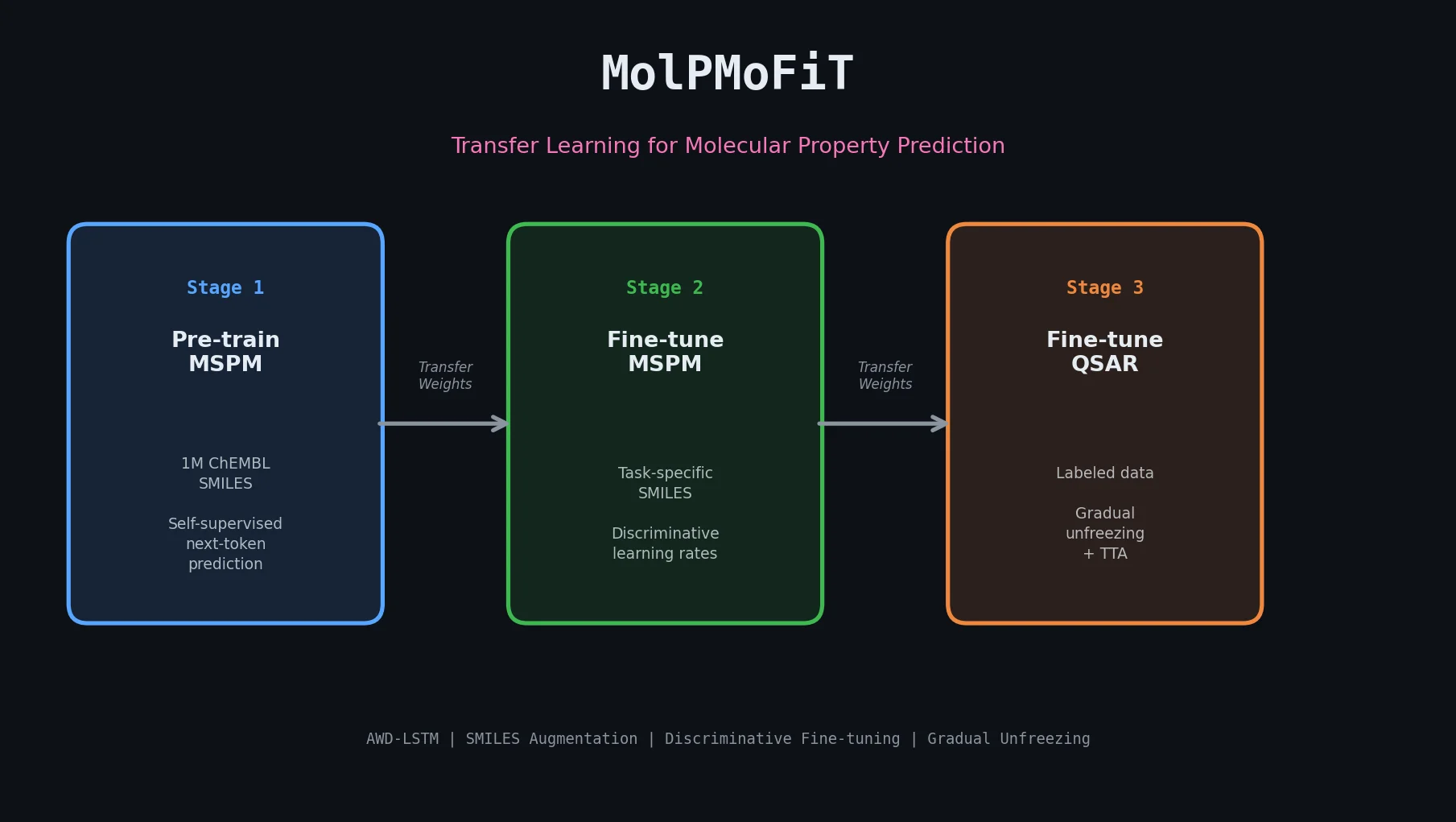
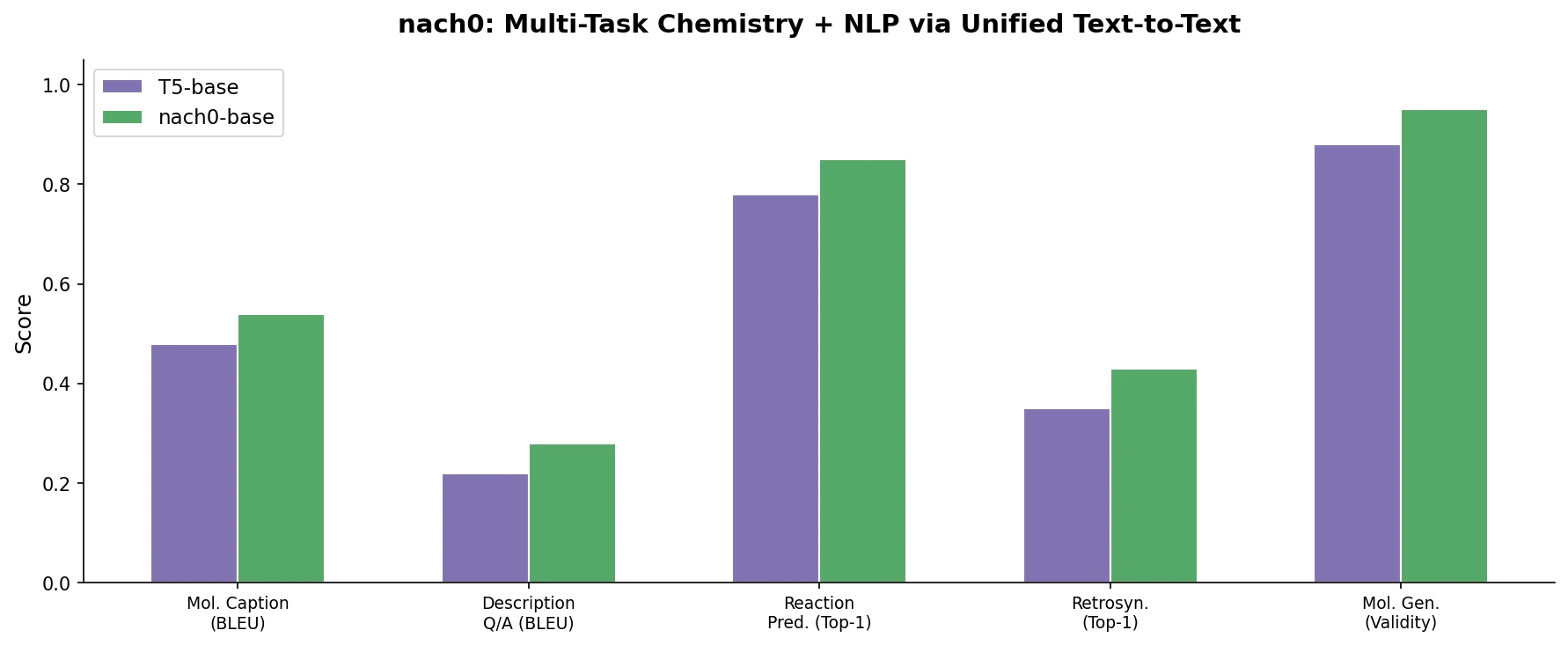
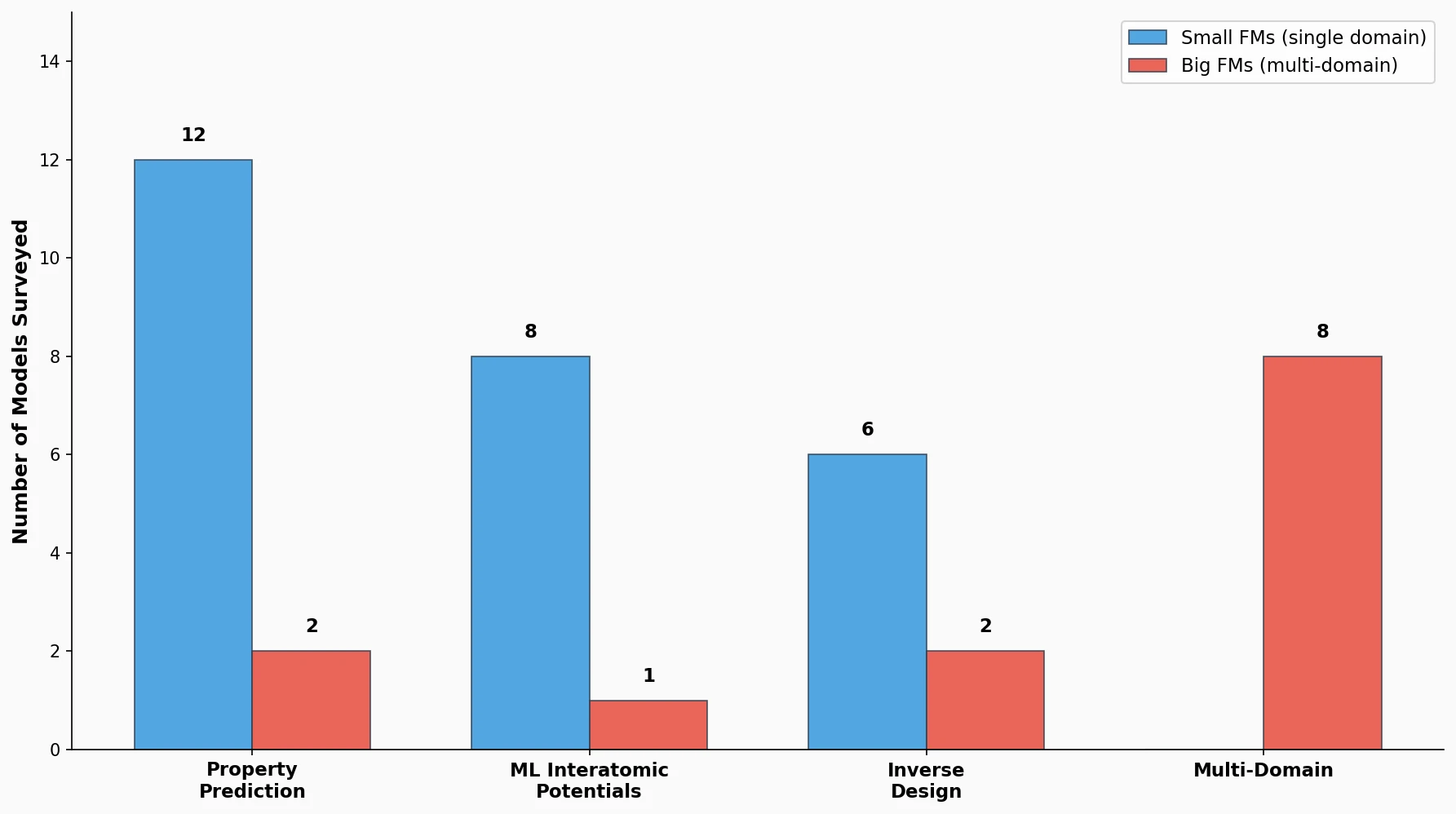
This perspective from Choi et al. reviews foundation models in chemistry, categorizing them as ‘small’ (domain-specific, e.g., property prediction, MLIPs, inverse design) and ‘big’ (multi-domain, e.g., multimodal and LLM-based). It surveys pretraining strategies, key architectures (GNNs and language models), and outlines future directions for scaling, efficiency, and interpretability.