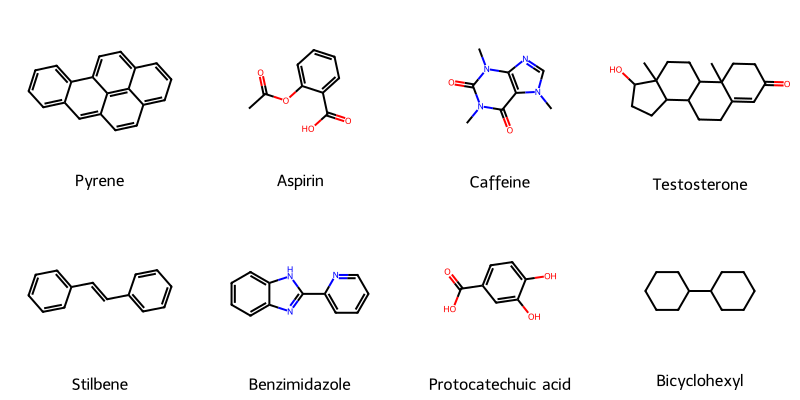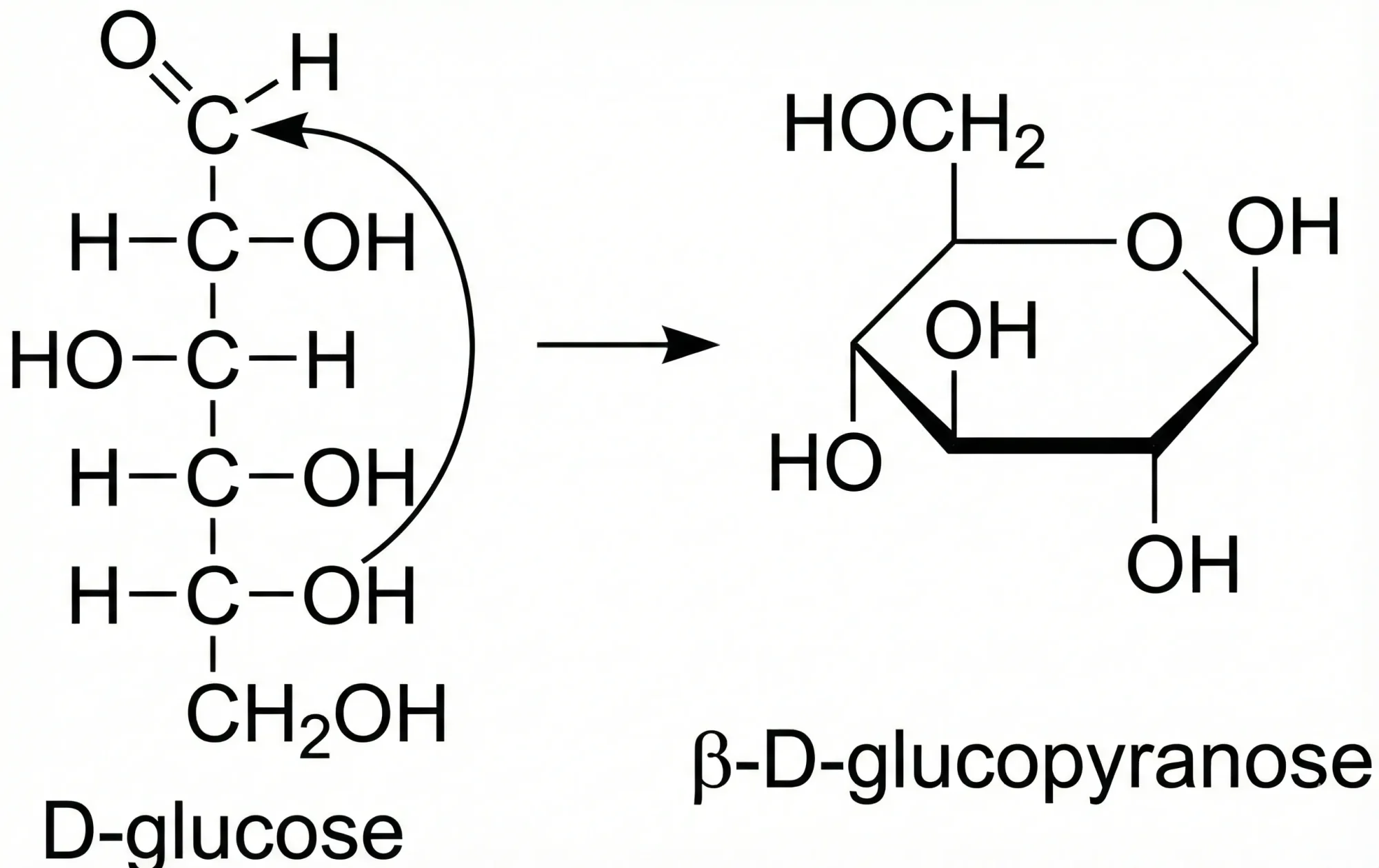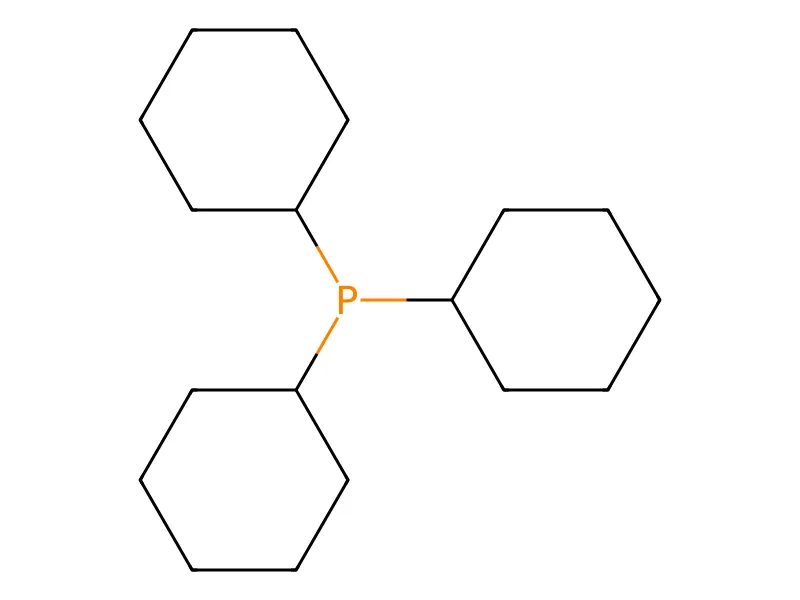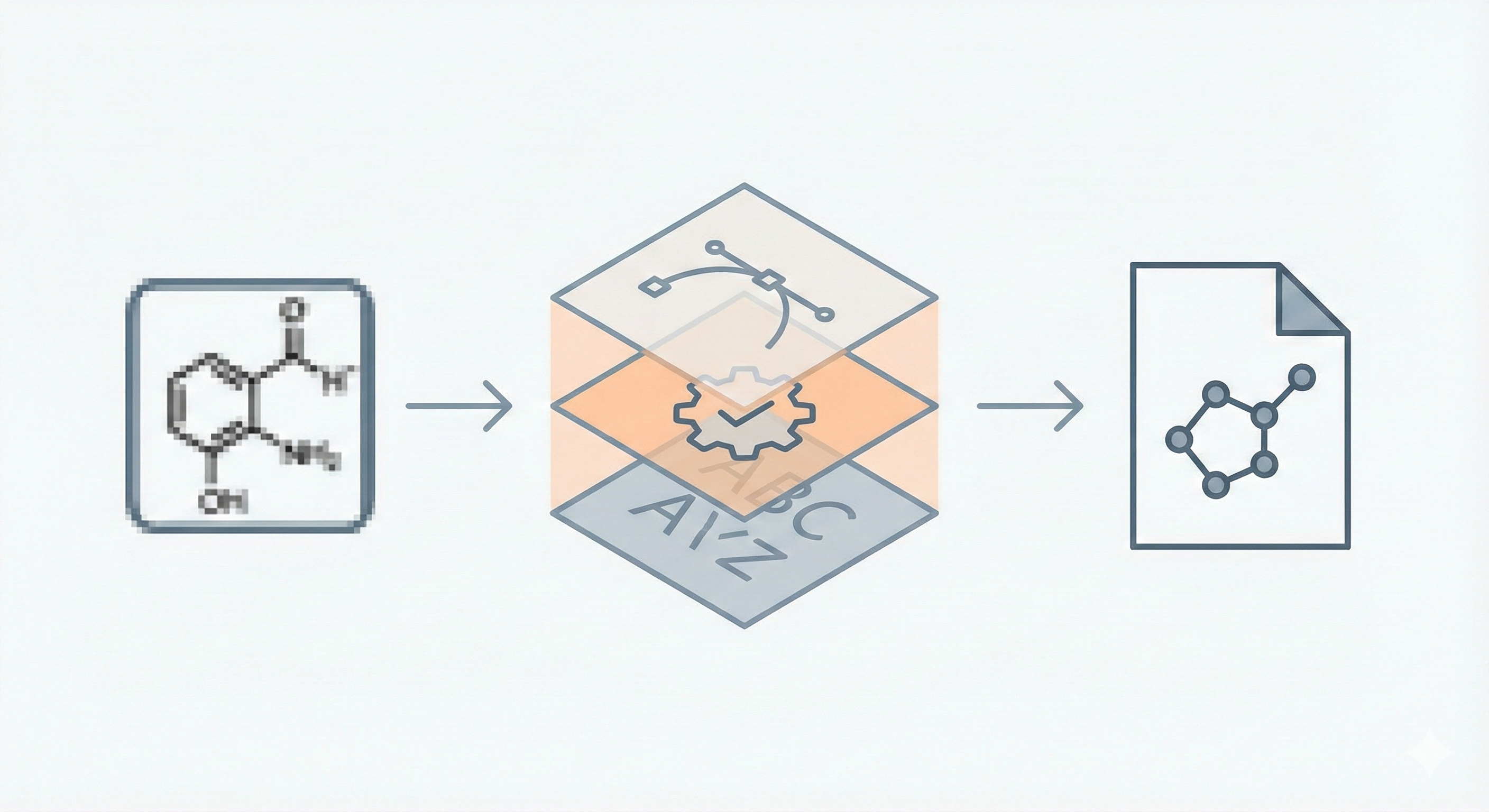
Automatic Recognition of Chemical Images
This methodological paper presents a system for digitizing chemical images into SDF files. It utilizes a custom vectorization algorithm and chemical rule validation, achieving 94% accuracy on benchmark datasets compared to 50% for commercial tools.