MARCEL: Molecular Conformer Ensemble Learning Benchmark
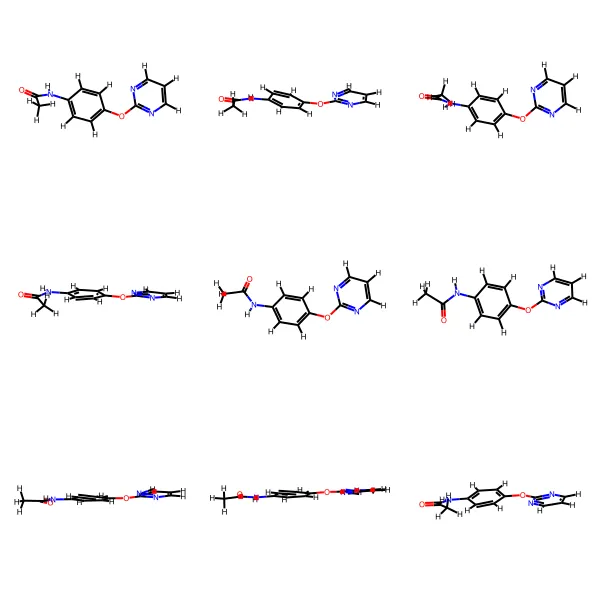
MARCEL provides a comprehensive benchmark for molecular representation learning with 722K+ conformers across four diverse subsets (Drugs-75K, Kraken, EE, BDE), enabling evaluation of conformer ensemble methods for property prediction in drug discovery and catalysis.