GuacaMol: Benchmarking Models for De Novo Molecular Design
GuacaMol provides an open-source benchmarking framework with 5 distribution-learning and 20 goal-directed tasks to standardize evaluation of de novo molecular design models.

GuacaMol provides an open-source benchmarking framework with 5 distribution-learning and 20 goal-directed tasks to standardize evaluation of de novo molecular design models.
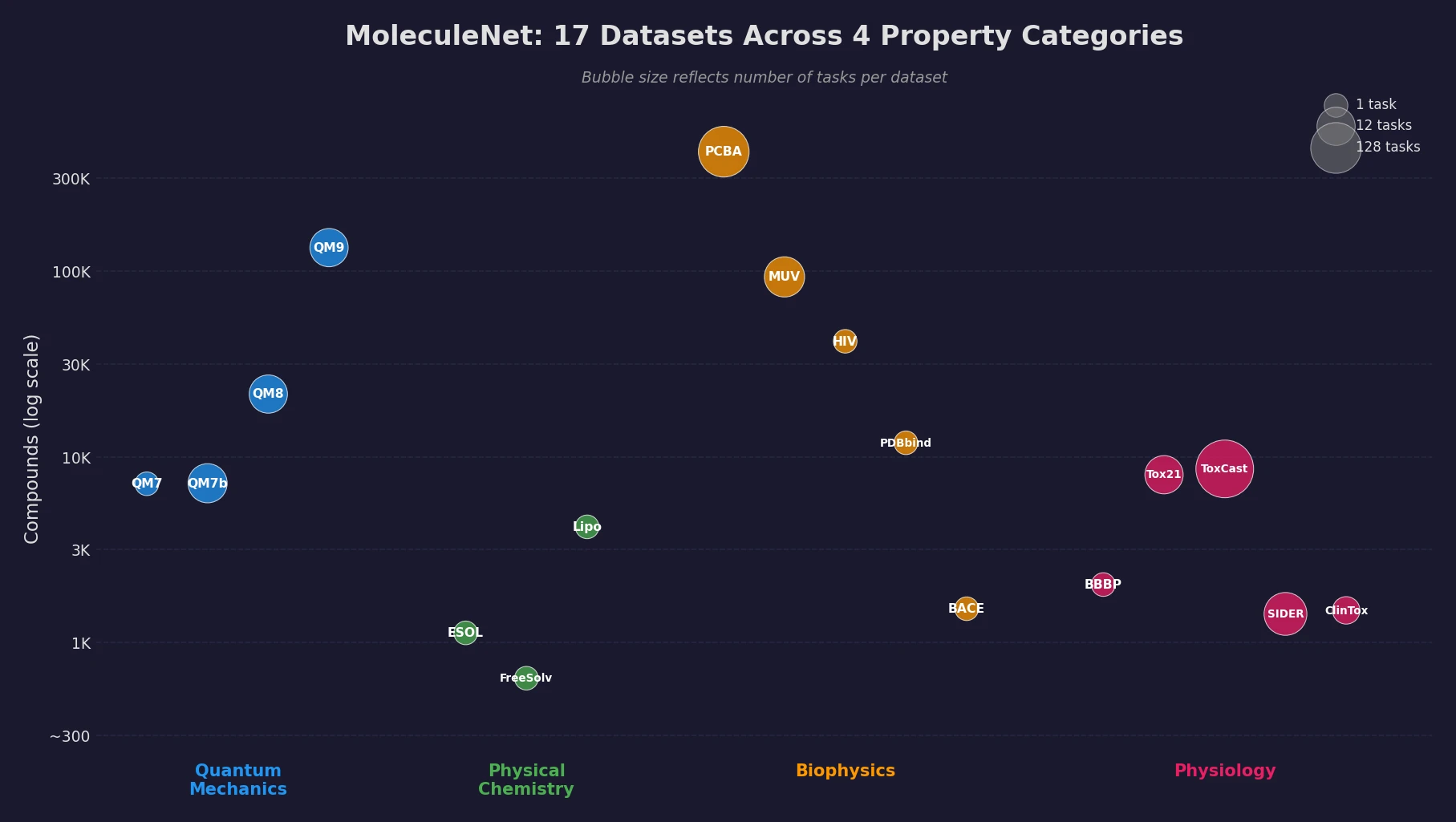
MoleculeNet introduces a large-scale benchmark suite for molecular machine learning, curating over 700,000 compounds across 17 datasets with standardized metrics, data splits, and featurization methods integrated into the DeepChem open-source library.
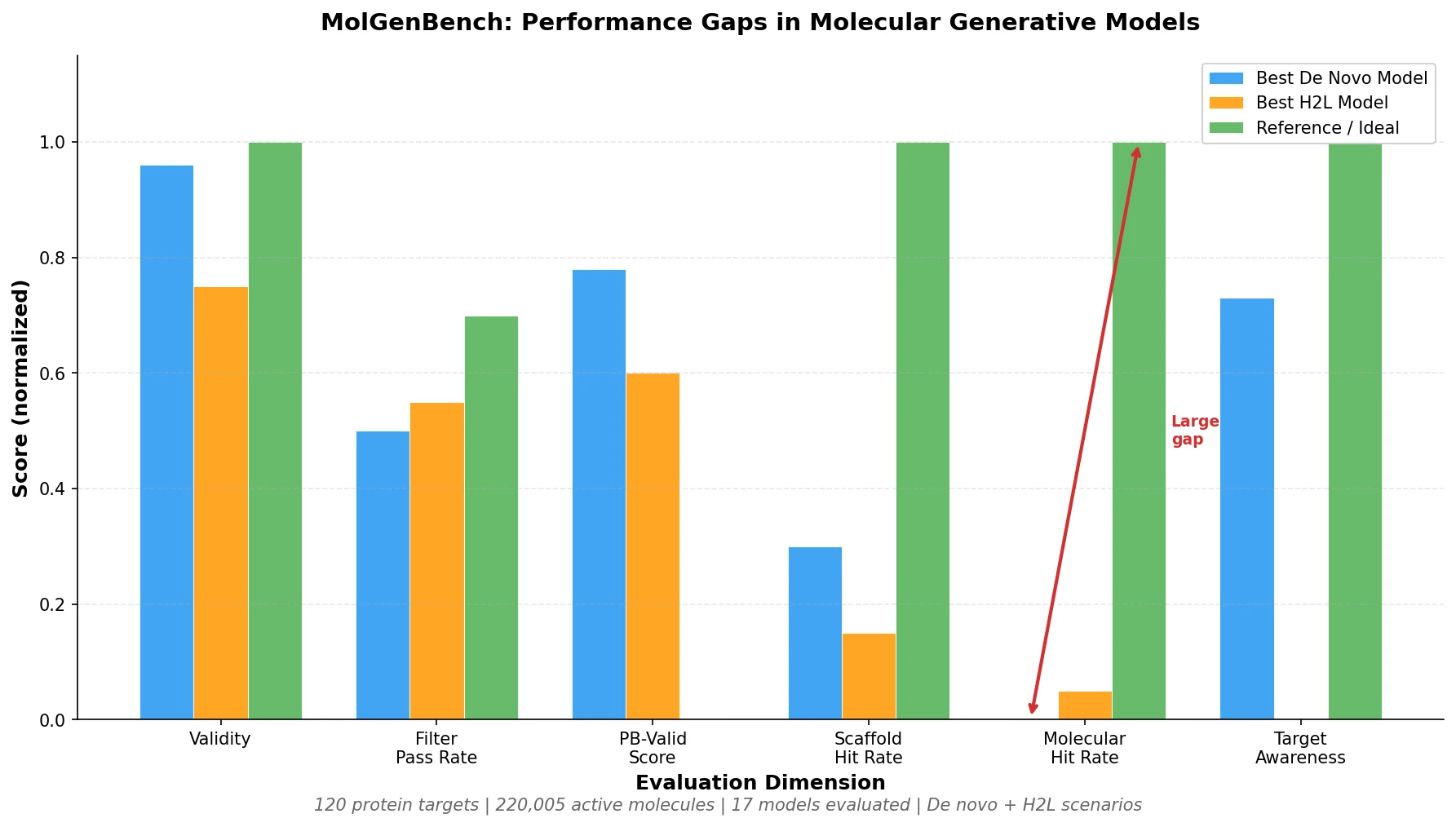
MolGenBench introduces a comprehensive benchmark for evaluating molecular generative models in realistic drug discovery settings, spanning de novo design and hit-to-lead optimization across 120 protein targets with 220,005 experimentally validated actives.
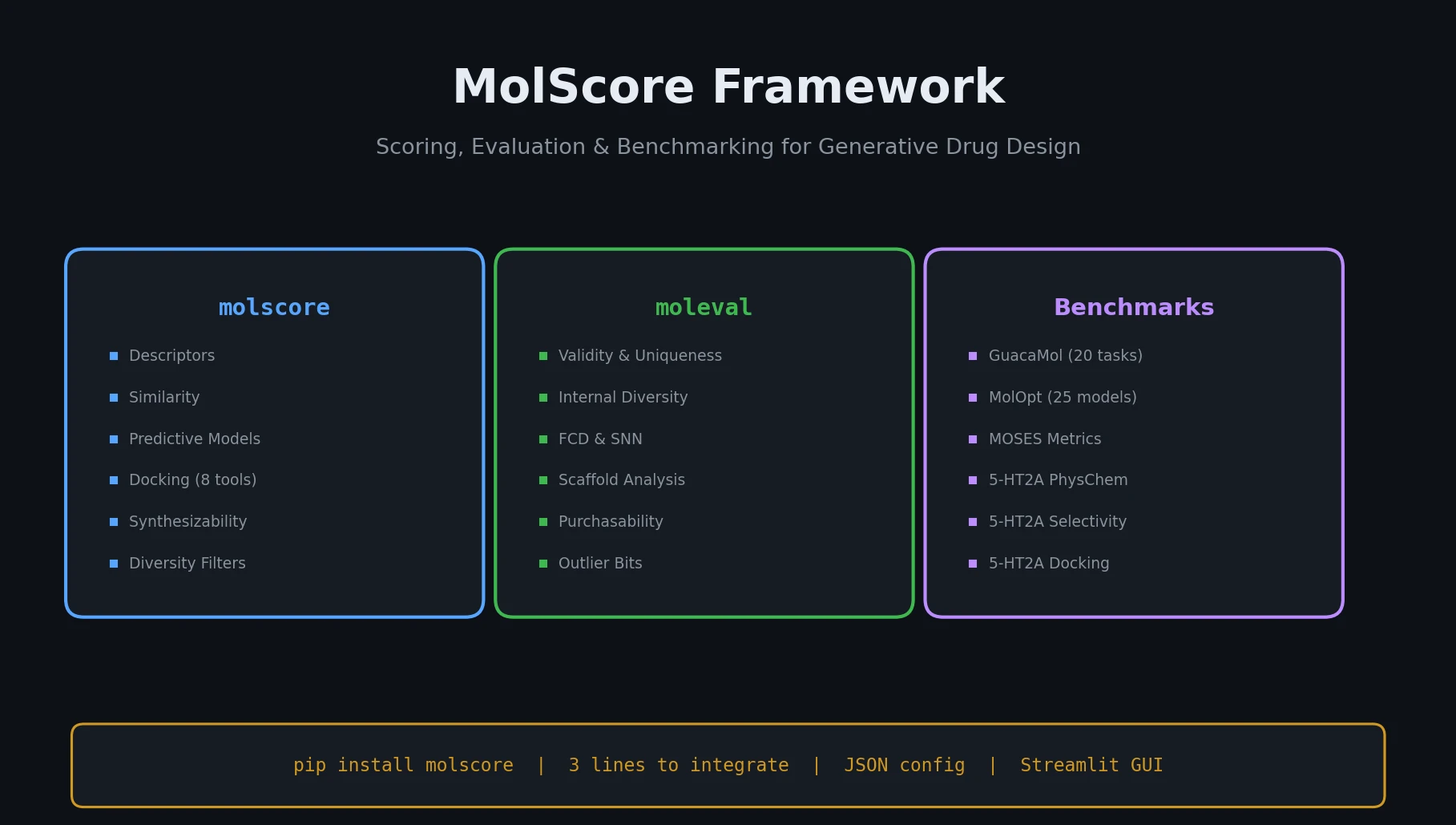
MolScore is an open-source framework that unifies scoring functions, evaluation metrics, and benchmarks for generative molecular design, with configurable objectives and GUI support.
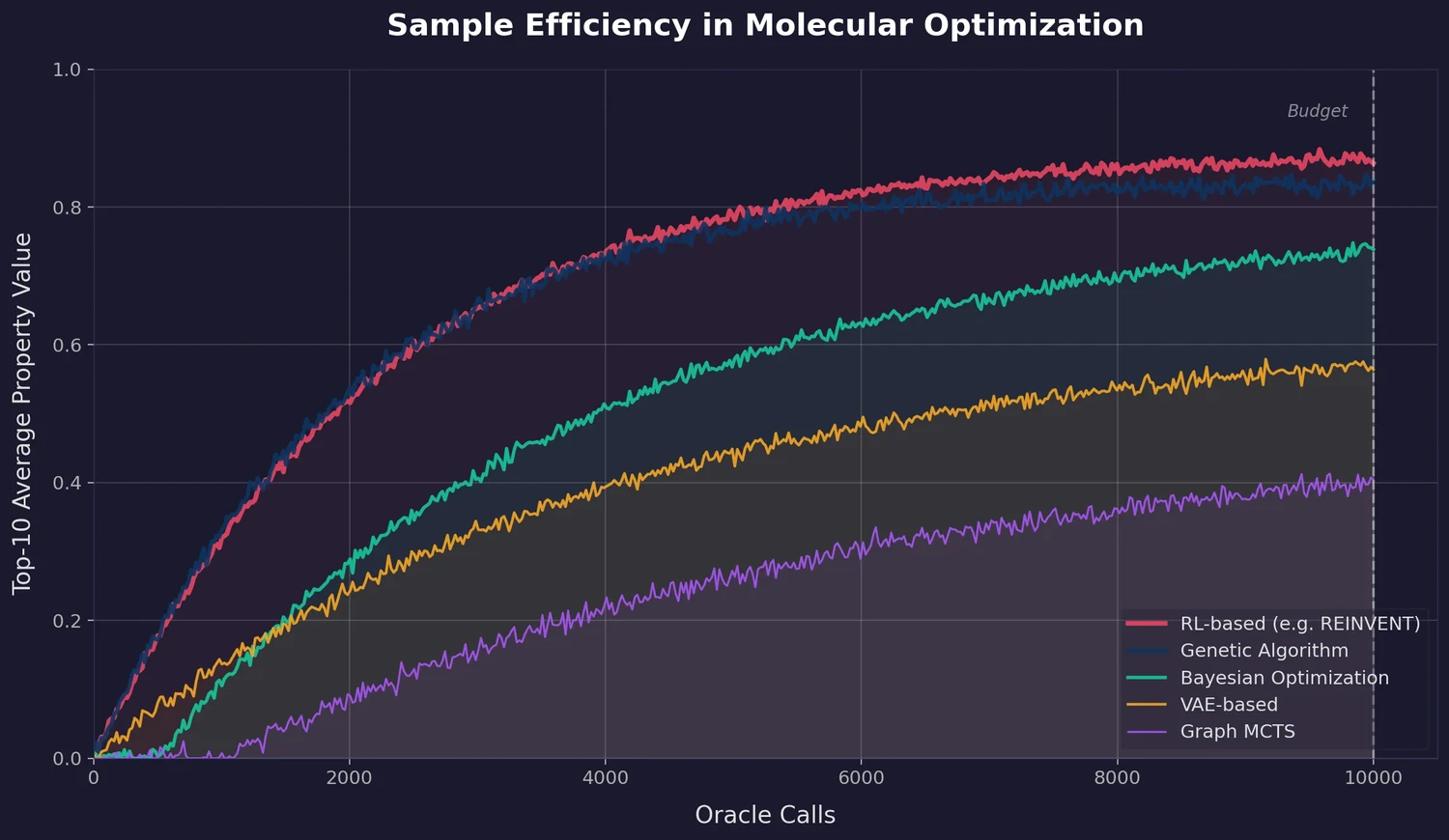
A large-scale benchmark of 25 molecular optimization methods on 23 oracles under constrained oracle budgets, showing that sample efficiency is a critical and often neglected dimension of evaluation.
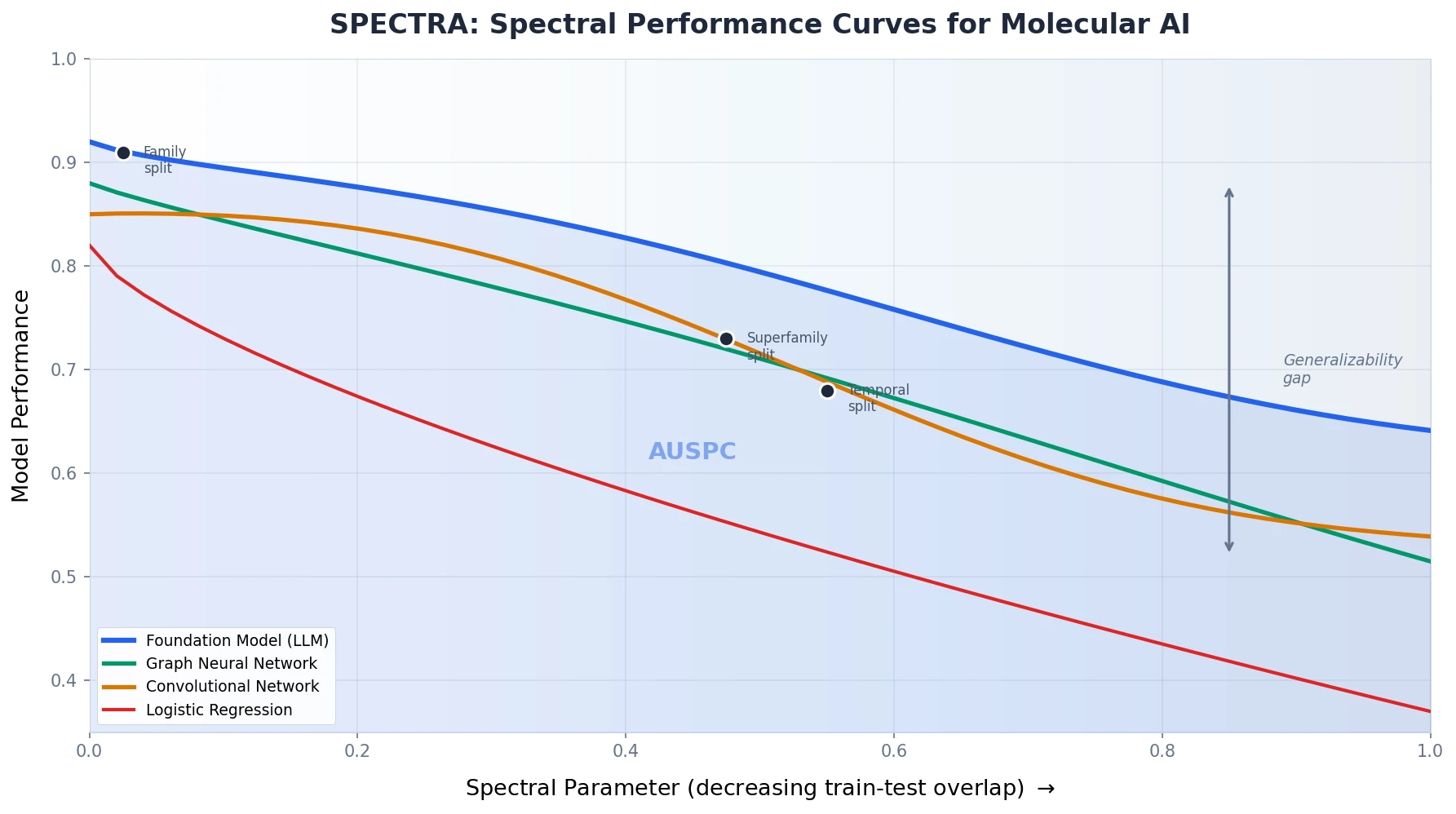
Introduces SPECTRA, a framework that generates spectral performance curves to measure how ML model accuracy degrades as train-test overlap decreases across molecular sequencing tasks.
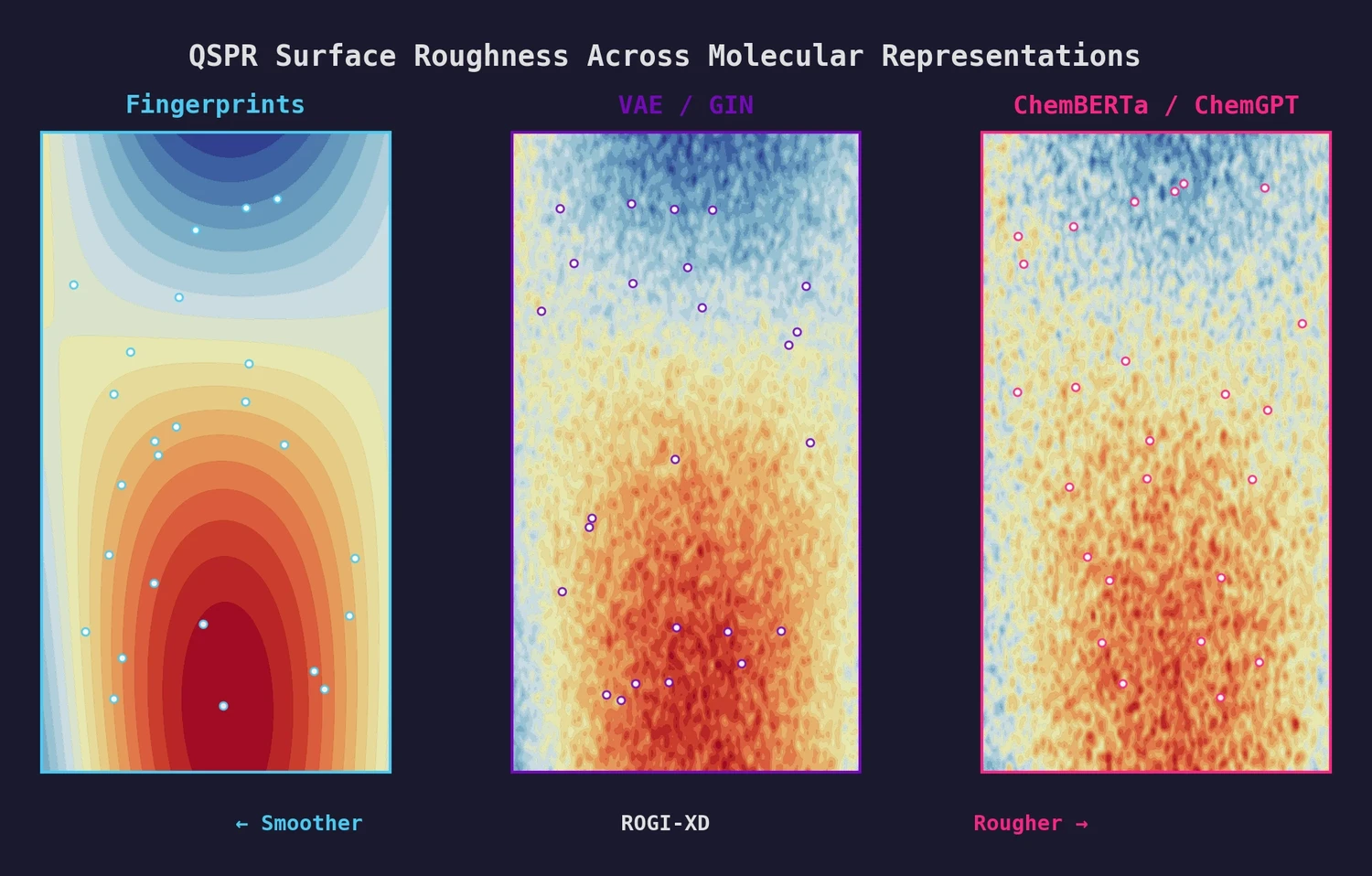
This paper introduces ROGI-XD, a reformulation of the ROuGhness Index that enables fair comparison of QSPR surface roughness across molecular representations of different dimensionalities. Evaluating VAE, GIN, ChemBERTa, and ChemGPT representations, the authors show that pretrained chemical models do not produce smoother structure-property landscapes than simple molecular fingerprints or descriptors.
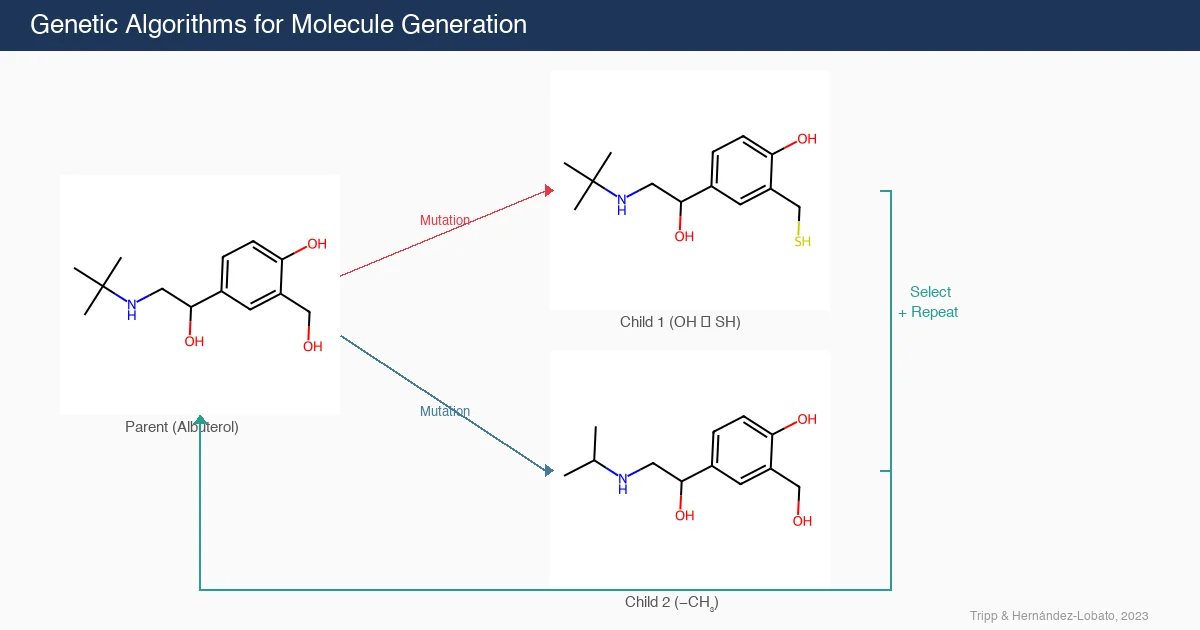
This position paper demonstrates that genetic algorithms (GAs) perform surprisingly well on molecular generation benchmarks, often outperforming complex deep learning methods. The authors propose the GA criterion: new molecule generation algorithms should demonstrate a clear advantage over GAs.
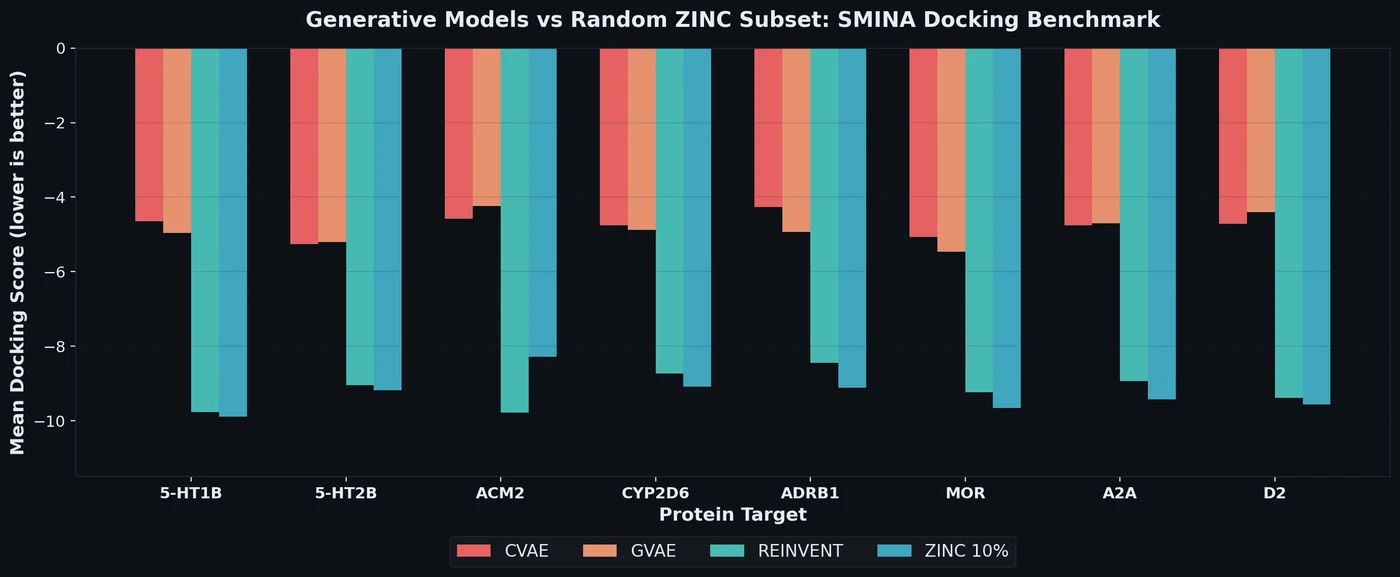
Proposes a benchmark for de novo drug design using SMINA docking scores across eight drug targets, revealing that popular generative models fail to outperform random ZINC subsets.
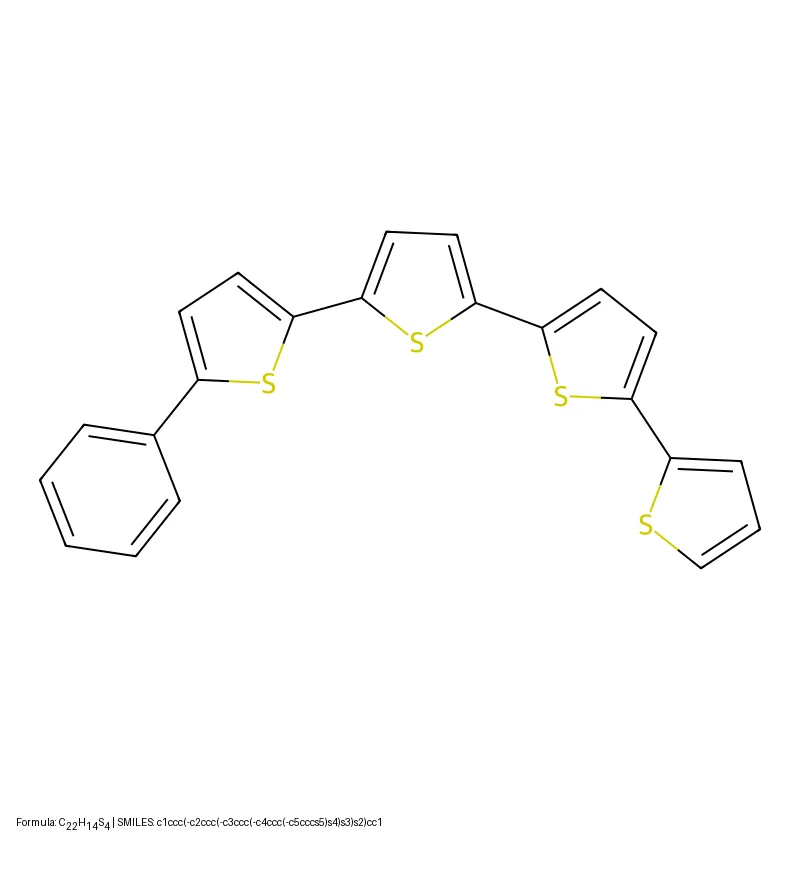
Tartarus introduces a modular suite of realistic molecular design benchmarks grounded in computational chemistry simulations. Benchmarking eight generative models reveals that no single algorithm dominates all tasks, and simple genetic algorithms often outperform deep generative models.
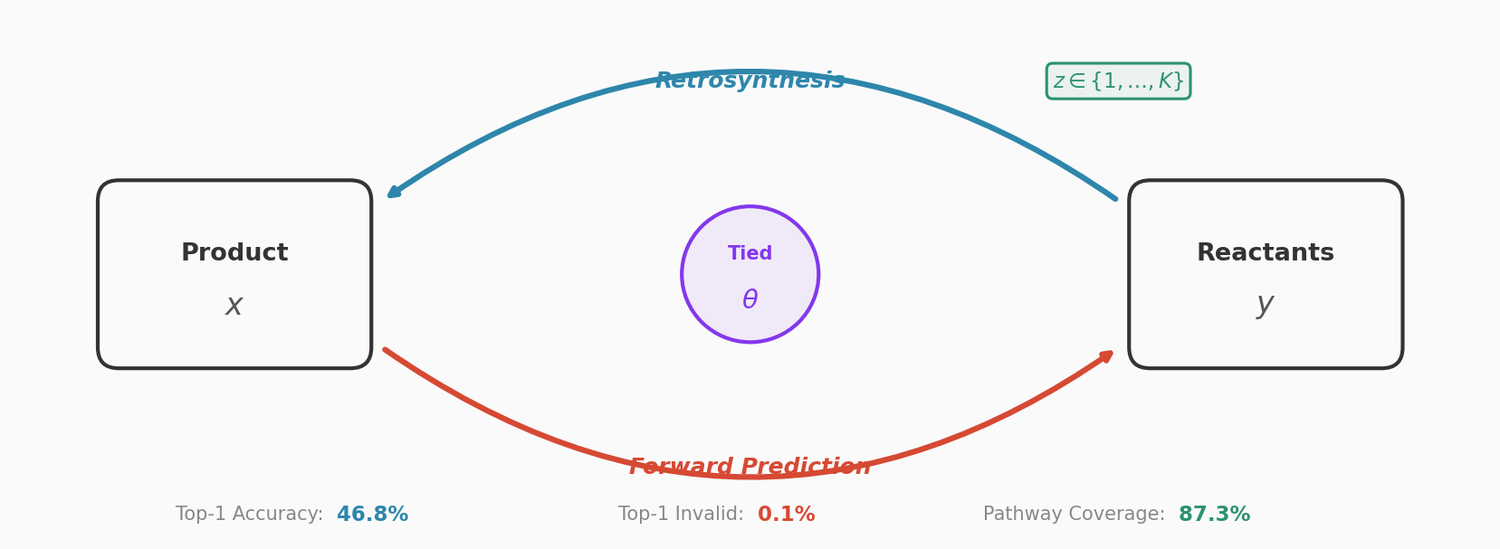
This paper couples a retrosynthesis transformer with a forward reaction transformer through parameter sharing, cycle consistency checks, and multinomial latent variables. The combined approach reduces top-1 SMILES invalidity to 0.1% on USPTO-50K, improves top-10 accuracy to 78.5%, and achieves 87.3% pathway coverage on a multi-pathway in-house dataset.
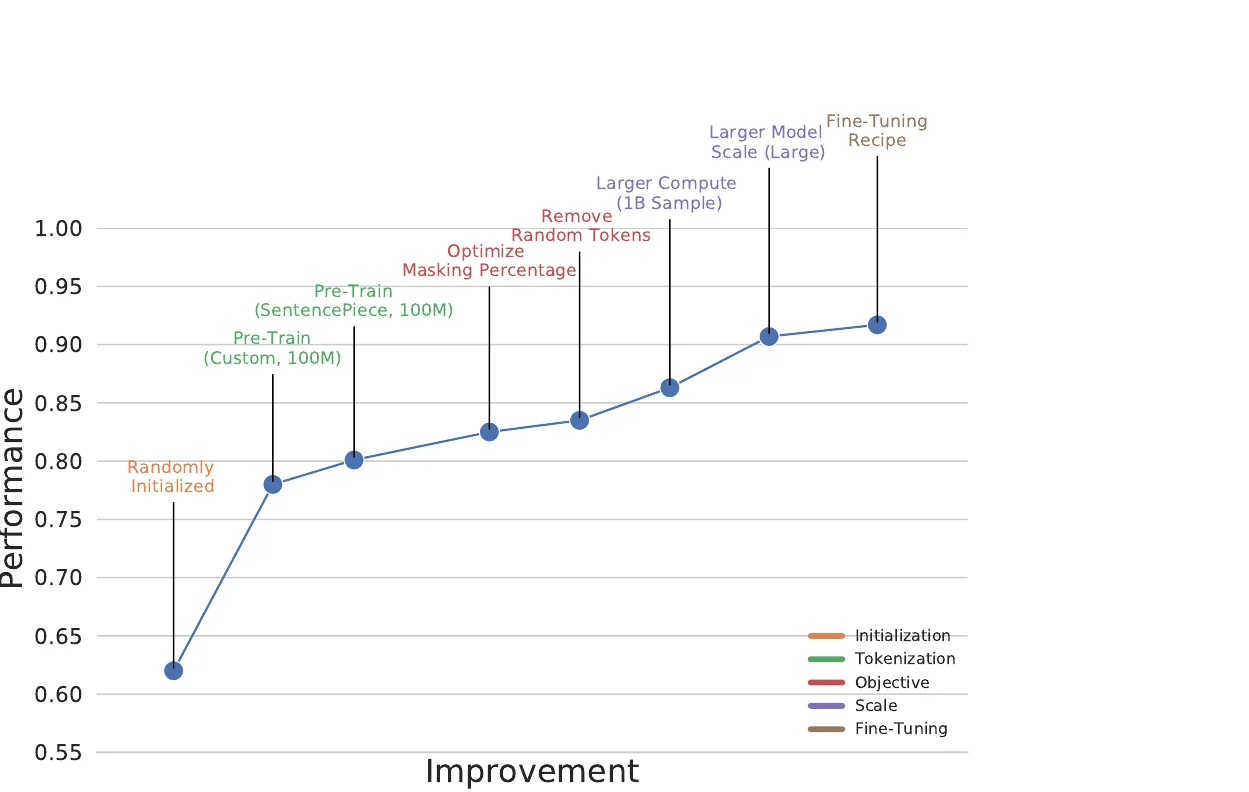
BARTSmiles pre-trains a BART-large model on 1.7 billion SMILES strings from ZINC20 and achieves the best reported results on 11 classification, regression, and generation benchmarks.