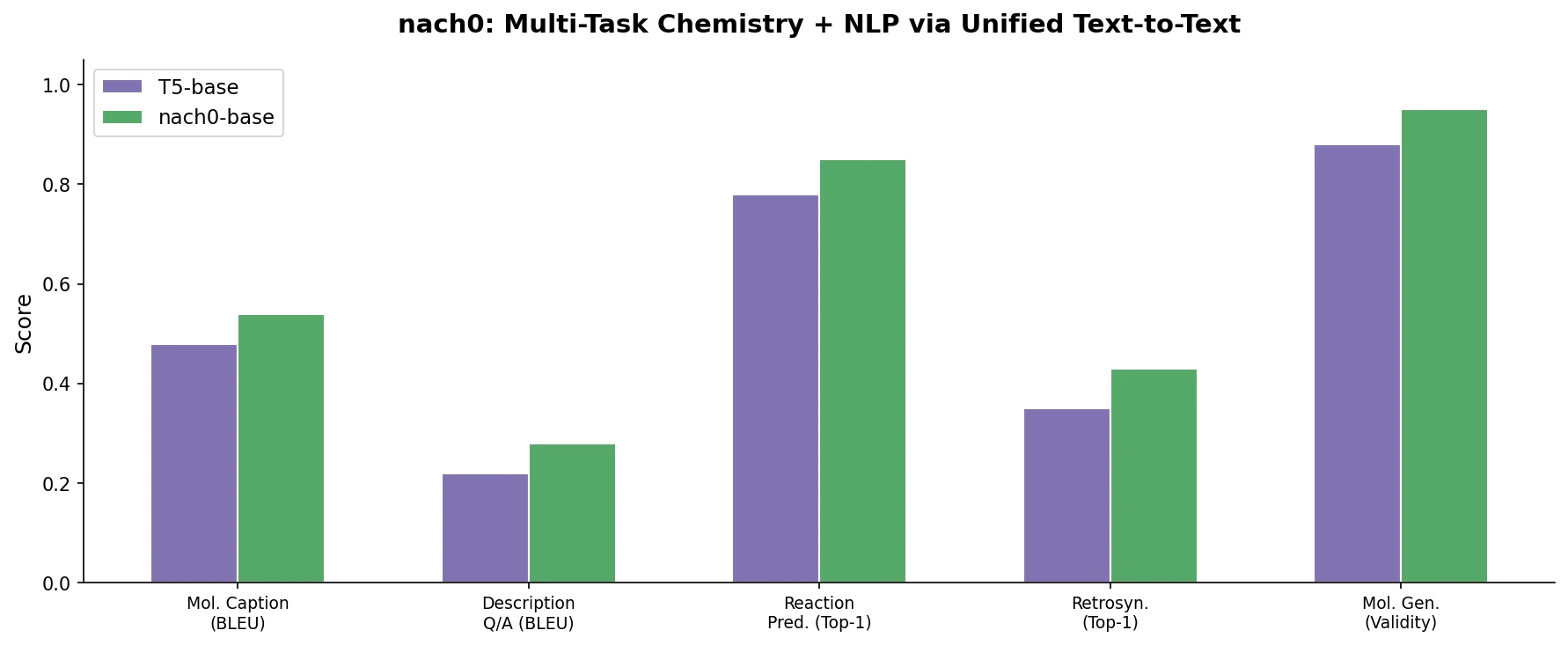
nach0: A Multimodal Chemical and NLP Foundation Model
nach0 unifies natural language and SMILES-based chemical tasks in a single encoder-decoder model, achieving competitive results across molecular property prediction, reaction prediction, molecular generation, and biomedical NLP benchmarks.