
GutenOCR: A Grounded Vision-Language Front-End for Documents
GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.

GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.
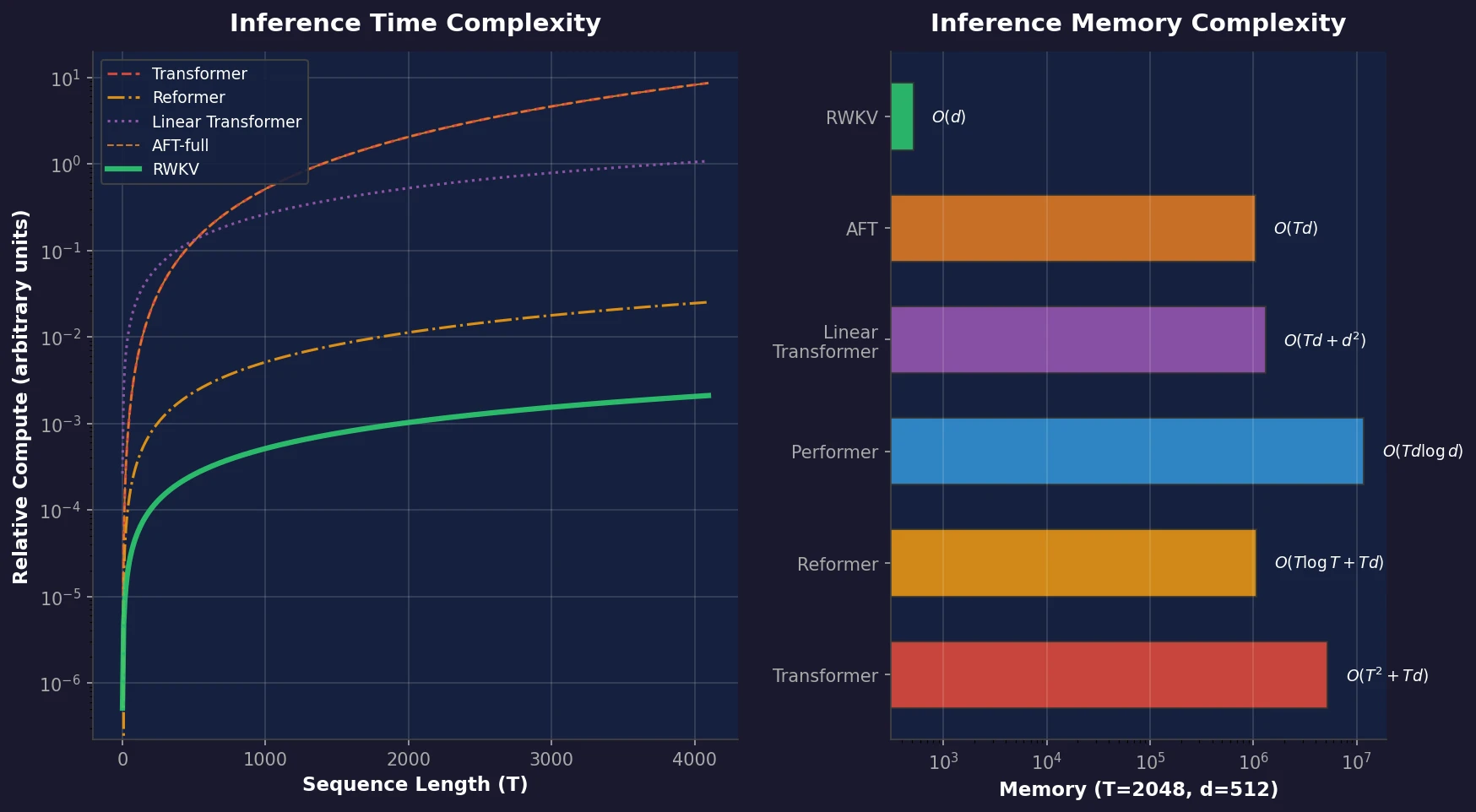
RWKV is a novel sequence model that achieves transformer-level performance while maintaining linear time and constant memory complexity during inference, scaled up to 14 billion parameters.
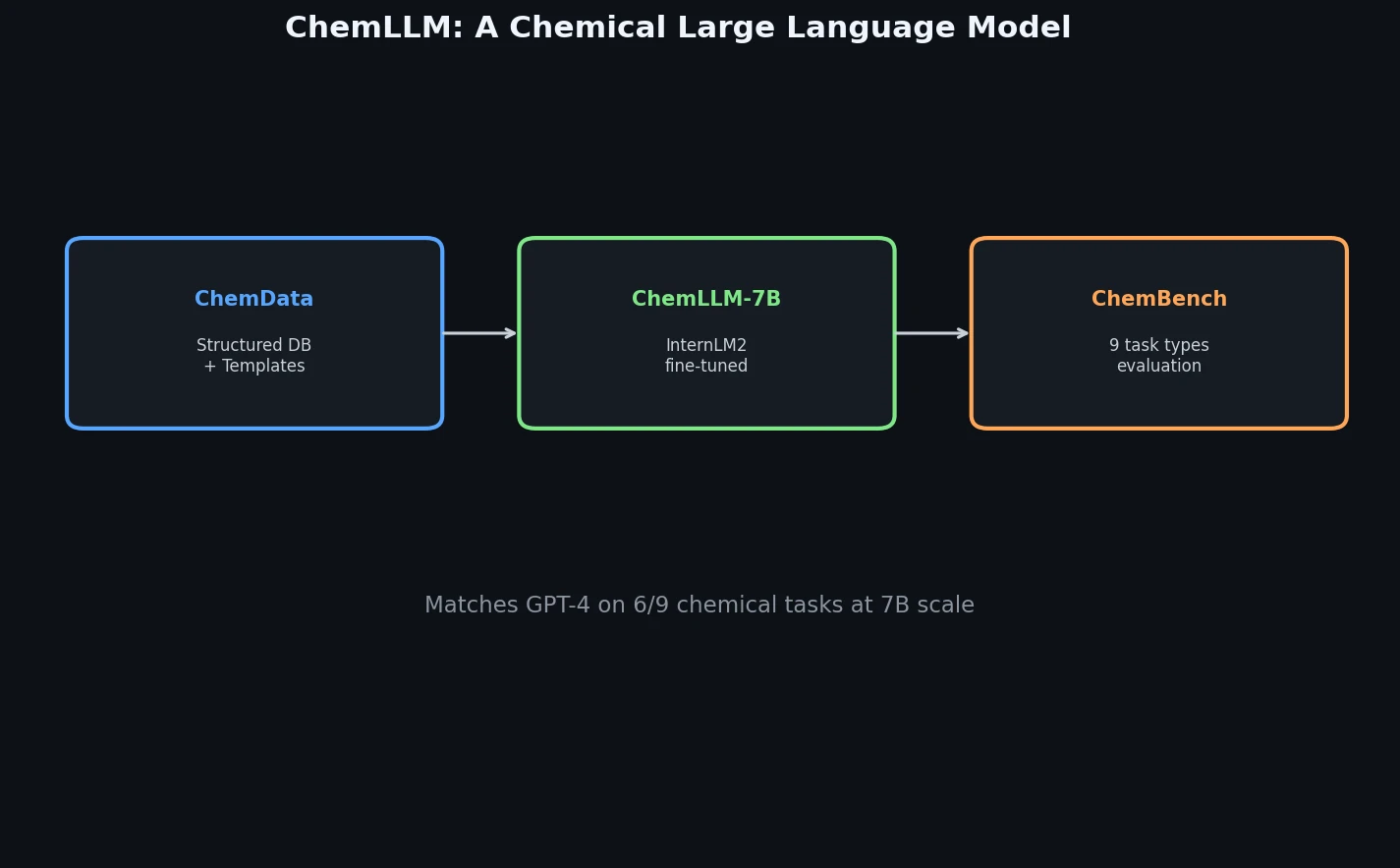
ChemLLM presents a comprehensive framework for chemistry-specific language modeling, including a 7M-sample instruction tuning dataset (ChemData), a 4,100-question benchmark (ChemBench), and a two-stage fine-tuned model that matches GPT-4 on core chemical tasks.
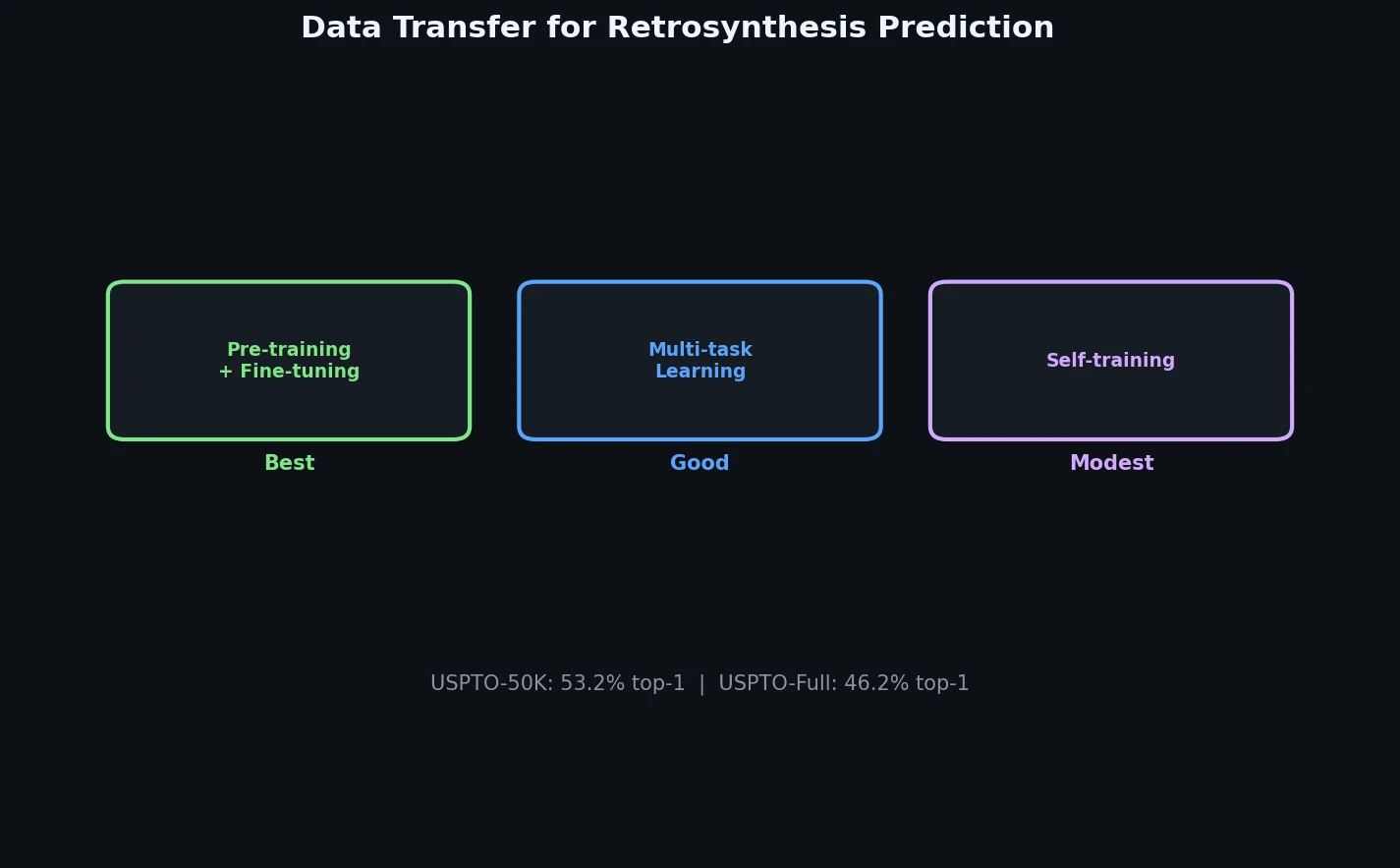
A systematic study of data transfer techniques (joint training, self-training, pre-training plus fine-tuning) applied to Transformer-based retrosynthesis. Pre-training on USPTO-Full followed by fine-tuning on USPTO-50K achieves the best results, improving top-1 accuracy from 35.3% to 57.4%.
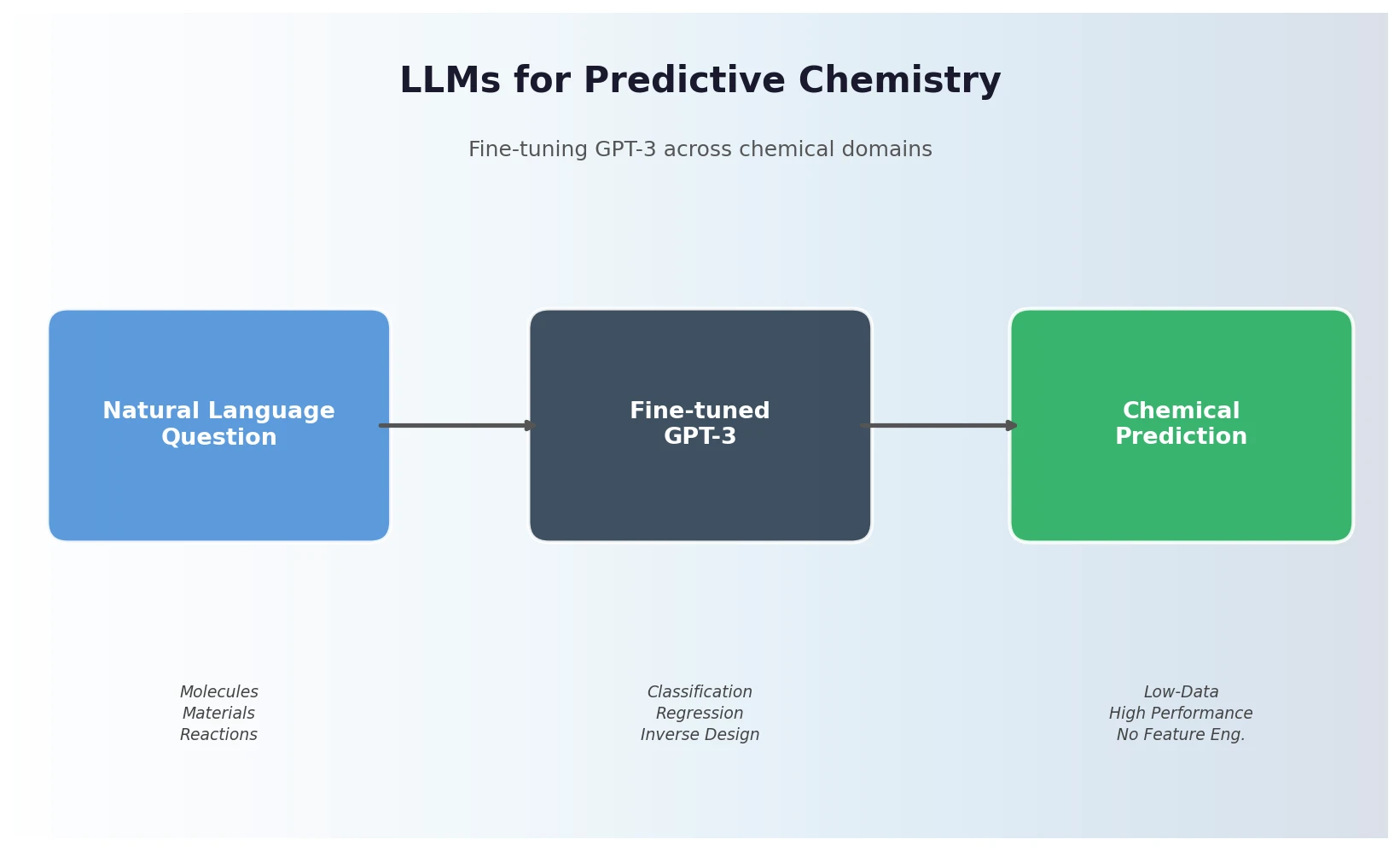
Jablonka et al. show that fine-tuning GPT-3 on natural language chemistry questions achieves competitive or superior performance to dedicated ML models across 15 benchmarks, with particular strength in low-data settings and inverse molecular design.
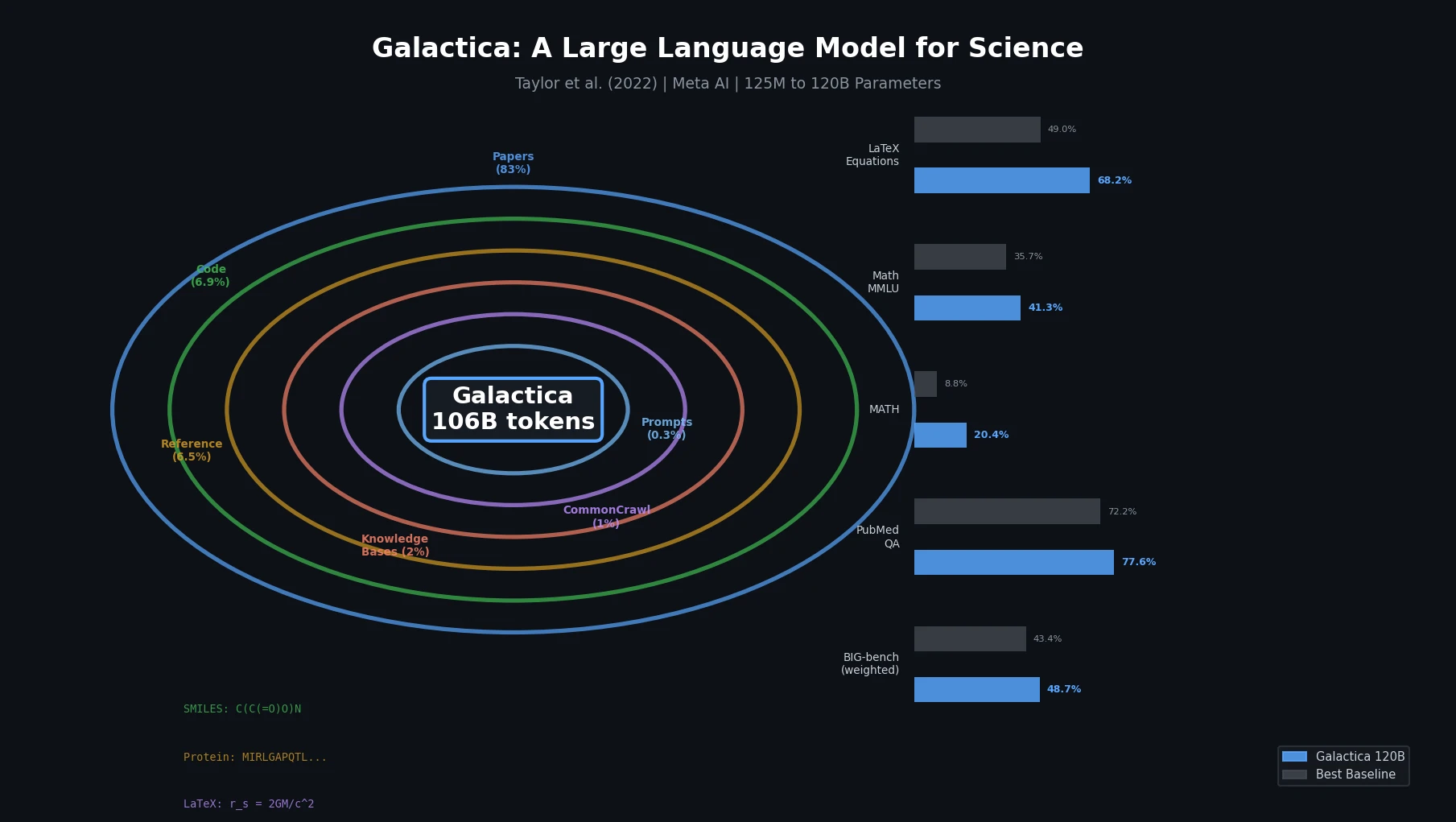
Galactica trains a decoder-only Transformer on a curated 106B-token scientific corpus spanning papers, proteins, and molecules, achieving strong results on scientific QA, mathematical reasoning, and citation prediction.
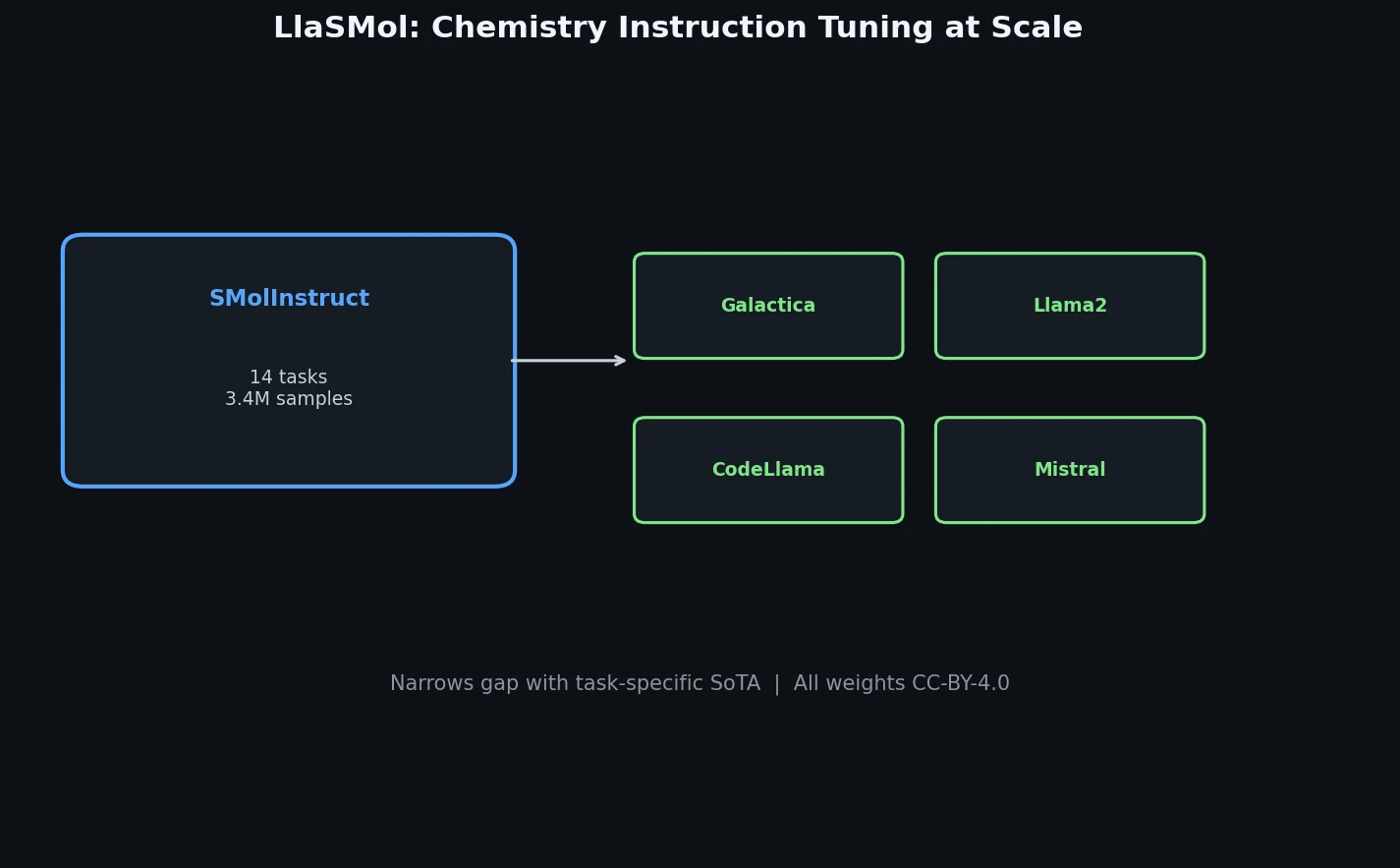
LlaSMol fine-tunes Mistral, Llama 2, and other open-source LLMs on SMolInstruct, a 3.3M-sample instruction tuning dataset covering 14 chemistry tasks. The Mistral-based model outperforms GPT-4 and Claude 3 Opus across all tasks.
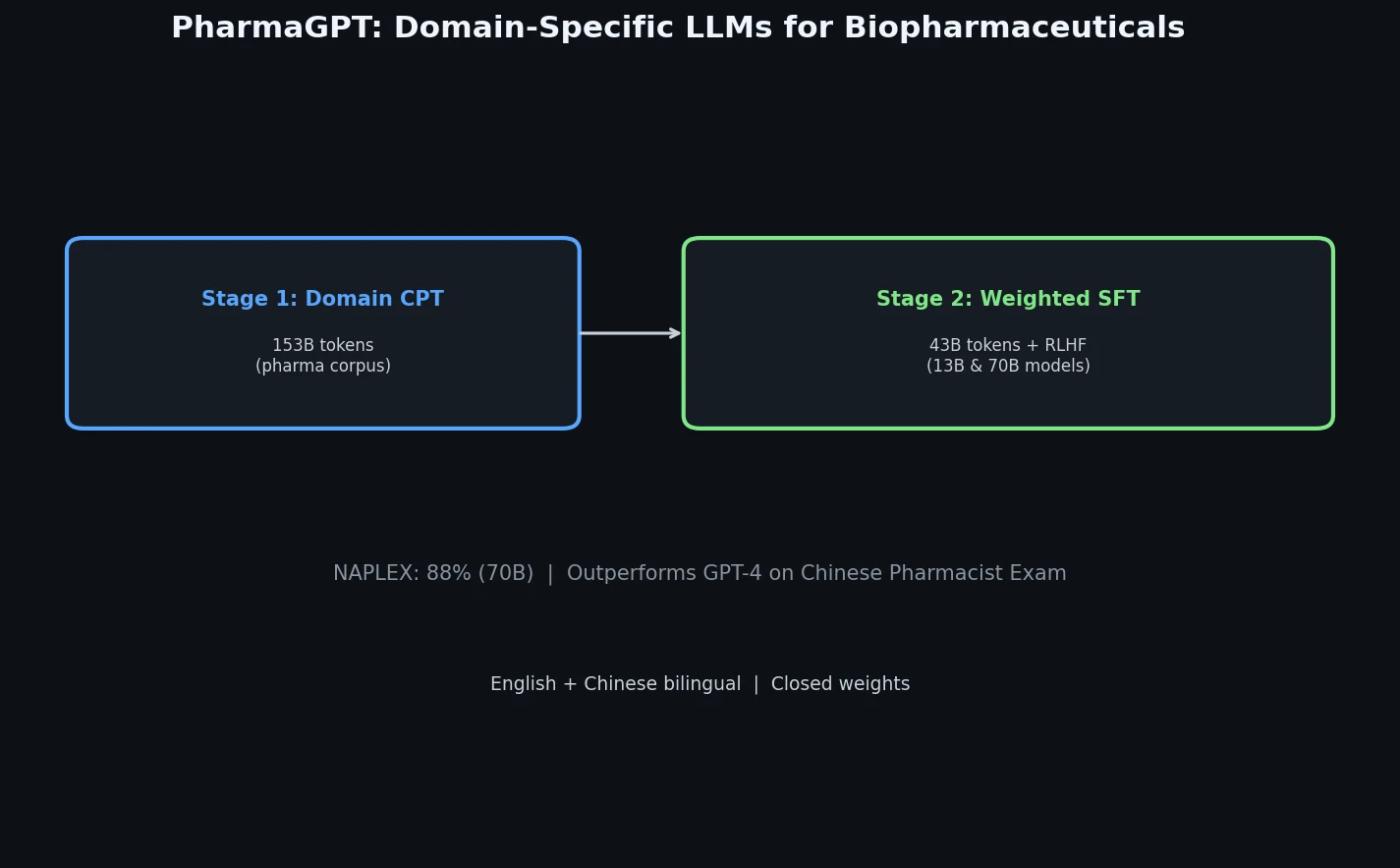

PharmaGPT is a suite of domain-specific LLMs (13B and 70B parameters) built on LLaMA with continued pretraining on biopharmaceutical and chemical data, achieving strong results on NAPLEX and Chinese pharmacist exams.
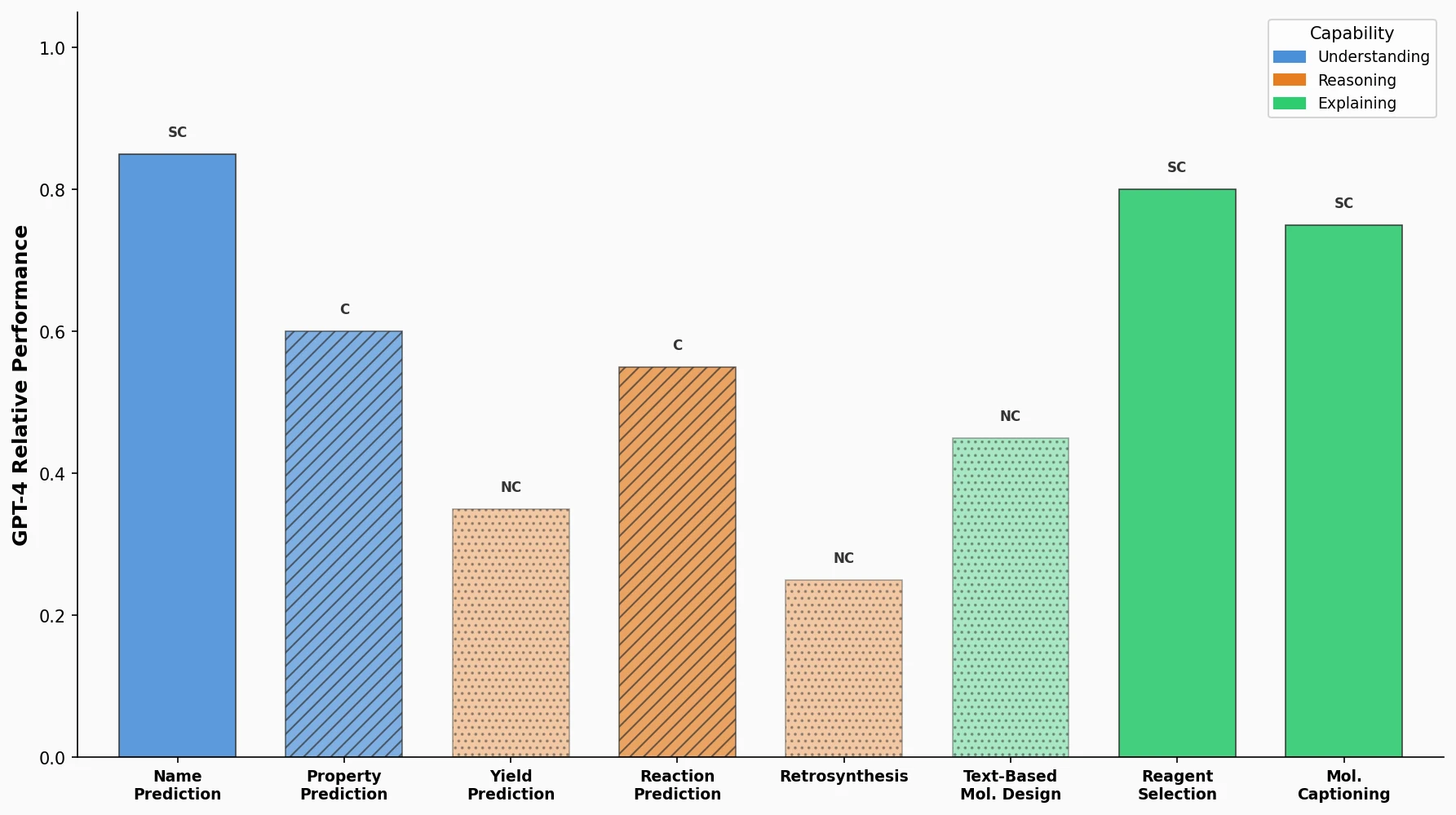
A comprehensive benchmark evaluating GPT-4, GPT-3.5, Davinci-003, Llama, and Galactica on eight practical chemistry tasks, revealing that LLMs are competitive on classification and text tasks but struggle with SMILES-dependent generation.
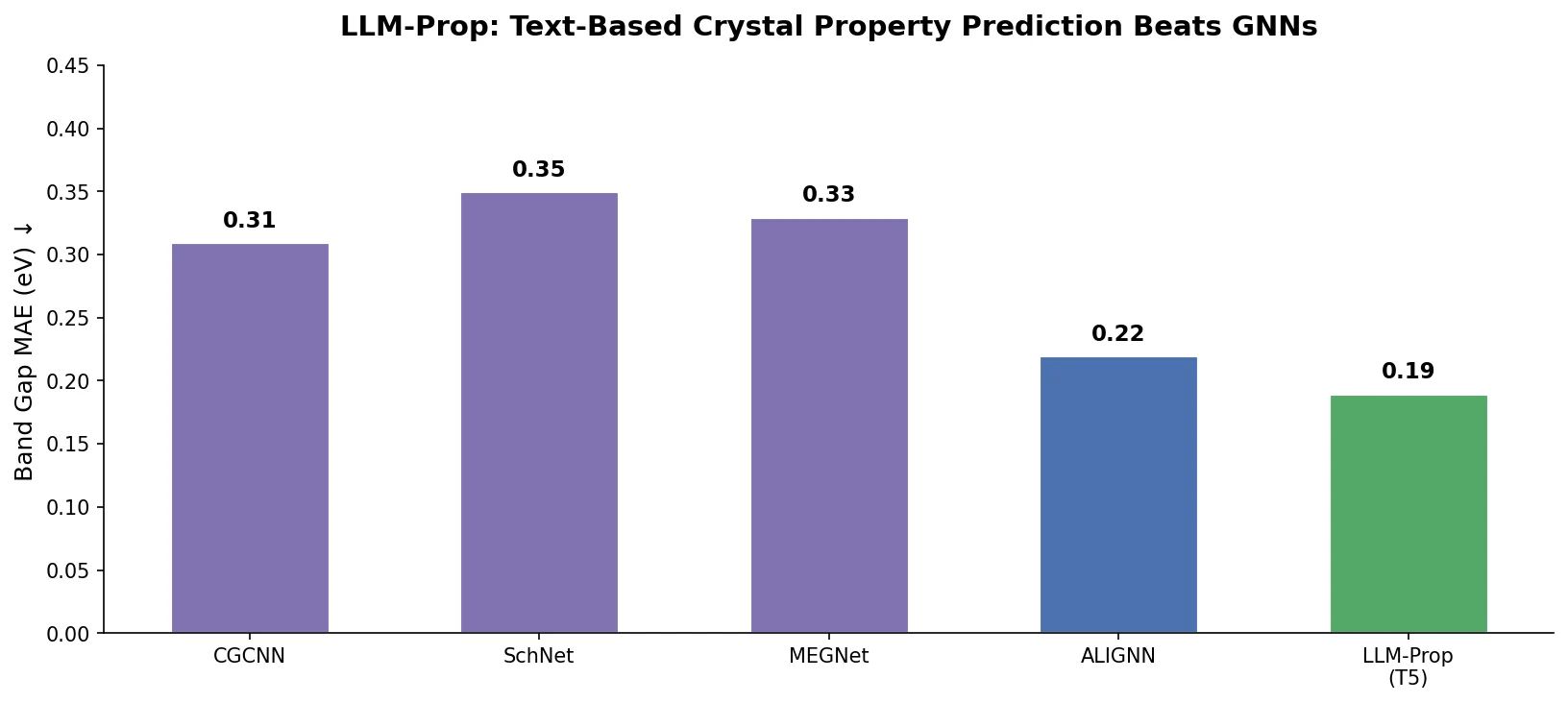
LLM-Prop uses the encoder half of T5, fine-tuned on Robocrystallographer text descriptions, to predict crystal properties. It outperforms GNN baselines like ALIGNN on band gap and volume prediction while using fewer parameters.
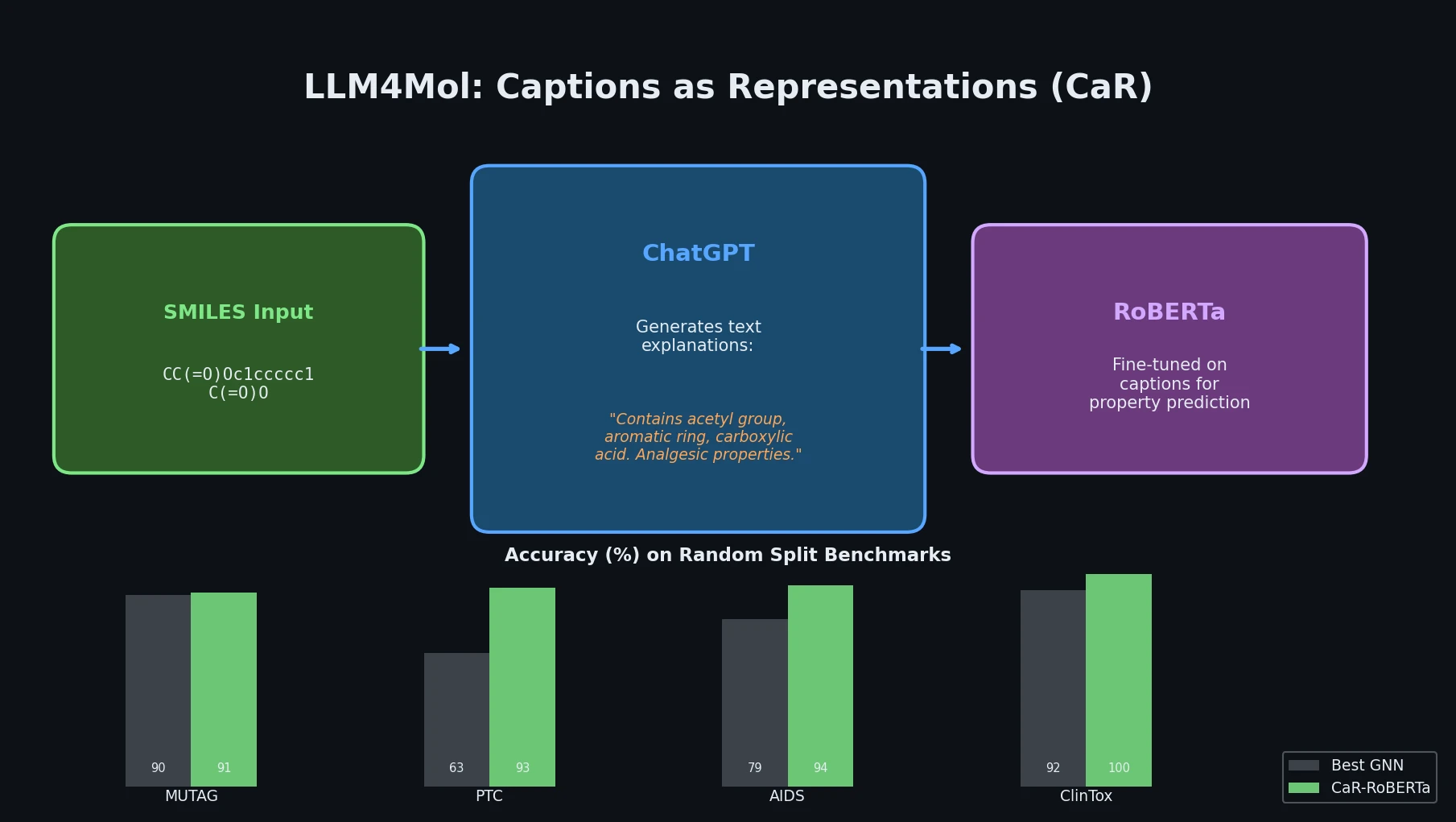
Proposes Captions as Representations (CaR), where ChatGPT generates textual explanations for SMILES strings that are then used to fine-tune small language models for molecular property prediction.
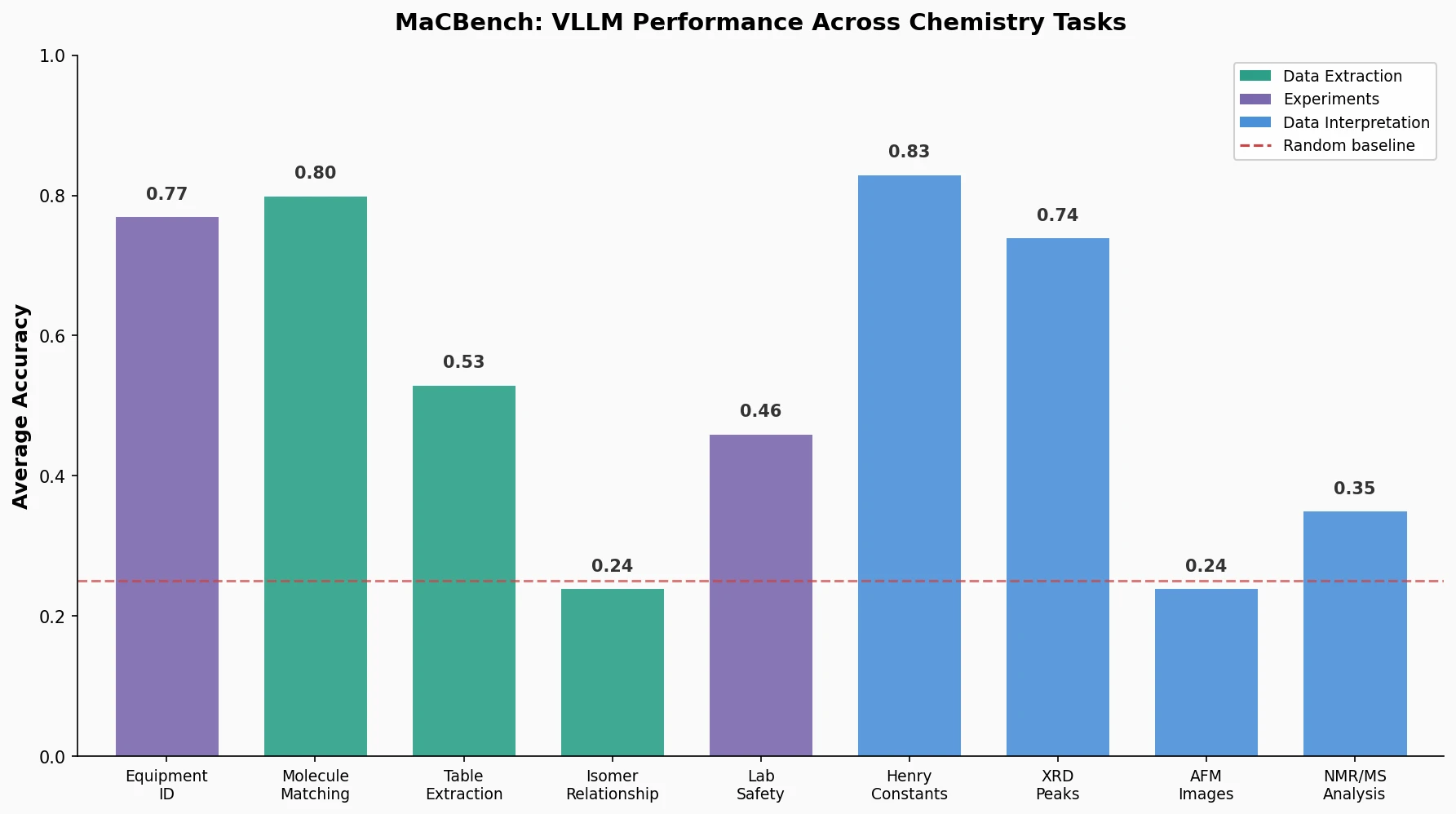
MaCBench evaluates frontier vision language models across 1,153 chemistry and materials science tasks spanning data extraction, experimental execution, and data interpretation, uncovering fundamental limitations in spatial reasoning and cross-modal integration.