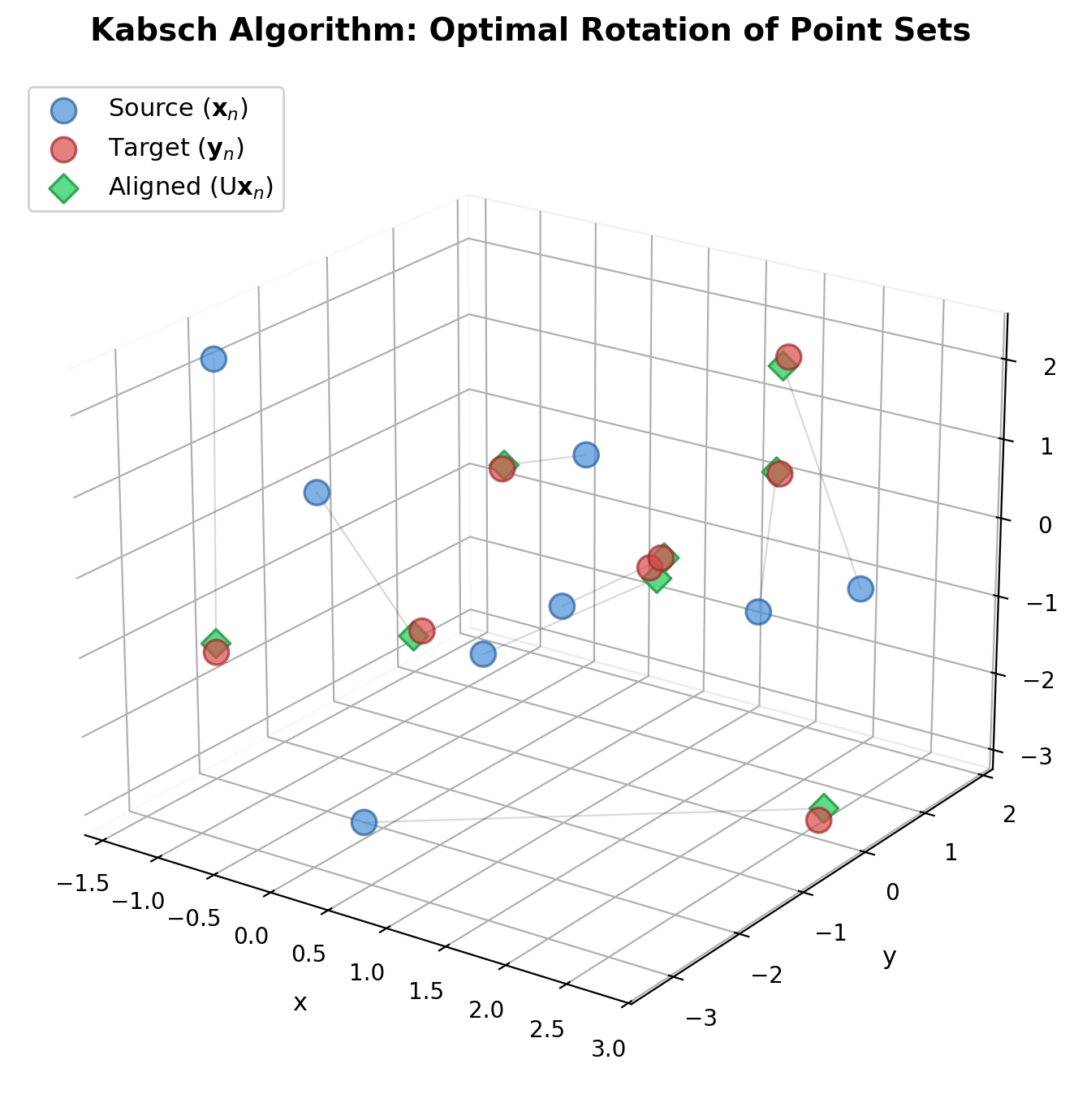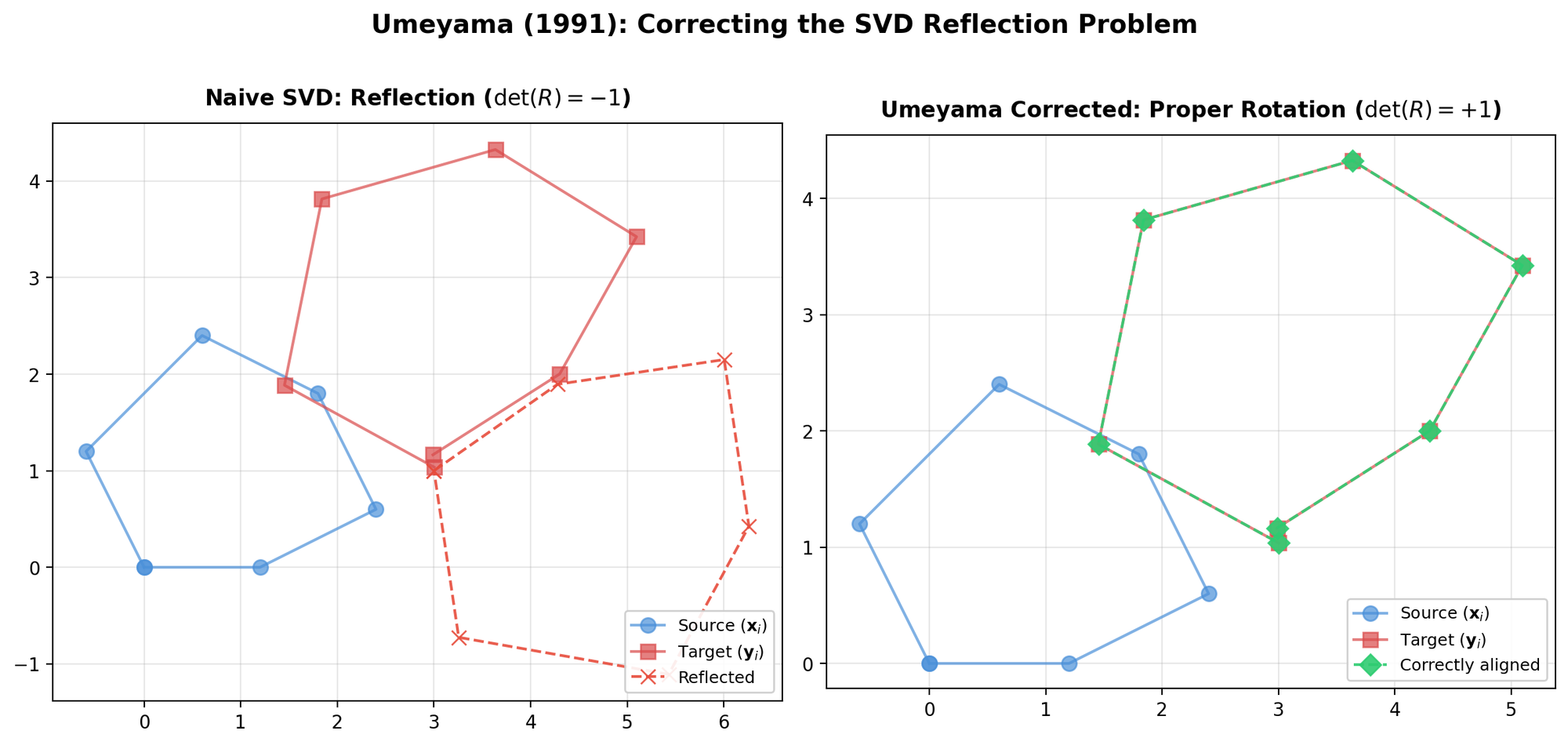
Umeyama's Method: Corrected SVD for Point Alignment
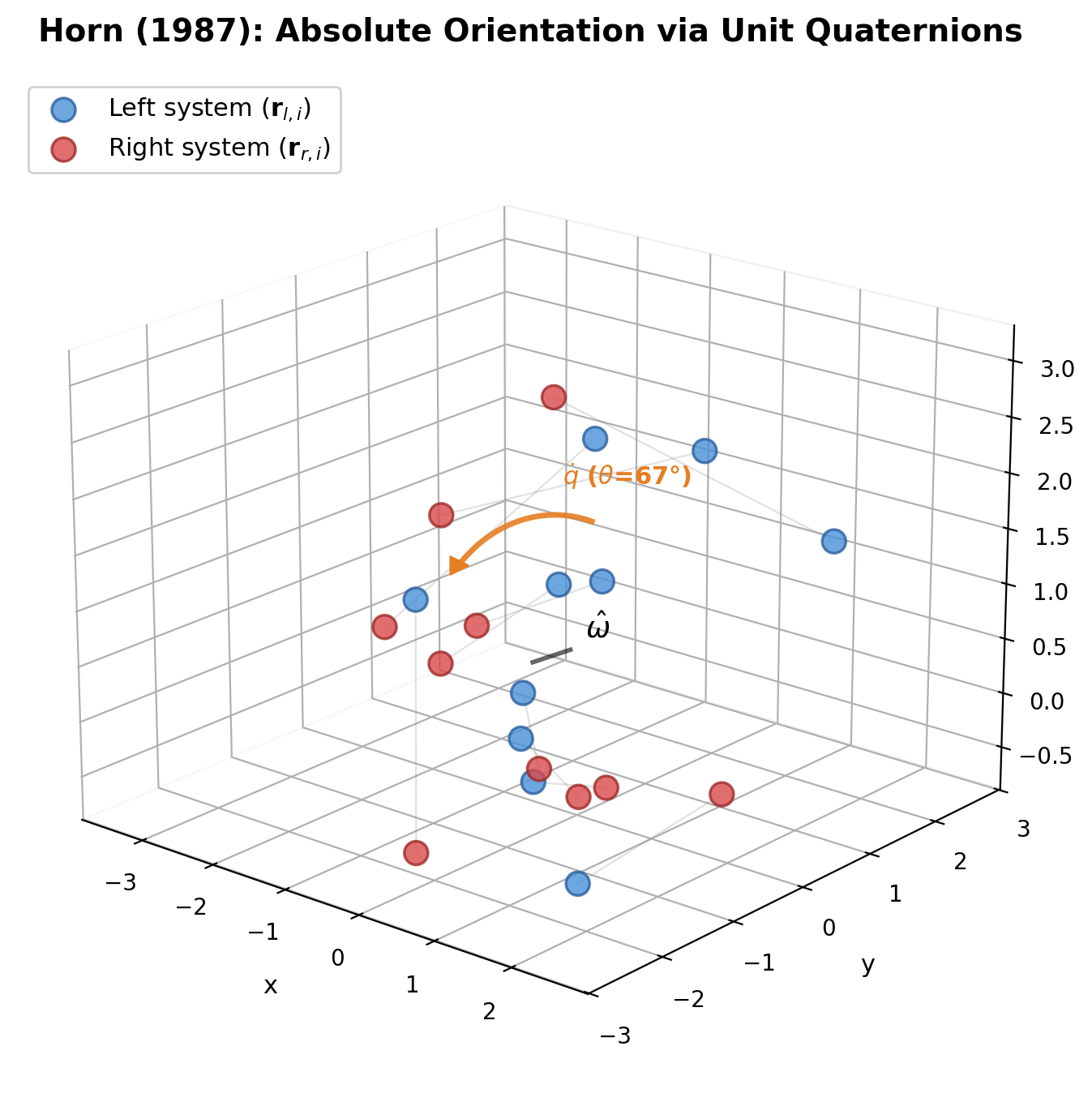
Corrects a flaw in prior SVD-based alignment methods (Arun et al., Horn et al.) that could produce reflections instead of rotations under noisy data, and provides a complete closed-form solution for similarity transformations in arbitrary dimensions.