Graph Grammar and ILP for Carbon Fixation Pathways
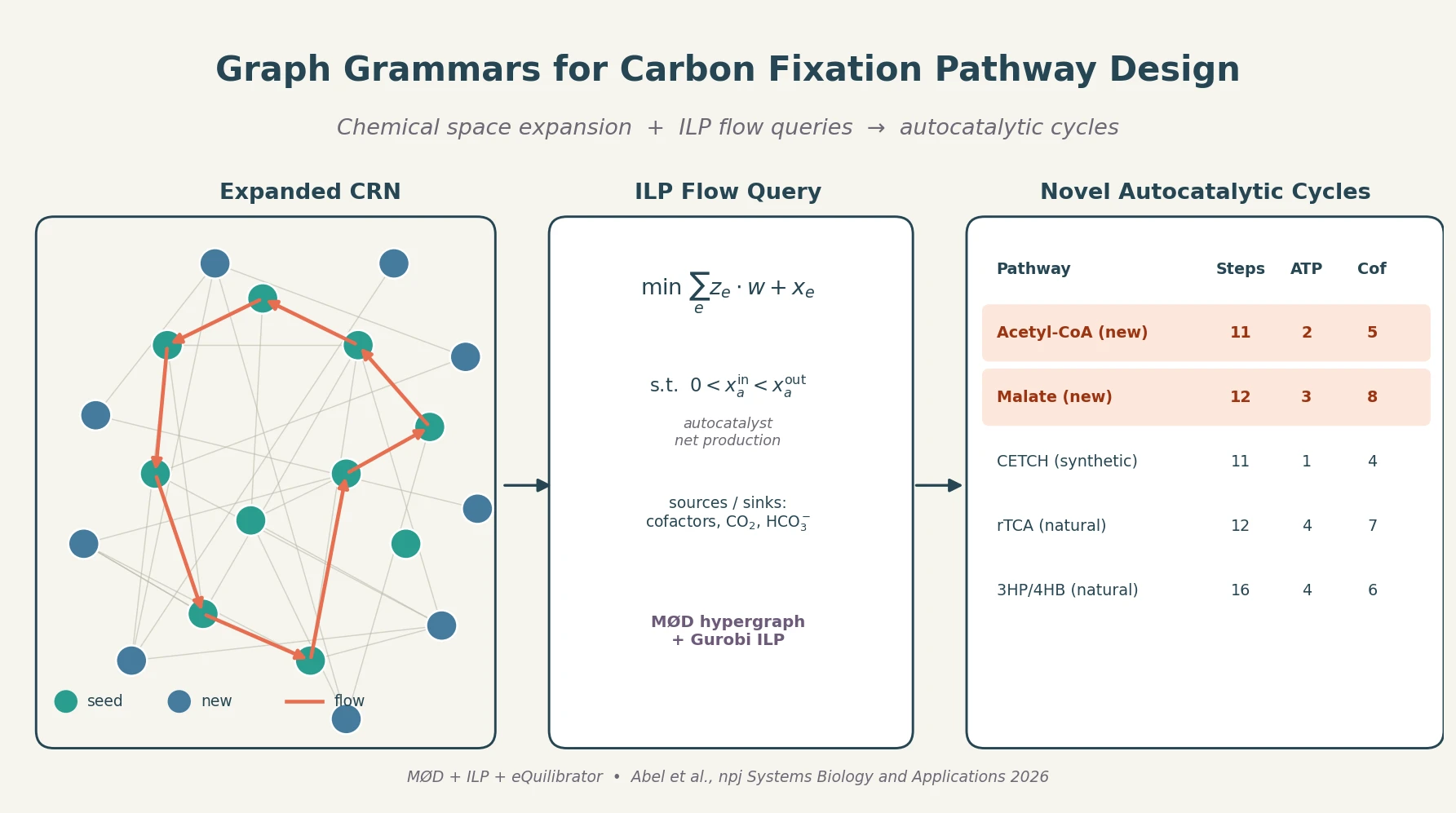
A graph-grammar cheminformatics workflow expands the carbon fixation reaction network, then uses integer linear programming flow queries to surface short autocatalytic pathways producing Acetyl-CoA and Malate with efficiencies approaching the CETCH cycle.