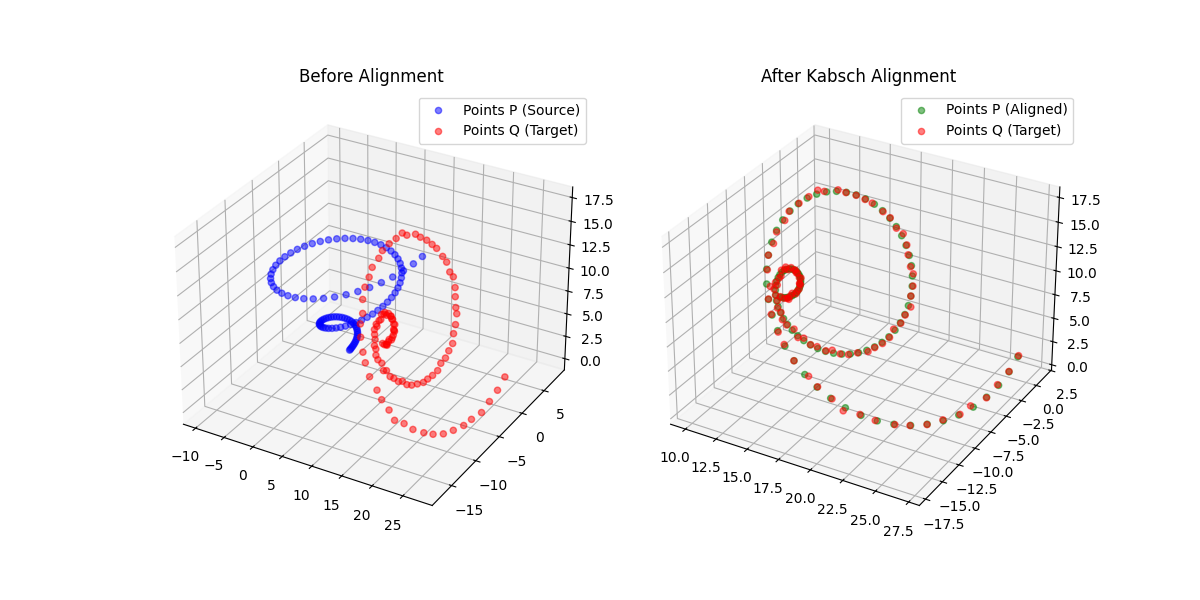
Overview
Aligning two sets of corresponding points, finding the optimal rotation (and optionally translation and scale) that maps one onto the other, is a fundamental operation across scientific computing. It appears in molecular dynamics (superimposing protein conformations), robotics (sensor registration), and computer vision (shape matching). The two dominant algorithm families are the Kabsch (SVD-based) method and the Horn (quaternion-based) method.
The Kabsch-Horn Cookbook is a Python library that implements both algorithm families across five numerical frameworks: NumPy, PyTorch, JAX, TensorFlow, and MLX. Every backend shares the same API, supports N-dimensional point sets, per-point weights, and arbitrary batch dimensions. The PyTorch, JAX, TensorFlow, and MLX backends are fully differentiable, with custom autograd rules that bypass the numerically unstable gradient of the standard SVD near degenerate singular values.
Features
Algorithms
- Kabsch: SVD-based optimal rotation for rigid alignment
- Kabsch-Umeyama: Kabsch with an additional optimal scaling factor $c$, solving $Q \approx cRP + t$
- Horn: Quaternion-based optimal rotation via the eigendecomposition of a $4 \times 4$ key matrix
- Horn + Scale: Horn’s method extended with optimal isotropic scaling
- RMSD Wrappers: Convenience functions that return RMSD directly alongside the alignment parameters
Framework Support
| Framework | Differentiable | Compile/JIT | Versions |
|---|---|---|---|
| NumPy | 1.24+ | ||
| PyTorch | Yes | torch.compile | 2.0+ |
| JAX | Yes | jax.jit | 0.4+ |
| TensorFlow | Yes | tf.function | 2.13+ |
| MLX | Yes | mx.compile | 0.1+ |
Numerical Robustness
Standard SVD and eigendecomposition backward passes produce NaN gradients when singular values collide or are near-zero. The library provides custom autograd primitives to handle these cases:
- SafeSVD (PyTorch, JAX, TF, MLX): Custom backward pass that clamps the singular value gap, preventing division-by-zero in the gradient
- SafeEigh (PyTorch, JAX, TF, MLX): Analogous safe backward for the symmetric eigendecomposition used in Horn’s method
- Per-point weights: Weighted centroids and weighted cross-covariance for mass-weighted or confidence-weighted alignment
- Batch dimensions: All functions broadcast over leading batch dimensions without explicit loops
- Mixed-dtype promotion: Inputs are promoted to a common floating-point dtype automatically
Testing
The test suite uses Hypothesis-based property testing across 12 modules covering:
- Round-trip correctness (align then compare)
- Gradient finiteness and correctness (finite-difference checks)
- Reflection handling (proper vs. improper rotations)
- Weighted alignment consistency
- Batch broadcasting
- 4 differentiable backends $\times$ 4 precisions (float32, float64, and where supported, float16, bfloat16)
Usage
Install from GitHub:
pip install git+https://github.com/hunter-heidenreich/Kabsch-Cookbook.git
Basic alignment with NumPy:
import numpy as np
from kabsch_horn_cookbook.numpy import kabsch
# Two sets of corresponding 3D points
P = np.random.randn(100, 3)
R_true = np.linalg.qr(np.random.randn(3, 3))[0] # random rotation matrix
Q = (P @ R_true.T) + np.random.randn(1, 3)
R, t = kabsch(P, Q)
aligned = P @ R.T + t
RMSD loss for training in PyTorch:
import torch
from kabsch_horn_cookbook.torch import kabsch_rmsd
pred_coords = model(input_features) # (B, N, 3), requires_grad=True
target_coords = batch["target"] # (B, N, 3)
rmsd = kabsch_rmsd(pred_coords, target_coords) # (B,)
loss = rmsd.mean()
loss.backward() # safe gradients via SafeSVD
For the full API reference and additional examples, see the documentation site.
Results
Gradient Stability
The standard SVD backward pass computes terms of the form $\frac{1}{\sigma_i^2 - \sigma_j^2}$, which diverges when two singular values are close. In molecular alignment this happens frequently: planar molecules, symmetric structures, and noisy coordinates can all produce near-degenerate singular values. The SafeSVD primitive clamps the denominator to a configurable epsilon, producing finite (if slightly biased) gradients in these edge cases. Property-based tests confirm that gradients remain finite across thousands of random rotations, scales, and noise levels for all four differentiable backends.
Framework Parity
All five backends produce numerically equivalent results (up to floating-point tolerance) on the same inputs. The shared API means switching from NumPy prototyping to PyTorch training requires changing only the import path.
Related Work
This project builds on the foundational alignment algorithms described in these papers:
- Kabsch (1976): the original SVD-based rotation alignment
- Arun et al. (1987): SVD formulation for 3D point set fitting
- Horn (1987): quaternion-based closed-form absolute orientation
- Horn et al. (1988): orthonormal matrix (polar decomposition) approach
- Umeyama (1991): extension to include optimal scaling
For a detailed walkthrough of the Kabsch algorithm with code examples, see the companion blog post: The Kabsch Algorithm.