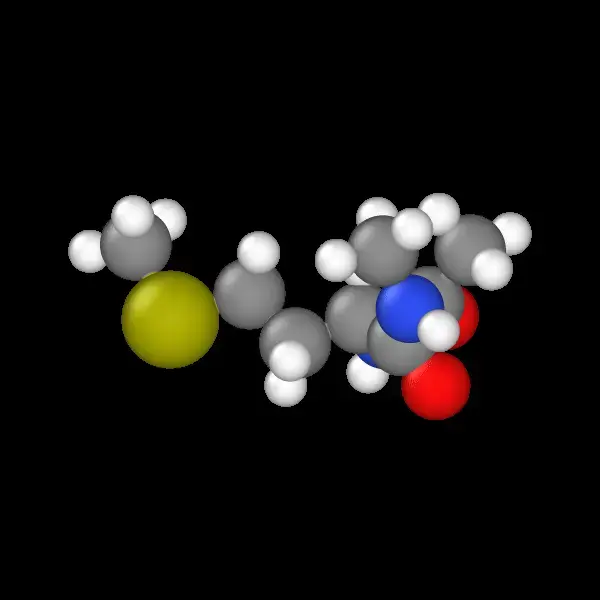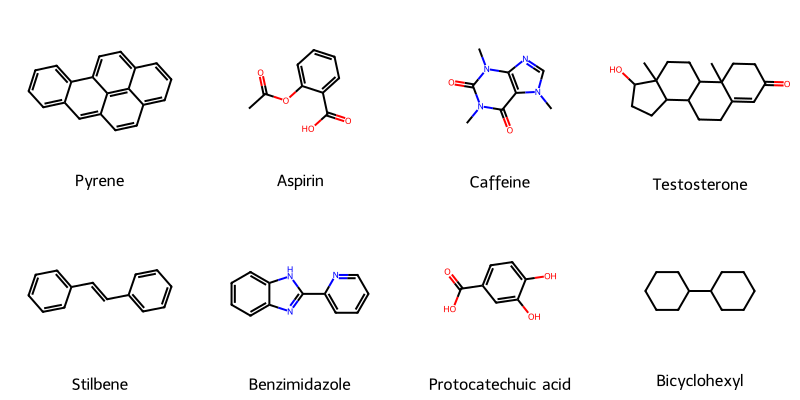
Molecular String Renderer: Robust Visualization Tool
A fault-tolerant RDKit wrapper treating molecular visualization as a software engineering problem, implementing strategy pattern for SVG generation with automatic raster fallback, native SELFIES support for generative AI workflows, and strict type safety for reliable batch processing of millions of molecules in training pipelines.